

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123570.7 + phase: 0
(601 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
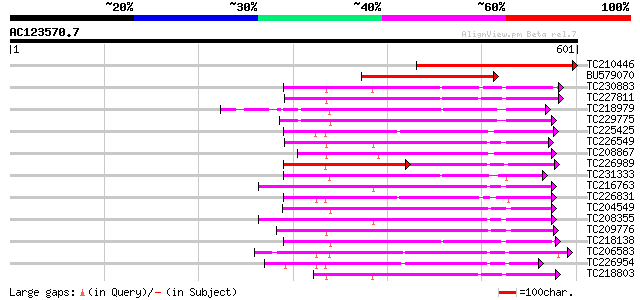
Score E
Sequences producing significant alignments: (bits) Value
TC210446 similar to UP|Q7EZR1 (Q7EZR1) Receptor protein kinase P... 231 9e-61
BU579070 similar to GP|11994399|db contains similarity to recept... 228 6e-60
TC230883 UP|Q8LKR3 (Q8LKR3) Receptor-like kinase RHG4, complete 216 2e-56
TC227811 similar to PIR|JQ1674|JQ1674 protein kinase TMK1 , rece... 204 9e-53
TC218979 similar to UP|Q9FUU9 (Q9FUU9) Leaf senescence-associate... 199 3e-51
TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable serin... 195 4e-50
TC225425 similar to GB|AAN31120.1|23506217|AY149966 At2g17220/T2... 193 2e-49
TC226549 UP|Q84P43 (Q84P43) Protein kinase Pti1, complete 192 4e-49
TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific... 191 1e-48
TC226989 similar to UP|PBS1_ARATH (Q9FE20) Serine/threonine-prot... 113 1e-48
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 190 1e-48
TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete 189 4e-48
TC226831 similar to PIR|T52285|T52285 serine/threonine-specific ... 188 5e-48
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 187 9e-48
TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete 187 2e-47
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 186 2e-47
TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis r... 185 4e-47
TC206583 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 182 3e-46
TC226954 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B ... 182 4e-46
TC218803 similar to UP|Q84P72 (Q84P72) Receptor-like protein kin... 181 1e-45
>TC210446 similar to UP|Q7EZR1 (Q7EZR1) Receptor protein kinase PERK1-like
protein, partial (29%)
Length = 741
Score = 231 bits (588), Expect = 9e-61
Identities = 118/171 (69%), Positives = 134/171 (78%), Gaps = 1/171 (0%)
Frame = +3
Query: 432 KNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYA-YGSVSSKIDVYAFGVVLYE 490
+NF KVADFGLTKL + G SS+PT + GTFGYMPPEYA YG VS K+DVYAFGVVLYE
Sbjct: 3 ENFRGKVADFGLTKLTEVGSSSLPTGRLVGTFGYMPPEYAQYGDVSPKVDVYAFGVVLYE 182
Query: 491 LISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQ 550
LISAK A++ DS AD KGLV LF+ V QP E L KLVDPRLGDNYPID V KMAQ
Sbjct: 183 LISAKEAIVKTNDSVADSKGLVALFDGVLSQPDLTEELCKLVDPRLGDNYPIDSVRKMAQ 362
Query: 551 LAKVCTNSDPQQRPNMSSVVVALTTLTSTTEDWDITSIFKNPNLVNLMSGR 601
LAK CT +PQ RP+M S+VVAL TL+STT+DWD+ S ++N NLVNLMSGR
Sbjct: 363 LAKACTQDNPQLRPSMRSIVVALMTLSSTTDDWDVGSFYENQNLVNLMSGR 515
>BU579070 similar to GP|11994399|db contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(19%)
Length = 437
Score = 228 bits (581), Expect = 6e-60
Identities = 115/145 (79%), Positives = 124/145 (85%)
Frame = +3
Query: 374 IDNGNLGQHLRSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKN 433
I+NGNLGQHLR S PL WS RV+IALDSARGL+YIHEHTVP YIHRDIKSENIL+DKN
Sbjct: 3 IENGNLGQHLRKSGFNPLPWSTRVQIALDSARGLQYIHEHTVPVYIHRDIKSENILIDKN 182
Query: 434 FCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAYGSVSSKIDVYAFGVVLYELIS 493
F AKVA FGLTKLID G SS+PTVNM GTFGYMPPEYAYG+VS KIDVYAFGVVLYELIS
Sbjct: 183 FGAKVAYFGLTKLIDVGSSSLPTVNMKGTFGYMPPEYAYGNVSPKIDVYAFGVVLYELIS 362
Query: 494 AKAAVIMGEDSGADLKGLVVLFEEV 518
K A+ G SGA+L GLV LF+EV
Sbjct: 363 GKEALSRGGVSGAELXGLVSLFDEV 437
>TC230883 UP|Q8LKR3 (Q8LKR3) Receptor-like kinase RHG4, complete
Length = 3053
Score = 216 bits (551), Expect = 2e-56
