

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123898.1 - phase: 0 /pseudo
(177 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
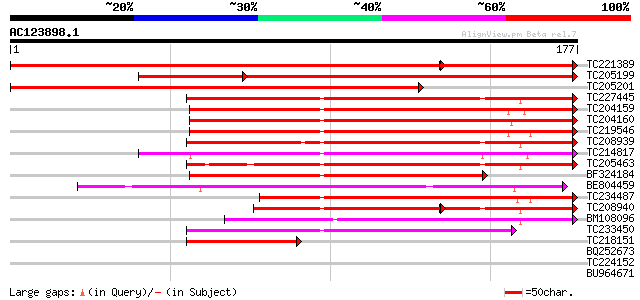
Sequences producing significant alignments: (bits) Value
TC221389 homologue to UP|HS7M_PHAVU (Q01899) Heat shock 70 kDa p... 231 2e-79
TC205199 homologue to UP|HS7M_PEA (P37900) Heat shock 70 kDa pro... 191 1e-62
TC205201 similar to UP|HS7M_PEA (P37900) Heat shock 70 kDa prote... 215 1e-56
TC227445 homologue to UP|O22639 (O22639) Endoplasmic reticulum H... 116 6e-27
TC204159 homologue to UP|Q40151 (Q40151) Hsc70 protein, partial ... 108 1e-24
TC204160 homologue to UP|HS72_LYCES (P27322) Heat shock cognate ... 108 1e-24
TC219546 UP|HS70_SOYBN (P26413) Heat shock 70 kDa protein, complete 107 3e-24
TC208939 UP|Q39830 (Q39830) BiP isoform A, complete 106 5e-24
TC214817 homologue to UP|Q9M4E7 (Q9M4E7) Heat shock protein 70, ... 105 1e-23
TC205463 UP|Q39804 (Q39804) BiP isoform B, complete 101 2e-22
BF324184 homologue to PIR|S53126|S53 dnaK-type molecular chapero... 95 2e-20
BE804459 similar to PIR|T10248|T10 heat shock protein 70K chlo... 95 2e-20
TC234487 similar to UP|Q9SAU8 (Q9SAU8) HSP70, partial (23%) 84 3e-17
TC208940 UP|Q39803 (Q39803) BiP isoform C (Fragment), partial (26%) 62 3e-16
BM108096 77 4e-15
TC233450 72 2e-13
TC218151 homologue to UP|O22639 (O22639) Endoplasmic reticulum H... 42 2e-04
BQ252673 homologue to SP|Q02028|HS7S_ Stromal 70 kDa heat shock-... 38 0.003
TC224152 similar to UP|Q8L8N8 (Q8L8N8) 70kD heat shock protein, ... 37 0.005
BU964671 32 0.12
>TC221389 homologue to UP|HS7M_PHAVU (Q01899) Heat shock 70 kDa protein,
mitochondrial precursor, partial (36%)
Length = 827
Score = 231 bits (590), Expect(2) = 2e-79
Identities = 114/136 (83%), Positives = 124/136 (90%)
Frame = +3
Query: 1 MAAATALLRSLRRRDAASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGID 60
+ A +LLRSLRRRD AS++++A+RSLTG+TKPAYV HNWSSLSRPFSSR AGNDVIGID
Sbjct: 87 LPAMASLLRSLRRRDVASASLSAYRSLTGSTKPAYVAHNWSSLSRPFSSRPAGNDVIGID 266
Query: 61 LGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISG 120
LGTTNSCVSVMEGKNPKV+ENSEGARTTPSVVAF QKGELLVGTP KRQAVTNP NT+ G
Sbjct: 267 LGTTNSCVSVMEGKNPKVIENSEGARTTPSVVAFNQKGELLVGTPXKRQAVTNPTNTLFG 446
Query: 121 AKRLIGRRFDDPQTQK 136
KRLIGRRFDD QTQK
Sbjct: 447 TKRLIGRRFDDAQTQK 494
Score = 81.3 bits (199), Expect(2) = 2e-79
Identities = 37/43 (86%), Positives = 41/43 (95%)
Frame = +1
Query: 135 QKEMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE 177
++EMKMVP+KIVKAPNG+AWVE GQQYSPSQIGAFVLTKMKE
Sbjct: 490 KREMKMVPFKIVKAPNGNAWVETNGQQYSPSQIGAFVLTKMKE 618
>TC205199 homologue to UP|HS7M_PEA (P37900) Heat shock 70 kDa protein,
mitochondrial precursor, partial (94%)
Length = 2204
Score = 191 bits (485), Expect(2) = 1e-62
Identities = 93/104 (89%), Positives = 97/104 (92%)
Frame = +3
Query: 74 KNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQ 133
