

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123571.7 - phase: 0
(247 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
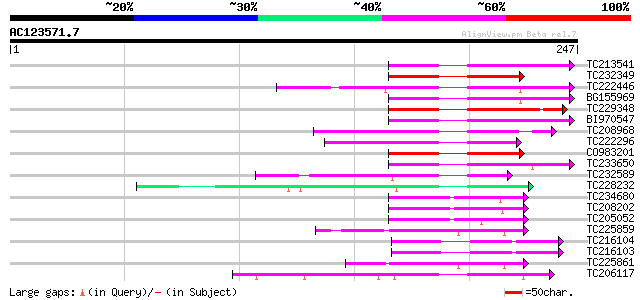
Score E
Sequences producing significant alignments: (bits) Value
TC213541 similar to UP|Q9LES3 (Q9LES3) Promoter-binding factor-l... 69 2e-12
TC232349 similar to UP|O81241 (O81241) Dc3 promoter-binding fact... 67 1e-11
TC222446 similar to UP|O81241 (O81241) Dc3 promoter-binding fact... 66 1e-11
BG155969 similar to PIR|T12622|T126 Dc3 promoter-binding factor-... 66 2e-11
TC229348 homologue to UP|Q94KA6 (Q94KA6) BZIP transcription fact... 63 1e-10
BI970547 weakly similar to GP|29027731|dbj bZIP transcription fa... 55 3e-08
TC208968 similar to UP|GBF4_ARATH (P42777) G-box binding factor ... 54 9e-08
TC222296 similar to UP|P93426 (P93426) Leucine zipper protein, p... 54 9e-08
CO983201 53 1e-07
TC233650 similar to UP|Q84JC9 (Q84JC9) BZIP transcription factor... 53 1e-07
TC232589 similar to UP|Q8LK78 (Q8LK78) ABA response element bind... 46 1e-05
TC228232 homologue to UP|Q43449 (Q43449) G-box binding factor, c... 45 4e-05
TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor... 43 2e-04
TC208202 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor... 42 3e-04
TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 42 3e-04
TC225859 similar to UP|O22677 (O22677) BZIP DNA-binding protein,... 42 3e-04
TC216104 42 3e-04
TC216103 similar to GB|AAK84220.1|15100049|AF401297 transcriptio... 42 3e-04
TC225861 similar to UP|Q941L6 (Q941L6) BZIP transcription factor... 41 4e-04
TC206117 weakly similar to UP|Q9FUD3 (Q9FUD3) BZIP protein BZO2H... 41 4e-04
>TC213541 similar to UP|Q9LES3 (Q9LES3) Promoter-binding factor-like protein
(ABA-responsive element binding protein 3) (AREB3),
partial (31%)
Length = 741
Score = 69.3 bits (168), Expect = 2e-12
Identities = 45/81 (55%), Positives = 47/81 (57%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR KRMIKNRESAARSRARKQ AYT ELE KV L+EEN RLRRQ +
Sbjct: 61 ERRQKRMIKNRESAARSRARKQ------------AYTQELEIKVSQLEEENERLRRQNEI 204
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
S K L RTSS
Sbjct: 205 ERALPSAPPPDPKHQLRRTSS 267
>TC232349 similar to UP|O81241 (O81241) Dc3 promoter-binding factor-3
(Fragment), partial (50%)
Length = 834
Score = 66.6 bits (161), Expect = 1e-11
Identities = 37/59 (62%), Positives = 42/59 (70%)
Frame = +3
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQ 224
ERR KRMIKNRESAARSRARKQ AYTNELE KV L+EEN RLR++++
Sbjct: 408 ERRQKRMIKNRESAARSRARKQ------------AYTNELENKVSRLEEENERLRKRKE 548
>TC222446 similar to UP|O81241 (O81241) Dc3 promoter-binding factor-3