Identities = 137/310 (44%), Positives = 185/310 (59%), Gaps = 13/310 (4%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKKMDM-----KATKEFLAEL 344
FS + L TNNF+ N +G+GGFG VY +L+ G K A+K+M+ K KEF AE+
Sbjct: 1594 FSIQVLQQVTNNFSEENILGRGGFGVVYKGQLHDGTKIAVKRMESVAMGNKGLKEFEAEI 1773
Query: 345 KVLTRVHHVNLVRLIGYCVEG-SLFLVYEYIDNGNLGQHL---RSSDGEPLSWSIRVKIA 400
VL++V H +LV L+GYC+ G LVYEY+ G L QHL + PL+W RV IA
Sbjct: 1774 AVLSKVRHRHLVALLGYCINGIERLLVYEYMPQGTLTQHLFEWQEQGYVPLTWKQRVVIA 1953
Query: 401 LDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMA 460
LD ARG+EY+H ++IHRD+K NILL + AKVADFGL K G SV T +A
Sbjct: 1954 LDVARGVEYLHSLAQQSFIHRDLKPSNILLGDDMRAKVADFGLVKNAPDGKYSVET-RLA 2130
Query: 461 GTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKG-LVVLFEEV 518
GTFGY+ PEY A G V++K+D+YAFG+VL ELI+ + A+ +D+ D + LV F V
Sbjct: 2131 GTFGYLAPEYAATGRVTTKVDIYAFGIVLMELITGRKAL---DDTVPDERSHLVTWFRRV 2301
Query: 519 FDQPHPIEGLKKLVDPRLG-DNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLT 577
E + K +D L D ++ ++K+A+LA CT +P QRP+M V L L
Sbjct: 2302 LINK---ENIPKAIDQTLNPDEETMESIYKVAELAGHCTAREPYQRPDMGHAVNVLVPL- 2469
Query: 578 STTEDWDITS 587
E W +S
Sbjct: 2470 --VEQWKPSS 2493
>TC227811 similar to PIR|JQ1674|JQ1674 protein kinase TMK1 , receptor type
precursor - Arabidopsis thaliana {Arabidopsis thaliana;}
, partial (40%)
Length = 1436
Score = 204 bits (519), Expect = 9e-53
Identities = 128/308 (41%), Positives = 184/308 (59%), Gaps = 12/308 (3%)
Frame = +2
Query: 292 SYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKKMDM-----KATKEFLAELK 345
S + L N T+NF+ N +GQGGFG VY EL+ G + A+K+M+ K EF +E+
Sbjct: 77 SIQVLKNVTDNFSEKNVLGQGGFGTVYRGELHDGTRIAVKRMECGAIAGKGAAEFKSEIA 256
Query: 346 VLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRS--SDG-EPLSWSIRVKIAL 401
VLT+V H +LV L+GYC++G+ LVYEY+ G L +HL +G EPL W+ R+ IAL
Sbjct: 257 VLTKVRHRHLVSLLGYCLDGNEKLLVYEYMPQGTLSRHLFDWPEEGLEPLEWNRRLTIAL 436
Query: 402 DSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAG 461
D ARG+EY+H ++IHRD+K NILL + AKVADFGL +L G +S+ T +AG
Sbjct: 437 DVARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKASIET-RIAG 613
Query: 462 TFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFD 520
TFGY+ PEYA G V++K+DV++FGV+L ELI+ + A + E D LV F +
Sbjct: 614 TFGYLAPEYAVTGRVTTKVDVFSFGVILMELITGRKA--LDETQPEDSMHLVTWFRRMSI 787
Query: 521 QPHPIEGLKKLVDPRLGDN-YPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTST 579
+ +K +D + N + + +A+LA C +P QRP+M V L++L
Sbjct: 788 NK---DSFRKAIDSTIELNEETLASIHTVAELAGHCGAREPYQRPDMGHAVNVLSSLVEL 958
Query: 580 TEDWDITS 587
+ D S
Sbjct: 959 WKPSDQNS 982
>TC218979 similar to UP|Q9FUU9 (Q9FUU9) Leaf senescence-associated
receptor-like protein kinase, partial (34%)
Length = 1343
Score = 199 bits (506), Expect = 3e-51
Identities = 132/358 (36%), Positives = 203/358 (55%), Gaps = 8/358 (2%)
Frame = +3
Query: 224 LILLSFCIYVTYYRKKKIRKQEFLSEESSAIFGQVKNDEVSGNATYGTSDSASPANMIGI 283
++L++ I T R+K E+S+A+ E+S + DS +
Sbjct: 3 ILLVAVAILWTLKRRKS-------KEKSTALMEVNDESEISRLQSTKKDDSL-------L 140
Query: 284 RVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKAT---KEF 340
+V+K +SY ++ TNNFN IG+GGFG VY ++ A+K + + ++F
Sbjct: 141 QVKKQ-IYSYSDVLKITNNFNTI--IGKGGFGTVYLGYIDDSPVAVKVLSPSSVNGFQQF 311
Query: 341 LAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL--RSSDGEPLSWSIRV 397
AE+K+L +VHH NL LIGYC EG+ L+YEY+ NGNL +HL + S LSW R+
Sbjct: 312 QAEVKLLVKVHHKNLTSLIGYCNEGTNKALIYEYMANGNLQEHLSGKHSKSTFLSWEDRL 491