+NPKV+ENSEGARTTPSVVAF QK ELLVGTPAKRQAVTNP NT+ G KRLIGRRFDD Q
Sbjct: 177 QNPKVIENSEGARTTPSVVAFNQKAELLVGTPAKRQAVTNPTNTLFGTKRLIGRRFDDSQ 356
Query: 134 TQKEMKMVPYKIVKAPNGDAWVEAKGQQYSPSQIGAFVLTKMKE 177
TQKEMKMVPYKIVKAPNGDAWVEA GQQYSPSQ+GAFVLTKMKE
Sbjct: 357 TQKEMKMVPYKIVKAPNGDAWVEANGQQYSPSQVGAFVLTKMKE 488
Score = 65.5 bits (158), Expect(2) = 1e-62
Identities = 31/34 (91%), Positives = 33/34 (96%)
Frame = +1
Query: 41 SSLSRPFSSRAAGNDVIGIDLGTTNSCVSVMEGK 74
+SLSRPFSS+ AGNDVIGIDLGTTNSCVSVMEGK
Sbjct: 1 ASLSRPFSSKPAGNDVIGIDLGTTNSCVSVMEGK 102
>TC205201 similar to UP|HS7M_PEA (P37900) Heat shock 70 kDa protein,
mitochondrial precursor, partial (18%)
Length = 467
Score = 215 bits (547), Expect = 1e-56
Identities = 108/129 (83%), Positives = 117/129 (89%)
Frame = +1
Query: 1 MAAATALLRSLRRRDAASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIGID 60
MAAATALLRSLRRRD SS+++AFRSLT TK +YVG+ W+SLSRPFSS+ AGNDVIGID
Sbjct: 79 MAAATALLRSLRRRDLPSSSLSAFRSLTSGTKTSYVGNKWASLSRPFSSKPAGNDVIGID 258
Query: 61 LGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISG 120
LGTTNSCVSVMEGKNPKV+ENSEGARTTPSVVAF QK ELLVGTPAKRQAVTNP NT+ G
Sbjct: 259 LGTTNSCVSVMEGKNPKVIENSEGARTTPSVVAFNQKAELLVGTPAKRQAVTNPTNTLFG 438
Query: 121 AKRLIGRRF 129
KRLIGRRF
Sbjct: 439 TKRLIGRRF 465
>TC227445 homologue to UP|O22639 (O22639) Endoplasmic reticulum HSC70-cognate
binding protein precursor, complete
Length = 2254
Score = 116 bits (290), Expect = 6e-27
Identities = 63/127 (49%), Positives = 84/127 (65%), Gaps = 5/127 (3%)
Frame = +2
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V + + +++ N +G R TPS VAFT E L+G AK A NPE
Sbjct: 215 VIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPSWVAFTDS-ERLIGEAAKNLAAVNPE 391
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK-----GQQYSPSQIGAF 170
TI KRLIGR+F+D + Q++MK+VPYKIV +G +++ K + +SP +I A
Sbjct: 392 RTIFDVKRLIGRKFEDKEVQRDMKLVPYKIVN-KDGKPYIQVKIKDGETKVFSPEEISAM 568
Query: 171 VLTKMKE 177
+LTKMKE
Sbjct: 569 ILTKMKE 589
>TC204159 homologue to UP|Q40151 (Q40151) Hsc70 protein, partial (22%)
Length = 769
Score = 108 bits (271), Expect = 1e-24
Identities = 61/125 (48%), Positives = 77/125 (60%), Gaps = 4/125 (3%)
Frame = +2
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 107 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPIN 283
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEAKG--QQYSPSQIGAFVL 172
T+ AKRLIGRRF D Q +MK+ P+K++ P V KG +Q+S +I + VL
Sbjct: 284 TVFDAKRLIGRRFSDASVQSDMKLWPFKVIPGPADKPMIVVNYKGDEKQFSAEEISSMVL 463
Query: 173 TKMKE 177
KMKE
Sbjct: 464 IKMKE 478
>TC204160 homologue to UP|HS72_LYCES (P27322) Heat shock cognate 70 kDa
protein 2, partial (66%)
Length = 1390
Score = 108 bits (271), Expect = 1e-24
Identities = 58/125 (46%), Positives = 77/125 (61%), Gaps = 4/125 (3%)
Frame = +2
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP N
Sbjct: 122 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPTN 298
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWV----EAKGQQYSPSQIGAFVL 172