(Fragment), partial (46%)
Length = 765
Score = 66.2 bits (160), Expect = 1e-11
Identities = 53/134 (39%), Positives = 67/134 (49%), Gaps = 4/134 (2%)
Frame = +3
Query: 117 QQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDI--SPGERRNKRMIK 174
QQ +GV+ V + +P A S G++ G + + ERR KRMIK
Sbjct: 222 QQPFQVGVNLVLDAAYSETPASLK---GALSDTQTLGRKRGVSGIVVEKTVERRQKRMIK 392
Query: 175 NRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRR--QQQELWEAESG 232
NRESAARSRAR+Q AYT ELE KV L+EEN RLRR + + +
Sbjct: 393 NRESAARSRARRQ------------AYTQELEIKVSRLEEENERLRRLNEMERALPSVPP 536
Query: 233 GQQKKKSSLYRTSS 246
+ K K L RTSS
Sbjct: 537 PEPKPKHQLRRTSS 578
>BG155969 similar to PIR|T12622|T126 Dc3 promoter-binding factor-2 - common
sunflower, partial (43%)
Length = 414
Score = 65.9 bits (159), Expect = 2e-11
Identities = 43/83 (51%), Positives = 49/83 (58%), Gaps = 2/83 (2%)
Frame = +2
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRR--QQ 223
ERR KRMIKNRESAARSRAR+Q AYT ELE KV L+EEN RLRR +
Sbjct: 14 ERRQKRMIKNRESAARSRARRQ------------AYTQELEIKVSRLEEENERLRRLNEM 157
Query: 224 QELWEAESGGQQKKKSSLYRTSS 246
+ + + K K L RTSS
Sbjct: 158 ERALPSVPPPEPKPKQQLRRTSS 226
>TC229348 homologue to UP|Q94KA6 (Q94KA6) BZIP transcription factor 6,
partial (34%)
Length = 786
Score = 62.8 bits (151), Expect = 1e-10
Identities = 39/78 (50%), Positives = 49/78 (62%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
ERR +RMIKNRESAARSRARKQ AYT ELE +V L+EEN L+++Q E
Sbjct: 241 ERRQRRMIKNRESAARSRARKQ------------AYTMELEAEVAKLKEENEELQKKQAE 384
Query: 226 LWEAESGGQQKKKSSLYR 243
+ E + Q K+ +L R
Sbjct: 385 IMEIQK-NQVKEMMNLQR 435
>BI970547 weakly similar to GP|29027731|dbj bZIP transcription factor
{Arabidopsis thaliana}, partial (15%)
Length = 697
Score = 55.1 bits (131), Expect = 3e-08
Identities = 34/81 (41%), Positives = 46/81 (55%)
Frame = -1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
++R+ R+IKNRESA RSRARKQ AY LE ++ L EEN+RL+RQ +E
Sbjct: 493 DQRHTRVIKNRESAVRSRARKQ------------AYRKGLEVEISRLTEENSRLKRQLKE 350
Query: 226 LWEAESGGQQKKKSSLYRTSS 246
L + ++ RTSS
Sbjct: 349 LQRCLCSSHTPRMAAPCRTSS 287
>TC208968 similar to UP|GBF4_ARATH (P42777) G-box binding factor 4
(AtbZIP40), partial (27%)
Length = 1199
Score = 53.5 bits (127), Expect = 9e-08
Identities = 40/106 (37%), Positives = 53/106 (49%)
Frame = +2
Query: 133 NASPCDQNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITS 192
+ P V S+S +R E P ++ +RMIKNRESAARSR RKQ
Sbjct: 437 SVEPFANGVSAAPSNSVQKGKRRAVEEPVDKATLQKQRRMIKNRESAARSRERKQ----- 601
Query: 193 FLFSKFSAYTNELEQKVQLLQEENARLRRQQQELWEAESGGQQKKK 238
AYT+ELE V L++EN +L + EAE Q+KK+
Sbjct: 602 -------AYTSELEYLVHQLEQENVQLLNE-----EAEMRRQRKKQ 703