Query: 398 KIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDA-GISSVPT 456
+IA+D+A GLEY+ P IHRD+KS NILL+++F AK++DFGL+K I G S V T
Sbjct: 492 RIAVDAALGLEYLQNGCKPPIIHRDVKSTNILLNEHFQAKLSDFGLSKAIPTDGESHVST 671
Query: 457 VNMAGTFGYMPPEYAYGS-VSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLF 515
V +AGT GY+ P S ++ K DVY+FGVVL E+I+ + + ++ G + + L
Sbjct: 672 V-LAGTPGYLDPHCHISSRLTQKSDVYSFGVVLLEIITNQPVMERNQEKGHIRERVRSLI 848
Query: 516 EEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVAL 573
E+ ++ +VD RL +Y I+ +K ++A C + +P +RP MS + + L
Sbjct: 849 EK--------GDIRAIVDSRLEGDYDINSAWKALEIAMACVSQNPNERPIMSEIAIEL 998
>TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable
serine/threonine-protein kinase pelle , partial (8%)
Length = 1250
Score = 195 bits (496), Expect = 4e-50
Identities = 111/300 (37%), Positives = 178/300 (59%), Gaps = 7/300 (2%)
Frame = +1
Query: 287 KSGEFSYEELANATNNFNMANKIGQGGFGEVYYAELNGEKAAIKKMDMKATK---EFLAE 343
K ++++ EL TNNF +G+GGFG+VY+ ++ + A+K + A + +FLAE
Sbjct: 37 KQRQYTFNELVKITNNFTRI--LGRGGFGKVYHGFIDDTQVAVKMLSPSAVRGYEQFLAE 210
Query: 344 LKVLTRVHHVNLVRLIGYC-VEGSLFLVYEYIDNGNLGQHL--RSSDGEPLSWSIRVKIA 400
+K+L RVHH NL L+GYC E ++ L+YEY+ NGNL + + +SS + L+W R++IA
Sbjct: 211 VKLLMRVHHRNLTSLVGYCNEENNIGLIYEYMANGNLDEIVSGKSSRAKFLTWEDRLQIA 390
Query: 401 LDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMA 460
LD+A+GLEY+H P IHRD+K NILL++NF AK+ADFGL+K S + +A
Sbjct: 391 LDAAQGLEYLHNGCKPPIIHRDVKCANILLNENFQAKLADFGLSKSFPTDGGSYMSTVVA 570
Query: 461 GTFGYMPPEYAYGS-VSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVF 519
GT GY+ PEY+ S ++ K DVY+FGVVL E+++ + A+ D + + +
Sbjct: 571 GTPGYLDPEYSISSRLTEKSDVYSFGVVLLEMVTGQPAIAKTPDKTHISQWVKSMLSN-- 744
Query: 520 DQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTST 579
+K + D R +++ V+++ ++ + P +RP+MS +V L +T
Sbjct: 745 ------GDIKNIADSRFKEDFDTSSVWRIVEIGMASVSISPFKRPSMSYIVNELKECLTT 906
>TC225425 similar to GB|AAN31120.1|23506217|AY149966 At2g17220/T23A1.8
{Arabidopsis thaliana;} , partial (80%)
Length = 1386
Score = 193 bits (491), Expect = 2e-49
Identities = 121/307 (39%), Positives = 175/307 (56%), Gaps = 16/307 (5%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL---------NGEKAAIKKMD---MKATK 338
F++ EL AT NF +G+GGFG+V+ L +G A+KK++ ++ +
Sbjct: 112 FTFAELKAATRNFKADTVLGEGGFGKVFKGWLEEKATSKGGSGTVIAVKKLNSESLQGLE 291
Query: 339 EFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHL--RSSDGEPLSWSI 395
E+ +E+ L R+ H NLV+L+GYC+E S L LVYE++ G+L HL R S +PL W I
Sbjct: 292 EWQSEVNFLGRLSHTNLVKLLGYCLEESELLLVYEFMQKGSLENHLFGRGSAVQPLPWDI 471
Query: 396 RVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVP 455
R+KIA+ +ARGL ++H T I+RD K+ NILLD ++ AK++DFGL KL + S
Sbjct: 472 RLKIAIGAARGLAFLH--TSEKVIYRDFKASNILLDGSYNAKISDFGLAKLGPSASQSHV 645
Query: 456 TVNMAGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVL 514
T + GT GY PEY A G + K DVY FGVVL E+++ A+ SG L
Sbjct: 646 TTRVMGTHGYAAPEYVATGHLYVKSDVYGFGVVLVEILTGLRALDSNRPSGQH-----KL 810
Query: 515 FEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALT 574
E V H LK ++D RL +P F++AQL+ C S+P+ RP+M V+ L
Sbjct: 811 TEWVKPYLHDRRKLKGIMDSRLEGKFPSKAAFRIAQLSMKCLASEPKHRPSMKDVLENLE 990
Query: 575 TLTSTTE 581
+ + E
Sbjct: 991 RIQAANE 1011
>TC226549 UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1627
Score = 192 bits (488), Expect = 4e-49
Identities = 119/299 (39%), Positives = 172/299 (56%), Gaps = 14/299 (4%)
Frame = +3
Query: 292 SYEELANATNNFNMANKIGQGGFGEVYYAELNGEKA-AIKKMDM----KATKEFLAELKV 346
