

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123575.7 - phase: 0 /pseudo
(378 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
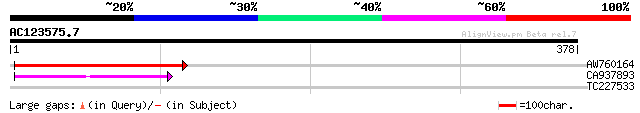
AW760164 similar to GP|11994422|dbj oxidoreductase short-chain ... 95 5e-20
CA937893 similar to GP|20805072|dbj retrovirus-related pol polyp... 53 2e-07
TC227533 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17... 30 2.3
>AW760164 similar to GP|11994422|dbj oxidoreductase short-chain
dehydrogenase/reductase family-like protein {Arabidopsis
thaliana}, partial (9%)
Length = 428
Score = 95.1 bits (235), Expect = 5e-20
Identities = 48/115 (41%), Positives = 73/115 (62%)
Frame = +3
Query: 4 SKWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAI 63
+K+ +EKFTG NDFGL +KMR +L+QQ VEAL G+ ++ + +K + KA +AI
Sbjct: 84 AKYEVEKFTGQNDFGLC*LKMRALLVQQGLVEALDGEIKLEKMMADGDKKALLQKAYNAI 263
Query: 64 VLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLYFYRMVESKTN 118
+L LGDKVLR+V++ET V +W+KL+ L + T C + Y + E ++N
Sbjct: 264 ILSLGDKVLRQVSKETTAVGVWSKLEVLDIDSVGTFTMCHEALKYLKKGGEGRSN 428
>CA937893 similar to GP|20805072|dbj retrovirus-related pol polyprotein from
transposon TNT 1-94-like, partial (7%)
Length = 412
Score = 53.1 bits (126), Expect = 2e-07
Identities = 31/105 (29%), Positives = 55/105 (51%)
Frame = -2
Query: 4 SKWNIEKFTGSNDFGLWKVKMRIILIQQKCVEALKGKAQMPAHLTPAEKTEMNDKAISAI 63
+K+ + KF G+ +F LW+ +++ +L Q ++ L+ E E+ ++ + I
Sbjct: 312 AKFEVGKFDGTGNFRLWQKRVKDLLA*QGLLKVLRDSKSNNTEALDWE--EL*ERTATTI 139
Query: 64 VLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQCLKQQLY 108
LCL D+ L + +W KL+S YM KS T++ L Q+LY
Sbjct: 138 RLCLVDEFLYHMMELAFPGEVWKKLESQYMLKSLTNKLYLMQKLY 4
>TC227533 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17H15.1
{Arabidopsis thaliana;} , partial (4%)
Length = 1519
Score = 29.6 bits (65), Expect = 2.3
Identities = 12/42 (28%), Positives = 24/42 (56%)
Frame = -3
Query: 60 ISAIVLCLGDKVLREVARETIVVSMWNKLDSLYMTKSWTHRQ 101
IS + LG LR TI++++W + ++++ W+HR+
Sbjct: 512 ISLSITLLGKSCLRISCLMTIILALWLSIVVIHLSWGWSHRR 387
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.356 0.157 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,226,733
Number of Sequences: 63676
Number of extensions: 186168
Number of successful extensions: 1698
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 1694
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1698
length of query: 378
length of database: 12,639,632
effective HSP length: 99
effective length of query: 279
effective length of database: 6,335,708
effective search space: 1767662532
effective search space used: 1767662532
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC123575.7