>TC222296 similar to UP|P93426 (P93426) Leucine zipper protein, partial (24%)
Length = 945
Score = 53.5 bits (127), Expect = 9e-08
Identities = 36/86 (41%), Positives = 48/86 (54%)
Frame = +3
Query: 138 DQNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFLFSK 197
+ VG S+S +R E P ++ +RMIKNRESAARSR RKQ
Sbjct: 405 NNGVGSAPSNSVQKGKRRAVEEPVDKATLQKLRRMIKNRESAARSRERKQ---------- 554
Query: 198 FSAYTNELEQKVQLLQEENARLRRQQ 223
AYT+ELE V L++ENARL +++
Sbjct: 555 --AYTSELEYLVHQLEQENARLLKEE 626
Score = 32.3 bits (72), Expect = 0.20
Identities = 28/83 (33%), Positives = 41/83 (48%), Gaps = 10/83 (12%)
Frame = +3
Query: 1 MEEVWKDINLSSLNDQNTRPMIMSTRNSTFGGGVILQDFLTR--PLTLD--------PTK 50
M+++ K I +S++D ST N+ GGV L+DFLT+ P+T + P
Sbjct: 183 MDDLLKSITPNSVDDFWKGIAAASTDNA---GGVTLEDFLTKAIPVTEEDVRGAPPPPPS 353
Query: 51 SLDYSSNNSSSSVASDQNNNNAS 73
L + + SSSSV NN S
Sbjct: 354 FLPFPAEGSSSSVEPFANNGVGS 422
>CO983201
Length = 774
Score = 52.8 bits (125), Expect = 1e-07
Identities = 30/59 (50%), Positives = 38/59 (63%)
Frame = -3
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQ 224
ERR +R IKNRE AARSRARKQ AY NEL KV L+EEN +L+++++
Sbjct: 718 ERRLRRKIKNREXAARSRARKQ------------AYHNELVGKVSRLEEENVKLKKEKE 578
>TC233650 similar to UP|Q84JC9 (Q84JC9) BZIP transcription factor, partial
(17%)
Length = 792
Score = 52.8 bits (125), Expect = 1e-07
Identities = 35/84 (41%), Positives = 48/84 (56%), Gaps = 3/84 (3%)
Frame = +3
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQE 225
++R+ R+IKNRESA RSRARKQ AY LE ++ L EEN+RL+RQ +E
Sbjct: 333 DQRHARIIKNRESAVRSRARKQ------------AYRKGLEVEIARLTEENSRLKRQLKE 476
Query: 226 L---WEAESGGQQKKKSSLYRTSS 246
L + + ++L RTSS
Sbjct: 477 LQCCLSSSDNPPTPRMAALCRTSS 548
>TC232589 similar to UP|Q8LK78 (Q8LK78) ABA response element binding factor,
partial (10%)
Length = 464
Score = 46.2 bits (108), Expect = 1e-05
Identities = 39/116 (33%), Positives = 55/116 (46%), Gaps = 4/116 (3%)
Frame = +2
Query: 108 HNNSQLLLQQQQHNIGVSNVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDISPG-- 165
+++ L L H++ VS+ F+N SSS + F K +S
Sbjct: 164 NSSVDLDLNNNNHSVSVSS----FLNQPLSTFLTLTSTSSSSSVFHKHDHSLLSVSDPNT 331
Query: 166 --ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARL 219
++R+ R+IKNRESA RSRARKQ AY LE ++ L EEN+RL
Sbjct: 332 LQDQRHTRVIKNRESAVRSRARKQ------------AYRKGLEVEISRLTEENSRL 463
>TC228232 homologue to UP|Q43449 (Q43449) G-box binding factor, complete
Length = 1723
Score = 44.7 bits (104), Expect = 4e-05
Identities = 53/188 (28%), Positives = 73/188 (38%), Gaps = 15/188 (7%)
Frame = +1
Query: 56 SNNSSSSVASDQNNNNASFYCPISTTPPPLVTALSLNSRPDFLYDPLIRHNKHNNSQLLL 115
S + SS ASD+N N + N + F + N NNS ++