T+ AKRLIGRRF D Q +MK+ P+K++ P + + + +Q+S +I + VL
Sbjct: 299 TVFDAKRLIGRRFSDASVQGDMKLWPFKVIPGPAEKPMIVVNYKGEEKQFSAEEISSMVL 478
Query: 173 TKMKE 177
KMKE
Sbjct: 479 MKMKE 493
>TC219546 UP|HS70_SOYBN (P26413) Heat shock 70 kDa protein, complete
Length = 2099
Score = 107 bits (267), Expect = 3e-24
Identities = 59/125 (47%), Positives = 78/125 (62%), Gaps = 4/125 (3%)
Frame = +1
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS VAFT E L+G AK Q NP+N
Sbjct: 25 IGIDLGTTYSCVGVWQNDRVEIIPNDQGNRTTPSYVAFTDT-ERLIGDAAKNQVAMNPQN 201
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAW--VEAKGQQ--YSPSQIGAFVL 172
T+ AKRLIGRRF D Q +MK+ P+K+ +P V KG++ +S +I + VL
Sbjct: 202 TVFDAKRLIGRRFSDSSVQNDMKLWPFKVGGSPCDKPMIVVNYKGEEKKFSAEEISSMVL 381
Query: 173 TKMKE 177
KM+E
Sbjct: 382 VKMRE 396
>TC208939 UP|Q39830 (Q39830) BiP isoform A, complete
Length = 2452
Score = 106 bits (265), Expect = 5e-24
Identities = 60/127 (47%), Positives = 82/127 (64%), Gaps = 5/127 (3%)
Frame = +3
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDLGTT SCV V + + +++ N +G R TPS +FT E L+G AK A NPE
Sbjct: 177 VIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPSW-SFTDS-ERLIGEAAKNLAAVNPE 350
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK-----GQQYSPSQIGAF 170
I KRLIGR+F+D + Q++MK+VPYKIV +G +++ K + +SP +I A
Sbjct: 351 RVIFDVKRLIGRKFEDKEVQRDMKLVPYKIVN-KDGKPYIQEKIKDGETKVFSPEEISAM 527
Query: 171 VLTKMKE 177
+LTKMKE
Sbjct: 528 ILTKMKE 548
>TC214817 homologue to UP|Q9M4E7 (Q9M4E7) Heat shock protein 70, complete
Length = 2364
Score = 105 bits (262), Expect = 1e-23
Identities = 63/144 (43%), Positives = 83/144 (56%), Gaps = 7/144 (4%)
Frame = +2
Query: 41 SSLSRPFSSRAAGND---VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQK 97
SS S P + AG IGIDLGTT SCV V + +++ N +G RTTPS V FT
Sbjct: 56 SSFSTPHAETMAGKGEGPAIGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVGFTDT 235
Query: 98 GELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKMVPYKIV--KAPNGDAWV 155
E L+G AK Q NP NT+ AKRLIGRRF D Q ++K+ P+K++ A V
Sbjct: 236 -ERLIGDAAKNQVAMNPINTVFDAKRLIGRRFSDSSVQSDIKLWPFKVIPGAADKPMIVV 412
Query: 156 EAKGQ--QYSPSQIGAFVLTKMKE 177
KG+ Q++ +I + VL KM+E
Sbjct: 413 NYKGEEKQFAAEEISSMVLIKMRE 484
>TC205463 UP|Q39804 (Q39804) BiP isoform B, complete
Length = 2383
Score = 101 bits (251), Expect = 2e-22
Identities = 61/127 (48%), Positives = 81/127 (63%), Gaps = 5/127 (3%)
Frame = +3
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
VIGIDL TT SCV V + +++ N +G R TPS VAFT E L+G AK A NP
Sbjct: 198 VIGIDL-TTYSCVGVTDAV--EIIANDQGNRITPSWVAFTDS-ERLIGEAAKIVAAVNPV 365
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK-----GQQYSPSQIGAF 170
TI KRLIGR+F+D + Q++MK+VPYKIV +G +++ K + +SP +I A
Sbjct: 366 RTIFDVKRLIGRKFEDKEVQRDMKLVPYKIVN-KDGKPYIQVKIKDGETKVFSPEEISAM 542
Query: 171 VLTKMKE 177
+LTKMKE
Sbjct: 543 ILTKMKE 563
>BF324184 homologue to PIR|S53126|S53 dnaK-type molecular chaperone hsp70 -