S +EL T+NF IG+G +G VYYA LN KA A+KK+D+ ++ EFL ++ +
Sbjct: 375 SLDELKEKTDNFGSKALIGEGSYGRVYYATLNNGKAVAVKKLDVSSEPESNNEFLTQVSM 554
Query: 347 LTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGNLGQHLR-------SSDGEPLSWSIRVK 398
++R+ + N V L GYCVEG+L L YE+ G+L L + G L W RV+
Sbjct: 555 VSRLKNGNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVR 734
Query: 399 IALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVN 458
IA+D+ARGLEY+HE P IHRDI+S N+L+ +++ AK+ADF L+ + + +
Sbjct: 735 IAVDAARGLEYLHEKVQPPIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTR 914
Query: 459 MAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEE 517
+ GTFGY PEYA G ++ K DVY+FGVVL EL++ + V G + LV
Sbjct: 915 VLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQ--QSLVTWATP 1088
Query: 518 VFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTL 576
+ + +K+ VDP+L YP V K+ +A +C + + RPNMS VV AL L
Sbjct: 1089RLSE----DKVKQCVDPKLKGEYPPKGVAKLGAVAALCVQYEAEFRPNMSIVVKALQPL 1253
>TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific protein
kinase-like protein, partial (72%)
Length = 1118
Score = 191 bits (484), Expect = 1e-48
Identities = 117/285 (41%), Positives = 161/285 (56%), Gaps = 11/285 (3%)
Frame = +1
Query: 306 ANKIGQGGFGEVYYAELN-GEKAAIKKMDM---KATKEFLAELKVLTRVHHVNLVRLIGY 361
+N IG GGFG VY LN G K AIK MD + +EF E+++L+R+H L+ L+GY
Sbjct: 7 SNVIGHGGFGLVYRGVLNDGRKVAIKFMDQAGKQGEEEFKVEVELLSRLHSPYLLALLGY 186
Query: 362 CVEGS-LFLVYEYIDNGNLGQHLRSSDGE-----PLSWSIRVKIALDSARGLEYIHEHTV 415
C + + LVYE++ NG L +HL L W R++IAL++A+GLEY+HEH
Sbjct: 187 CSDSNHKLLVYEFMANGGLQEHLYPVSNSIITPVKLDWETRLRIALEAAKGLEYLHEHVS 366
Query: 416 PTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GS 474
P IHRD KS NILLDK F AKV+DFGL KL + + GT GY+ PEYA G
Sbjct: 367 PPVIHRDFKSSNILLDKKFHAKVSDFGLAKLGPDRAGGHVSTRVLGTQGYVAPEYALTGH 546
Query: 475 VSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDP 534
+++K DVY++GVVL EL++ + V M G VL E + K++DP
Sbjct: 547 LTTKSDVYSYGVVLLELLTGRVPVDMKRPPGEG-----VLVSWALPLLTDREKVVKIMDP 711
Query: 535 RLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTST 579
L Y + V ++A +A +C + RP M+ VV +L L T
Sbjct: 712 SLEGQYSMKEVVQVAAIAAMCVQPEADYRPLMADVVQSLVPLVKT 846
>TC226989 similar to UP|PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase
PBS1 (AvrPphB susceptible protein 1) , partial (86%)
Length = 1730
Score = 113 bits (282), Expect(2) = 1e-48
Identities = 60/142 (42%), Positives = 90/142 (63%), Gaps = 8/142 (5%)
Frame = +2
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELN--GEKAAIKKMD---MKATKEFLAELK 345
F++ ELA AT NF + +G+GGFG VY L + A+K++D ++ +EFL E+
Sbjct: 245 FTFRELAAATKNFRPESFVGEGGFGRVYKGRLETTAQIVAVKQLDKNGLQGNREFLVEVL 424
Query: 346 VLTRVHHVNLVRLIGYCVEG-SLFLVYEYIDNGNLGQHLRS--SDGEPLSWSIRVKIALD 402
+L+ +HH NLV LIGYC +G LVYE++ G+L HL D EPL W+ R+KIA+
Sbjct: 425 MLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEPLDWNTRMKIAVG 604
Query: 403 SARGLEYIHEHTVPTYIHRDIK 424
+A+GLEY+H+ P I + ++
Sbjct: 605 AAKGLEYLHDKANPPGIQQRLQ 670
Score = 99.4 bits (246), Expect(2) = 1e-48
Identities = 60/164 (36%), Positives = 88/164 (53%), Gaps = 1/164 (0%)
Frame = +3
Query: 420 HRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSK 478
+RD KS NILLD+ + K++DFGL KL G S + + GT+GY PEYA G ++ K
Sbjct: 657 NRDCKSSNILLDEGYHPKLSDFGLAKLGPVGDKSHVSTRVMGTYGYCAPEYAMTGQLTVK 836
Query: 479 IDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGD 538
DVY+FGVV ELI+ + A+ + G + LV +F+ KL DPRL
Sbjct: 837 SDVYSFGVVFLELITGRKAIDSTQPQGE--QNLVTWARPLFNDRRK---FSKLADPRLQG 1001