Sbjct: 499 SGSEGSSDASDENTNQQES---------------ATNKKGSFDQMLVDGANARNNSVSII 633
Query: 116 QQQQH----------NIGVS--NVSPCFVNASPCDQNVGVPASSSFTCFGKRFGEAPDIS 163
Q + NIG++ N SP A+ QN A + T G+
Sbjct: 634 PQPGNPAVSMSPTSLNIGMNLWNASPAGDEAAKMRQNQSSGAVTPPTIMGREVALGEHWI 813
Query: 164 PGER---RNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLR 220
ER + KR NRESA RSR RKQ A EL+++V+ L EN LR
Sbjct: 814 QDERELKKQKRKQSNRESARRSRLRKQ------------AECEELQKRVESLGSENQTLR 957
Query: 221 RQQQELWE 228
+ Q + E
Sbjct: 958 EELQRVSE 981
>TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor, partial
(49%)
Length = 571
Score = 42.7 bits (99), Expect = 2e-04
Identities = 29/71 (40%), Positives = 40/71 (55%), Gaps = 10/71 (14%)
Frame = +2
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLL----------QEE 215
ER+++RMI NRESA RSR RKQ+ + L+S+ NE Q + L +E
Sbjct: 344 ERKHRRMISNRESARRSRMRKQKHLDE-LWSQVVWLRNENHQLMDKLNHVSESHDQVMQE 520
Query: 216 NARLRRQQQEL 226
NA+L+ Q EL
Sbjct: 521 NAQLKEQALEL 553
>TC208202 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor, partial
(51%)
Length = 758
Score = 42.0 bits (97), Expect = 3e-04
Identities = 28/71 (39%), Positives = 39/71 (54%), Gaps = 10/71 (14%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQ----------EE 215
ER+++RMI NRESA RSR RKQ+ + L+S+ NE Q + L +E
Sbjct: 229 ERKHRRMISNRESARRSRMRKQKHLDE-LWSQVVWLRNENHQLMDKLNHVSESHDKVAQE 405
Query: 216 NARLRRQQQEL 226
N +LR + EL
Sbjct: 406 NVQLREEASEL 438
>TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (58%)
Length = 953
Score = 42.0 bits (97), Expect = 3e-04
Identities = 29/71 (40%), Positives = 38/71 (52%), Gaps = 10/71 (14%)
Frame = +1
Query: 166 ERRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNE----------LEQKVQLLQEE 215
+R+ KRMI NRESA RSR RKQ+ + L S+ + NE QK ++ E
Sbjct: 598 QRKRKRMISNRESARRSRMRKQKHLDD-LASQVTQLRNENHQILTSVNLTTQKYLAVEAE 774
Query: 216 NARLRRQQQEL 226
N+ LR Q EL
Sbjct: 775 NSVLRAQVNEL 807
>TC225859 similar to UP|O22677 (O22677) BZIP DNA-binding protein, partial
(59%)
Length = 1132
Score = 42.0 bits (97), Expect = 3e-04
Identities = 34/102 (33%), Positives = 49/102 (47%), Gaps = 9/102 (8%)
Frame = +1
Query: 134 ASPCDQNVGVPASSSFTCFGKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSF 193
ASP Q S+ T G G+ I ER+ KRM+ NRESA RSR RKQ+++
Sbjct: 421 ASPIQQQ----QRSTTTSSGSEGGDPHIID--ERKRKRMLSNRESARRSRMRKQKQLEDL 582
Query: 194 L--FSKFSAYTNELEQKVQLLQE-------ENARLRRQQQEL 226
S+ + +L + ++ +E N+ LR Q EL
Sbjct: 583 TDEVSRLQSANKKLAENIEAKEEACVETEAANSILRAQTMEL 708
>TC216104
Length = 760
Score = 41.6 bits (96), Expect = 3e-04
Identities = 29/75 (38%), Positives = 38/75 (50%)
Frame = +2