rice (fragment), partial (18%)
Length = 398
Score = 95.1 bits (235), Expect = 2e-20
Identities = 47/93 (50%), Positives = 60/93 (63%)
Frame = -3
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGIDLGTT SCV V + +++ N +G RTTPS V FT E L+G AK Q NP N
Sbjct: 342 IGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSYVGFTDT-ERLIGDAAKNQVAMNPIN 166
Query: 117 TISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAP 149
T+ AKRLIGRRF D Q ++K+ P+K++ P
Sbjct: 165 TVFDAKRLIGRRFSDASVQSDIKLWPFKVLSGP 67
>BE804459 similar to PIR|T10248|T10 heat shock protein 70K chloroplast -
cucumber, partial (18%)
Length = 478
Score = 94.7 bits (234), Expect = 2e-20
Identities = 60/156 (38%), Positives = 85/156 (54%), Gaps = 3/156 (1%)
Frame = +2
Query: 22 TAFRSLTGNTKPAYVGHNWSSLSRPFSSRAAGNDVIG-IDLGTTNSCVSVMEGKNPKVVE 80
T F NTK A++ S R N+ + IDLGTTN V+ G P ++
Sbjct: 8 TLFLGQRLNTKAAFI--KLKSTPRRLRPLIVVNEKVACIDLGTTNCSVAAFVGGKPTIIS 181
Query: 81 NSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEMKM 140
NS G +TTP V A+T G+ LVG AKRQ V NPENT+ +R IG + + +E K
Sbjct: 182 NSRGQKTTPFVRAYTNNGDRLVGLIAKRQVVVNPENTLFFVERFIGPKMS--EVDEESKH 355
Query: 141 VPYKIVKAPNGDAWVE--AKGQQYSPSQIGAFVLTK 174
V Y++++ NG+ ++ A G+Q + +I A VLTK
Sbjct: 356 VSYRVIRDDNGNVKLDCPAIGKQCAADEISAQVLTK 463
>TC234487 similar to UP|Q9SAU8 (Q9SAU8) HSP70, partial (23%)
Length = 457
Score = 84.0 bits (206), Expect = 3e-17
Identities = 46/103 (44%), Positives = 63/103 (60%), Gaps = 4/103 (3%)
Frame = +1
Query: 79 VENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQKEM 138
+ N +G TTPS VAFT + + L+G AK QA TNPENT+ AKRLIGR+F DP QK+
Sbjct: 1 IHNDQGNNTTPSCVAFTDQ-QRLIGEAAKNQAATNPENTVFDAKRLIGRKFSDPVIQKDK 177
Query: 139 KMVPYKIVKAPNGDAWVEA--KGQQ--YSPSQIGAFVLTKMKE 177
+ P+K+V N + KGQ+ ++ + VL KM+E
Sbjct: 178 MLWPFKVVAGINDKPMISLNYKGQEKHLLAEEVSSMVLIKMRE 306
>TC208940 UP|Q39803 (Q39803) BiP isoform C (Fragment), partial (26%)
Length = 447
Score = 61.6 bits (148), Expect(2) = 3e-16
Identities = 31/60 (51%), Positives = 39/60 (64%)
Frame = +1
Query: 77 KVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGRRFDDPQTQK 136
+++ N +G R TPS VAFT E L+G AK QA NPE TI KRLIGR+F+D + K
Sbjct: 88 EIIANDQGNRITPSWVAFTDS-ERLIGEAAKNQAAVNPERTIFDVKRLIGRKFEDKEISK 264
Score = 39.7 bits (91), Expect(2) = 3e-16
Identities = 22/48 (45%), Positives = 31/48 (63%), Gaps = 5/48 (10%)
Frame = +2
Query: 135 QKEMKMVPYKIVKAPNGDAWVEAK-----GQQYSPSQIGAFVLTKMKE 177
QK+MK+VPYKIV +G +++ K + +SP +I A VL KMKE
Sbjct: 260 QKDMKLVPYKIVN-KDGKPYIQVKIKDGETKVFSPEEISAMVLIKMKE 400
>BM108096
Length = 525
Score = 77.0 bits (188), Expect = 4e-15
Identities = 40/114 (35%), Positives = 66/114 (57%), Gaps = 4/114 (3%)
Frame = +3
Query: 68 VSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPENTISGAKRLIGR 127
V+V + VV N E R TP++V F K L GT + NP+N+IS KRLIGR
Sbjct: 9 VAVARQRGIDVVLNDESKRETPAIVCFGDKQRFL-GTAGAASTMMNPKNSISQIKRLIGR 185
Query: 128 RFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK----GQQYSPSQIGAFVLTKMKE 177