Query: 539 NYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTED 582
+P+ +++ +A +C RP + VV AL+ L + D
Sbjct: 1002RFPMRGLYQALAVASMCIQESAATRPLIGDVVTALSYLANQAYD 1133
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 190 bits (483), Expect = 1e-48
Identities = 115/290 (39%), Positives = 166/290 (56%), Gaps = 10/290 (3%)
Frame = +3
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDMKAT---KEFLAELKV 346
F++ EL AT F+ AN + +GGFG V+ L +G+ A+K+ + +T KEF +E++V
Sbjct: 6 FTFSELQLATGGFSQANFLAEGGFGSVHRGVLPDGQVIAVKQYKLASTQGDKEFCSEVEV 185
Query: 347 LTRVHHVNLVRLIGYCVE-GSLFLVYEYIDNGNLGQHLRSSDGEPLSWSIRVKIALDSAR 405
L+ H N+V LIG+CV+ G LVYEYI NG+L H+ L WS R KIA+ +AR
Sbjct: 186 LSCAQHRNVVMLIGFCVDDGRRLLVYEYICNGSLDSHIYRRKQNVLEWSARQKIAVGAAR 365
Query: 406 GLEYIHEH-TVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFG 464
GL Y+HE V +HRD++ NILL +F A V DFGL + G V T + GTFG
Sbjct: 366 GLRYLHEECRVGCIVHRDMRPNNILLTHDFEALVGDFGLARWQPDGDMGVET-RVIGTFG 542
Query: 465 YMPPEYAY-GSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPH 523
Y+ PEYA G ++ K DVY+FG+VL EL++ + AV + G + +
Sbjct: 543 YLAPEYAQSGQITEKADVYSFGIVLLELVTGRKAVDINRPKGQQC---------LSEWAR 695
Query: 524 PI---EGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVV 570
P+ + KL+DP L + Y V++M + + +C DP RP MS V+
Sbjct: 696 PLLEKQATYKLIDPSLRNCYVDQEVYRMLKCSSLCIGRDPHLRPRMSQVL 845
>TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete
Length = 1634
Score = 189 bits (479), Expect = 4e-48
Identities = 125/329 (37%), Positives = 180/329 (53%), Gaps = 13/329 (3%)
Frame = +1
Query: 264 SGNATY-GTSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL 322
+GN +Y G + + I + + +EL + T+NF IG+G +G+VY A L
Sbjct: 274 TGNPSYHGRHTAVTAPRTINFQPIAVPSITVDELKSLTDNFGSKYFIGEGAYGKVYQATL 453
Query: 323 -NGEKAAIKKMDM--KATKEFLAELKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGN 378
NG IKK+D + +EFL+++ +++R+ H N+V L+ YCV+G L YEY G+
Sbjct: 454 KNGHAVVIKKLDSSNQPEQEFLSQVSIVSRLKHENVVELVNYCVDGPFRALAYEYAPKGS 633
Query: 379 LGQHLR-------SSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLD 431
L L + G LSW+ RVKIA+ +ARGLEY+HE IHR IKS NILL
Sbjct: 634 LHDILHGRKGVKGAQPGPVLSWAQRVKIAVGAARGLEYLHEKAEIHIIHRYIKSSNILLF 813
Query: 432 KNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYE 490
+ AKVADF L+ + + + + GTFGY PEYA G ++SK DVY+FGV+L E
Sbjct: 814 DDDVAKVADFDLSNQAPDAAARLHSTRVLGTFGYHAPEYAMTGQLTSKSDVYSFGVILLE 993
Query: 491 LISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQ 550
L++ + V G + LV + + +K+ VD RL YP V KMA
Sbjct: 994 LLTGRKPVDHTLPRGQ--QSLVTWATPKLSE----DKVKQCVDVRLKGEYPSKSVAKMAA 1155
Query: 551 LAKVCTNSDPQQRPNMSSVVVALTTLTST 579
+A +C + + RPNMS +V AL L +T
Sbjct: 1156VAALCVQYEAEFRPNMSIIVKALQPLLNT 1242
>TC226831 similar to PIR|T52285|T52285 serine/threonine-specific protein
kinase APK2a [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (82%)
Length = 1693
Score = 188 bits (478), Expect = 5e-48
Identities = 116/309 (37%), Positives = 178/309 (57%), Gaps = 20/309 (6%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAELN-----------GEKAAIKKMD---MKA 336
F++ EL NAT NF + +G+GGFG VY ++ G A+KK+ ++
Sbjct: 439 FTFNELKNATRNFRPDSLLGEGGFGYVYKGWIDEHTFTASKPGSGMVVAVKKLKPEGLQG 618
Query: 337 TKEFLAELKVLTRVHHVNLVRLIGYCVEG-SLFLVYEYIDNGNLGQHLRSSDGEPLSWSI 395
KE+L E+ L ++HH NLV+LIGYC +G + LVYE++ G+L HL +PLSWS+
Sbjct: 619 HKEWLTEVDYLGQLHHQNLVKLIGYCADGENRLLVYEFMSKGSLENHLFRRGPQPLSWSV 798
Query: 396 RVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVP 455