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R KR++ NR+SAARS+ RK Y +ELE KVQ LQ E L Q L
Sbjct: 26 KRAKRILANRQSAARSKERKMR------------YISELEHKVQTLQTEATTL-SAQLTL 166
Query: 227 WEAESGGQQKKKSSL 241
+ +S G + S L
Sbjct: 167 LQRDSAGLTNQNSEL 211
>TC216103 similar to GB|AAK84220.1|15100049|AF401297 transcription factor
bZIP29 {Arabidopsis thaliana;} , partial (34%)
Length = 1336
Score = 41.6 bits (96), Expect = 3e-04
Identities = 29/75 (38%), Positives = 38/75 (50%)
Frame = +1
Query: 167 RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQKVQLLQEENARLRRQQQEL 226
+R KR++ NR+SAARS+ RK Y +ELE KVQ LQ E L Q L
Sbjct: 601 KRAKRILANRQSAARSKERKMR------------YISELEHKVQTLQTEATTL-SAQLTL 741
Query: 227 WEAESGGQQKKKSSL 241
+ +S G + S L
Sbjct: 742 LQRDSAGLTNQNSEL 786
>TC225861 similar to UP|Q941L6 (Q941L6) BZIP transcription factor BZI-3,
partial (43%)
Length = 1239
Score = 41.2 bits (95), Expect = 4e-04
Identities = 30/89 (33%), Positives = 44/89 (48%), Gaps = 9/89 (10%)
Frame = +3
Query: 147 SSFTCFGKRFGEAPDISPGERRNKRMIKNRESAARSRARKQEKITSFL--FSKFSAYTNE 204
S+ T G G P + ER+ KRM+ NRESA RSR RKQ+++ S+ +
Sbjct: 465 STATSSGSEGGGDPQMID-ERKRKRMLSNRESARRSRMRKQKQLEDLTDEVSRLQGANKK 641
Query: 205 LEQKVQLLQE-------ENARLRRQQQEL 226
L + ++ +E N+ LR Q EL
Sbjct: 642 LAENIKAKEEACVETEAANSILRAQTMEL 728
>TC206117 weakly similar to UP|Q9FUD3 (Q9FUD3) BZIP protein BZO2H2, partial
(48%)
Length = 1225
Score = 41.2 bits (95), Expect = 4e-04
Identities = 46/153 (30%), Positives = 66/153 (43%), Gaps = 13/153 (8%)
Frame = +3
Query: 98 LYDPLIRHN---KHNNSQLLLQQQQHNIGVSNV-SPCFVNASPCDQNVGVPASSSFTCFG 153
L DPL N KH+ + Q SNV SP N +N A+S +
Sbjct: 186 LIDPLCSQNLTPKHSTITATIDSQSSICATSNVGSPVSANKPEGRENHTKGATSGSSEPS 365
Query: 154 KRFGEA----PDISPGE-RRNKRMIKNRESAARSRARKQEKITSFLFSKFSAYTNELEQK 208
EA +P + +R +R + NR+SA RSR RKQ A ++LE +
Sbjct: 366 DEDDEAGACEQSTNPADMKRLRRKVSNRDSARRSRRRKQ------------AQLSDLELQ 509
Query: 209 VQLLQEENARLRRQ----QQELWEAESGGQQKK 237
V+ L+ ENA L +Q Q EA++ + K
Sbjct: 510 VEKLKVENATLYKQFTDASQHFREADTNNRVLK 608
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.127 0.361
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,356,842
Number of Sequences: 63676
Number of extensions: 166256
Number of successful extensions: 1252
Number of sequences better than 10.0: 117
Number of HSP's better than 10.0 without gapping: 1181
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1219
length of query: 247
length of database: 12,639,632
effective HSP length: 95
effective length of query: 152
effective length of database: 6,590,412
effective search space: 1001742624
effective search space used: 1001742624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC123571.7