+F DP+ Q+++K P+ + + P+G + A+ + ++P+Q+ +L+ +KE
Sbjct: 186 QFSDPELQRDLKTFPFVVTEGPDGYPLIXARYLGEARTFTPTQVFGMMLSNLKE 347
>TC233450
Length = 677
Score = 71.6 bits (174), Expect = 2e-13
Identities = 38/103 (36%), Positives = 61/103 (58%)
Frame = +3
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPE 115
V+G D+G N ++V+ + V+ N E R TP+VV F++K + + G+ A+ + +
Sbjct: 228 VVGFDIGNENCVIAVVRQRGIDVLLNYESKRETPAVVCFSEK-QRI*GSAGAASAMMHIK 404
Query: 116 NTISGAKRLIGRRFDDPQTQKEMKMVPYKIVKAPNGDAWVEAK 158
+TIS KRLIGR+F DP +KE+KM+P K + G + K
Sbjct: 405 STISQIKRLIGRKFADPDVKKELKMLPGKTSEGQEGGISIHLK 533
>TC218151 homologue to UP|O22639 (O22639) Endoplasmic reticulum HSC70-cognate
binding protein precursor, partial (35%)
Length = 922
Score = 42.0 bits (97), Expect = 2e-04
Identities = 18/36 (50%), Positives = 25/36 (69%)
Frame = +1
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSV 91
VIGIDLGTT SCV V + + +++ N +G R TP +
Sbjct: 196 VIGIDLGTTYSCVGVYKNGHVEIIANDQGNRITPQI 303
>BQ252673 homologue to SP|Q02028|HS7S_ Stromal 70 kDa heat shock-related
protein chloroplast precursor. [Garden pea] {Pisum
sativum}, partial (6%)
Length = 302
Score = 37.7 bits (86), Expect = 0.003
Identities = 27/74 (36%), Positives = 36/74 (48%), Gaps = 1/74 (1%)
Frame = +1
Query: 1 MAAATALLRSLRRRDAASSTVTAFRSLTGNTKPAYVGHNWSSLSRPFSS-RAAGNDVIGI 59
MA ++A + L S + T F NTK A++ + R R V+GI
Sbjct: 82 MACSSAQIHGL---GTPSFSRTLFLGQRLNTKAAFIKVKSAPTPRRLRPLRVVNEKVVGI 252
Query: 60 DLGTTNSCVSVMEG 73
DLGTTNS V+ MEG
Sbjct: 253 DLGTTNSAVAAMEG 294
>TC224152 similar to UP|Q8L8N8 (Q8L8N8) 70kD heat shock protein, partial
(33%)
Length = 563
Score = 37.0 bits (84), Expect = 0.005
Identities = 24/81 (29%), Positives = 40/81 (48%), Gaps = 5/81 (6%)
Frame = +1
Query: 57 IGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAFTQKGELLVGTPAKRQAVTNPEN 116
IGID+GT+ V+V G ++++N+ + S V F + + +++ +
Sbjct: 37 IGIDIGTSQCSVAVWNGSQVELLKNTRNQKIMKSYVTFKDN----IPSGGVSSQLSHEDE 204
Query: 117 TISGA-----KRLIGRRFDDP 132
+SGA KRLIGR DP
Sbjct: 205 MLSGATIFNMKRLIGRVDTDP 267
>BU964671
Length = 181
Score = 32.3 bits (72), Expect = 0.12
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +1
Query: 56 VIGIDLGTTNSCVSVMEGKNPKVVENSEGARTTPSVVAF 94
V+G D G + V+V + VV N E R TP++V F
Sbjct: 64 VVGFDFGNESCIVAVARQRGIDVVLNDESKRETPAIVCF 180
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.127 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,238,509
Number of Sequences: 63676
Number of extensions: 72604
Number of successful extensions: 321
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 306
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 306
length of query: 177
length of database: 12,639,632
effective HSP length: 91
effective length of query: 86
effective length of database: 6,845,116
effective search space: 588679976
effective search space used: 588679976
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC123898.1