R+K+A+ +ARGL ++H + I+RD K+ NILLD F AK++DFGL K G +
Sbjct: 799 RMKVAIGAARGLSFLH-NAKSQVIYRDFKASNILLDAEFNAKLSDFGLAKAGPTGDRTHV 975
Query: 456 TVNMAGTFGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVL 514
+ + GT GY PEY A G +++K DVY+FGVVL EL+S + AV + S A ++
Sbjct: 976 STQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLELLSGRRAV---DRSKAGVE----- 1131
Query: 515 FEEVFDQPHPIEG----LKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVV 570
+ + + P G L +++D +LG YP + A LA C N + + RP ++ V+
Sbjct: 1132-QNLVEWAKPYLGDKRRLFRIMDTKLGGQYPQKGAYMAATLALKCLNREAKGRPPITEVL 1308
Query: 571 VALTTLTST 579
L + ++
Sbjct: 1309QTLEQIAAS 1335
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 187 bits (476), Expect = 9e-48
Identities = 109/297 (36%), Positives = 173/297 (57%), Gaps = 7/297 (2%)
Frame = +3
Query: 290 EFSYEELANATNNFNMANKIGQGGFGEVYYAELN-GEKAAIKKMDMKATK---EFLAELK 345
+F + + ATN F+ NK+G+GGFGEVY L+ G+ A+K++ + + EF E+
Sbjct: 789 QFDFSTIEAATNKFSADNKLGEGGFGEVYKGTLSSGQVVAVKRLSKSSGQGGEEFKNEVV 968
Query: 346 VLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGE-PLSWSIRVKIALDS 403
V+ ++ H NLVRL+G+C++G LVYEY+ N +L L + + L W R KI
Sbjct: 969 VVAKLQHRNLVRLLGFCLQGEEKILVYEYVPNKSLDYILFDPEKQRELDWGRRYKIIGGI 1148
Query: 404 ARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTF 463
ARG++Y+HE + IHRD+K+ NILLD + K++DFG+ ++ + T + GT+
Sbjct: 1149 ARGIQYLHEDSRLRIIHRDLKASNILLDGDMNPKISDFGMARIFGVDQTQGNTSRIVGTY 1328
Query: 464 GYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQP 522
GYM PEYA +G S K DVY+FGV+L E++S K + GA+ L+ +++
Sbjct: 1329 GYMAPEYAMHGEFSVKSDVYSFGVLLMEILSGKKNSSFYQTDGAE--DLLSYAWQLWKDG 1502
Query: 523 HPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTST 579
P+E L+DP L ++Y + V + + +C DP RP M+++V+ L + T T
Sbjct: 1503 TPLE----LMDPILRESYNQNEVIRSIHIGLLCVQEDPADRPTMATIVLMLDSNTVT 1661
>TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete
Length = 1536
Score = 187 bits (474), Expect = 2e-47
Identities = 124/329 (37%), Positives = 179/329 (53%), Gaps = 13/329 (3%)
Frame = +2
Query: 264 SGNATY-GTSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL 322
+GN +Y G + + I ++ + +EL T+NF IG+G +G+VY A L
Sbjct: 284 TGNPSYHGRHAAVTAPRTINVQPIAVPSITVDELKPLTDNFGSKCFIGEGAYGKVYQATL 463
Query: 323 -NGEKAAIKKMDM--KATKEFLAELKVLTRVHHVNLVRLIGYCVEGSL-FLVYEYIDNGN 378
NG IKK+D + EFL+++ +++R+ H N+V L+ YCV+G L YEY G+
Sbjct: 464 KNGRAVVIKKLDSSNQPEHEFLSQVSIVSRLKHENVVELVNYCVDGPFRALAYEYAPKGS 643
Query: 379 LGQHLR-------SSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLD 431
L L + G LSW+ RVKIA+ +ARGLEY+HE IHR IKS NILL
Sbjct: 644 LHDILHGRKGVKGAQPGPVLSWAQRVKIAVGAARGLEYLHEKAEIHIIHRYIKSSNILLF 823
Query: 432 KNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYE 490
+ AK+ADF L+ + + + + GTFGY PEYA G ++SK DVY+FGV+L E
Sbjct: 824 DDDVAKIADFDLSNQAPDAAARLHSTRVLGTFGYHAPEYAMTGQLTSKSDVYSFGVILLE 1003
Query: 491 LISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQ 550
L++ + V G + LV + + +K+ VD RL YP V KMA
Sbjct: 1004LLTGRKPVDHTLPRGQ--QSLVTWATPKLSE----DKVKQCVDVRLKGEYPSKSVAKMAA 1165
Query: 551 LAKVCTNSDPQQRPNMSSVVVALTTLTST 579
+A +C + + RPNMS +V AL L +T
Sbjct: 1166VAALCVQYEAEFRPNMSIIVKALQPLLNT 1252
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 186 bits (473), Expect = 2e-47
Identities = 109/306 (35%), Positives = 178/306 (57%), Gaps = 7/306 (2%)
Frame = +2
Query: 283 IRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDM---KATK 338
I +S F + + AT F+ ANK+G+GGFGEVY L +G++ A+K++ + +
Sbjct: 20 ISAVESLRFDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQGGE 199
