

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122729.6 + phase: 0 /pseudo
(247 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
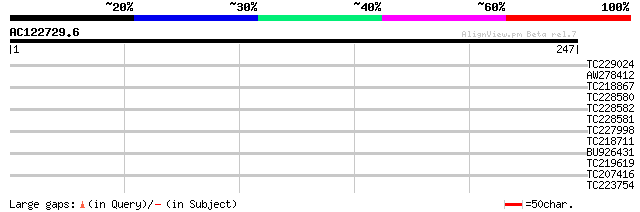
Score E
Sequences producing significant alignments: (bits) Value
TC229024 weakly similar to UP|GRX4_YEAST (P32642) Monothiol glut... 30 1.0
AW278412 30 1.0
TC218867 homologue to UP|Q43444 (Q43444) Phosphoinositide-specif... 29 1.7
TC228580 weakly similar to UP|Q944E5 (Q944E5) Cellulose synthase... 29 1.7
TC228582 similar to UP|Q944E5 (Q944E5) Cellulose synthase-like p... 29 2.3
TC228581 similar to UP|Q944E5 (Q944E5) Cellulose synthase-like p... 28 3.9
TC227998 weakly similar to UP|Q9LT99 (Q9LT99) FRO1-like protein;... 28 3.9
TC218711 similar to UP|Q90Z60 (Q90Z60) Rev-Erb beta (Fragment), ... 28 5.0
BU926431 28 5.0
TC219619 UP|NEU2_CATCO (P15211) Isoticin-neurophysin IT 2 precur... 27 6.6
TC207416 similar to UP|O49349 (O49349) Transcription factor IIA ... 27 8.6
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 27 8.6
>TC229024 weakly similar to UP|GRX4_YEAST (P32642) Monothiol glutaredoxin 4,
partial (30%)
Length = 1677
Score = 30.0 bits (66), Expect = 1.0
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = +1
Query: 58 GADICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHN 93
G DI + G + ++ ++G+D +EA EKE N
Sbjct: 742 GCDIAIAMHDSGELKEVFKDHGIDTTNEAKEKESGN 849
>AW278412
Length = 332
Score = 30.0 bits (66), Expect = 1.0
Identities = 19/56 (33%), Positives = 29/56 (50%)
Frame = +2
Query: 54 EEYVGADICEVFDEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEY 109
+EY + EV D+ YD++G E E+EEH E + D+E E+GE+
Sbjct: 191 DEYDNMEYEEVEDD-------YDDDG----EEEDEEEEHGEEVTDDEDGTGEHGEW 325
>TC218867 homologue to UP|Q43444 (Q43444) Phosphoinositide-specific
phospholipase C P25 , complete
Length = 2341
Score = 29.3 bits (64), Expect = 1.7
Identities = 21/69 (30%), Positives = 33/69 (47%), Gaps = 3/69 (4%)
Frame = +3
Query: 34 EREDIYESFEDETLEKPIYDEEYVGADICEVFDEEGNIDP---IYDENGLDDIHEAFEKE 90
E +++ E E+ EKP DEE G ++ + G I I D++ LDD + E E
Sbjct: 789 EAKEVQEKEEESQQEKPADDEEAWGKEVPSL--RGGTISDYKNIEDDDVLDDEEDIDEAE 962
Query: 91 EHNEPIYDE 99
+ + DE
Sbjct: 963 KSRQDAADE 989
>TC228580 weakly similar to UP|Q944E5 (Q944E5) Cellulose synthase-like
protein OsCslA9, partial (14%)
Length = 394
Score = 29.3 bits (64), Expect = 1.7
Identities = 15/48 (31%), Positives = 24/48 (49%)
Frame = -3
Query: 70 NIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKRFQ 117
NI +DE G + +KEEH E ++ + ++A E +E FQ
Sbjct: 203 NISKCHDEEG-----QRLDKEEHEELVFPKHNIIAAXKEEVECXXXFQ 75
>TC228582 similar to UP|Q944E5 (Q944E5) Cellulose synthase-like protein
OsCslA9, partial (19%)
Length = 835
Score = 28.9 bits (63), Expect = 2.3
Identities = 16/55 (29%), Positives = 26/55 (47%)
Frame = -1
Query: 70 NIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKRFQTSTNKDM 124
NI +DE G + +KEE E ++ E ++A E +E +FQ D+
Sbjct: 415 NISKCHDEEG-----KRLDKEEDEEMVFPEHNIIAATKEEVECDTKFQQVNPNDI 266
>TC228581 similar to UP|Q944E5 (Q944E5) Cellulose synthase-like protein