Query: 339 EFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDGEP-LSWSIR 396
EF E++++ ++ H NLVRL+G+C+EG LVYE++ N +L L + + L W+ R
Sbjct: 200 EFKNEVEIVAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTRR 379
Query: 397 VKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPT 456
KI ARG++Y+HE + IHRD+K+ N+LLD + K++DFG+ ++ + T
Sbjct: 380 YKIVEGIARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANT 559
Query: 457 VNMAGTFGYMPPEYA-YGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLF 515
+ GT+GYM PEYA +G S+K DVY+FGV++ E+IS K E A+ L+
Sbjct: 560 NRIVGTYGYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAE--DLLSYA 733
Query: 516 EEVFDQPHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTT 575
+++ P+E L+D L ++Y + V + + +C DP RP M+SVV+ L +
Sbjct: 734 WKLWKDEAPLE----LMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLMLDS 901
Query: 576 LTSTTE 581
+ T +
Sbjct: 902 YSVTLQ 919
>TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis receptor
kinase 1, partial (60%)
Length = 1444
Score = 185 bits (470), Expect = 4e-47
Identities = 118/303 (38%), Positives = 173/303 (56%), Gaps = 9/303 (2%)
Frame = +1
Query: 291 FSYEELANATNNFNMANKIGQGGFGEVYYAEL-NGEKAAIKKMDMKATK----EFLAELK 345
FS EL AT++F+ N +G+GGFG+VY L +G A+K++ + T +F E++
Sbjct: 121 FSLRELQVATDSFSNKNILGRGGFGKVYKGRLADGSLVAVKRLKEERTPGGELQFQTEVE 300
Query: 346 VLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDNGNLGQHLRSSDG--EPLSWSIRVKIALD 402
+++ H NL+RL G+C+ + LVY Y+ NG++ LR EPL W R ++AL
Sbjct: 301 MISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPPYQEPLDWPTRKRVALG 480
Query: 403 SARGLEYIHEHTVPTYIHRDIKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGT 462
SARGL Y+H+H P IHRD+K+ NILLD+ F A V DFGL KL+D + V T + GT
Sbjct: 481 SARGLSYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTA-VRGT 657
Query: 463 FGYMPPEY-AYGSVSSKIDVYAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQ 521
G++ PEY + G S K DV+ +G++L ELI+ + A + + D L+ + + +
Sbjct: 658 IGHIAPEYLSTGKSSEKTDVFGYGIMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKE 837
Query: 522 PHPIEGLKKLVDPRLGDNYPIDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTTE 581
+ L+ LVDP L NY V ++ Q+A +CT P RP MS VV L E
Sbjct: 838 ----KKLEMLVDPDLQTNYIETEVEQLIQVALLCTQGSPMDRPKMSEVVRMLEG-DGLAE 1002
Query: 582 DWD 584
WD
Sbjct: 1003RWD 1011
>TC206583 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(84%)
Length = 1629
Score = 182 bits (463), Expect = 3e-46
Identities = 128/364 (35%), Positives = 189/364 (51%), Gaps = 27/364 (7%)
Frame = +1
Query: 260 NDEVSGNATYGTSDSASPANMIGIRVEKSGEFSYEELANATNNFNMANKIGQGGFGEVYY 319
ND+VS N+ T S ++ F+ EL AT NF + +G+GGFG V+
Sbjct: 163 NDKVSANSIPQTPRSEGEI----LQSSNLKSFTLSELKTATRNFRPDSVLGEGGFGSVFK 330
Query: 320 AELN-----------GEKAAIKKMDMKAT---KEFLAELKVLTRVHHVNLVRLIGYCVEG 365
++ G A+K+++ +E+LAE+ L + H +LVRLIG+C+E
Sbjct: 331 GWIDENSLTATKPGTGIVIAVKRLNQDGIPGHREWLAEVNYLGQHFHSHLVRLIGFCLED 510
Query: 366 S-LFLVYEYIDNGNLGQHL--RSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRD 422
LVYE++ G+ HL R S +PLSWS+R+K+ALD+A+GL ++H I+RD
Sbjct: 511 EHRLLVYEFMPRGSWENHLFRRGSYFQPLSWSLRLKVALDAAKGLAFLHSAEAKV-IYRD 687
Query: 423 IKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEY-AYGSVSSKIDV 481
K+ N+LLD + AK++DFGL K G S + + GT+GY PEY A G +++K DV
Sbjct: 688 FKTSNVLLDSKYNAKLSDFGLAKDGPTGDKSHVSTRVMGTYGYAAPEYLATGHLTAKSDV 867
Query: 482 YAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYP 541
Y+FGVVL E++S K AV SG LV + I +++D RL Y
Sbjct: 868 YSFGVVLLEMLSGKRAVDKNRPSGQ--HNLVEWAKPFMANKRKI---FRVLDTRLQGQYS 1032
Query: 542 IDHVFKMAQLAKVCTNSDPQQRPNMSSVVVALTTLTSTT---------EDWDITSIFKNP 592
D +K+A LA C + + + RPNM VV L L + D+ +NP
Sbjct: 1033TDDAYKLATLALRCLSIESKFRPNMDQVVTTLEQLQLSNVNGGPRVRRRSADVNRGHQNP 1212
Query: 593 NLVN 596
+ VN
Sbjct: 1213SSVN 1224
>TC226954 similar to UP|APKB_ARATH (P46573) Protein kinase APK1B , partial
(80%)
Length = 1175
Score = 182 bits (462), Expect = 4e-46
Identities = 122/325 (37%), Positives = 181/325 (55%), Gaps = 29/325 (8%)
Frame = +2
Query: 271 TSDSASPANMIGIRVEKSGE---------FSYEELANATNNFNMANKIGQGGFGEVYYAE 321
+S+S S + I + GE FSY EL AT NF + +G+GGFG V+
Sbjct: 218 SSNSRSSSASIPVTSRSEGEILQSSNLKSFSYHELRAATRNFRPDSVLGEGGFGSVFKGW 397
Query: 322 LN-----------GEKAAIKKMD---MKATKEFLAELKVLTRVHHVNLVRLIGYCVEGS- 366
++ G+ A+KK++ ++ +E+LAE+ L ++ H NLV+LIGYC E
Sbjct: 398 IDEHSLAATKPGIGKIVAVKKLNQDGLQGHREWLAEINYLGQLQHPNLVKLIGYCFEDEH 577
Query: 367 LFLVYEYIDNGNLGQHL--RSSDGEPLSWSIRVKIALDSARGLEYIH--EHTVPTYIHRD 422
LVYE++ G++ HL R S +P SWS+R+KIAL +A+GL ++H EH V I+RD
Sbjct: 578 RLLVYEFMPKGSMENHLFRRGSYFQPFSWSLRMKIALGAAKGLAFLHSTEHKV---IYRD 748
Query: 423 IKSENILLDKNFCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEY-AYGSVSSKIDV 481
K+ NILLD N+ AK++DFGL + G S + + GT GY PEY A G +++K DV
Sbjct: 749 FKTSNILLDTNYNAKLSDFGLARDGPTGDKSHVSTRVMGTRGYAAPEYLATGHLTTKSDV 928
Query: 482 YAFGVVLYELISAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLGDNYP 541
Y+FGVVL E+IS + A+ + +G LV + + +++DPRL Y
Sbjct: 929 YSFGVVLLEMISGRRAIDKNQPTGE--HNLVEWAKPYLSNKRRV---FRVMDPRLEGQYS 1093
Query: 542 IDHVFKMAQLAKVCTNSDPQQRPNM 566
+ A LA C + +P+ RPNM
Sbjct: 1094QNRAQAAAALAMQCFSVEPKCRPNM 1168
>TC218803 similar to UP|Q84P72 (Q84P72) Receptor-like protein kinase-like
protein (Fragment), partial (35%)
Length = 1266
Score = 181 bits (458), Expect = 1e-45
Identities = 113/273 (41%), Positives = 158/273 (57%), Gaps = 11/273 (4%)
Frame = +1
Query: 323 NGEKAAIKKMDM-----KATKEFLAELKVLTRVHHVNLVRLIGYCVEGS-LFLVYEYIDN 376
+G K A+K+M+ KA EF +E+ VL++V H +LV L+GY EG+ LVYEY+
Sbjct: 25 DGTKIAVKRMEAGVISSKALDEFQSEIAVLSKVRHRHLVSLLGYSTEGNERILVYEYMPQ 204
Query: 377 GNLGQHL---RSSDGEPLSWSIRVKIALDSARGLEYIHEHTVPTYIHRDIKSENILLDKN 433
G L +HL +S D EPLSW R+ IALD ARG+EY+H ++IHRD+K NILL +
Sbjct: 205 GALSKHLFHWKSHDLEPLSWKRRLNIALDVARGMEYLHTLAHQSFIHRDLKPSNILLADD 384
Query: 434 FCAKVADFGLTKLIDAGISSVPTVNMAGTFGYMPPEYAY-GSVSSKIDVYAFGVVLYELI 492
F AKV+DFGL KL G + +AGTFGY+ PEYA G +++K DV++FGVVL EL+
Sbjct: 385 FKAKVSDFGLVKLAPEGEKASVVTRLAGTFGYLAPEYAVTGKITTKADVFSFGVVLMELL 564
Query: 493 SAKAAVIMGEDSGADLKGLVVLFEEVFDQPHPIEGLKKLVDPRLG-DNYPIDHVFKMAQL 551
+ A + ED + + L F + + L +DP L + V +A+L
Sbjct: 565 TGLMA--LDEDRPEESQYLAAWFWHIKSDK---KKLMAAIDPALDVKEETFESVSIIAEL 729
Query: 552 AKVCTNSDPQQRPNMSSVVVALTTLTSTTEDWD 584
A CT +P QRP+M V L L + +D
Sbjct: 730 AGHCTAREPSQRPDMGHAVNVLAPLVEKWKPFD 828
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,516,535
Number of Sequences: 63676
Number of extensions: 324101
Number of successful extensions: 3710
Number of sequences better than 10.0: 992
Number of HSP's better than 10.0 without gapping: 2681
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2711
length of query: 601
length of database: 12,639,632
effective HSP length: 103
effective length of query: 498
effective length of database: 6,081,004
effective search space: 3028339992
effective search space used: 3028339992
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC123570.7