OsCslA9, partial (22%)
Length = 912
Score = 28.1 bits (61), Expect = 3.9
Identities = 15/48 (31%), Positives = 24/48 (49%)
Frame = -1
Query: 70 NIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKRFQ 117
NI +DE G + +KEE E ++ E ++A E +E +FQ
Sbjct: 459 NISKCHDEEG-----KRLDKEEDEEMVFPEHNIIAATKEEVECDTKFQ 331
>TC227998 weakly similar to UP|Q9LT99 (Q9LT99) FRO1-like protein; NADPH
oxidase-like, partial (9%)
Length = 792
Score = 28.1 bits (61), Expect = 3.9
Identities = 11/34 (32%), Positives = 20/34 (58%)
Frame = +1
Query: 214 YMMVMPTSTLL*NMVLKSSWHVYHLMNSLKEKRN 247
Y++ M S ++ + + WH++ NSLK+K N
Sbjct: 190 YVICMVASVVIFGGSVVAMWHIWEKQNSLKDKSN 291
>TC218711 similar to UP|Q90Z60 (Q90Z60) Rev-Erb beta (Fragment), partial (6%)
Length = 1204
Score = 27.7 bits (60), Expect = 5.0
Identities = 14/47 (29%), Positives = 21/47 (43%), Gaps = 5/47 (10%)
Frame = -3
Query: 1 CVMCQGYGHIAIDCVNY-----KAVTIVNREINNIFEEEREDIYESF 42
C++C G+GH + +C V VN IN E R ++ F
Sbjct: 365 CLLCYGHGHFSYNCPQNGHGIDPKVLAVNGAINRTAVEARRELLPRF 225
>BU926431
Length = 457
Score = 27.7 bits (60), Expect = 5.0
Identities = 19/62 (30%), Positives = 28/62 (44%), Gaps = 1/62 (1%)
Frame = +3
Query: 40 ESFEDETLEKPIYDEEYVGADICEVFDEEGN-IDPIYDENGLDDIHEAFEKEEHNEPIYD 98
E EDE + D+ G D + D+E + +D +DE D+ E E+ E D
Sbjct: 198 EQEEDENKSNEVEDQRGGGDDEIDENDQEQSAVDTDHDEEFTDEEKEEESDEKEKENSRD 377
Query: 99 EE 100
EE
Sbjct: 378 EE 383
>TC219619 UP|NEU2_CATCO (P15211) Isoticin-neurophysin IT 2 precursor
[Contains: Isotocin (IT); Neurophysin IT 2], partial
(7%)
Length = 1063
Score = 27.3 bits (59), Expect = 6.6
Identities = 13/32 (40%), Positives = 22/32 (68%), Gaps = 1/32 (3%)
Frame = +3
Query: 60 DICEVFDE-EGNIDPIYDENGLDDIHEAFEKE 90
+IC + E E N+D IY ENG+ D+++ + +E
Sbjct: 54 NICHIRREREDNVD-IYGENGISDVNKIYPEE 146
>TC207416 similar to UP|O49349 (O49349) Transcription factor IIA large subunit
(TFIIA-L1), partial (62%)
Length = 1724
Score = 26.9 bits (58), Expect = 8.6
Identities = 14/53 (26%), Positives = 28/53 (52%)
Frame = +3
Query: 65 FDEEGNIDPIYDENGLDDIHEAFEKEEHNEPIYDEEYVLAEYGEYLETKKRFQ 117
FD++ + +P +E DD+ + + E+ N VLA++ + TK R++
Sbjct: 1158 FDDDDDDEPPLNEEDDDDLDDMEQGEDQN----THHLVLAQFDKVTRTKSRWK 1304
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 26.9 bits (58), Expect = 8.6
Identities = 9/15 (60%), Positives = 11/15 (73%)
Frame = +2
Query: 1 CVMCQGYGHIAIDCV 15
C C G+GH+A DCV
Sbjct: 134 CFNCGGFGHLARDCV 178
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.144 0.442
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,486,722
Number of Sequences: 63676
Number of extensions: 134683
Number of successful extensions: 870
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 850
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 870
length of query: 247
length of database: 12,639,632
effective HSP length: 95
effective length of query: 152
effective length of database: 6,590,412
effective search space: 1001742624
effective search space used: 1001742624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC122729.6


