

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122724.4 - phase: 0
(357 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
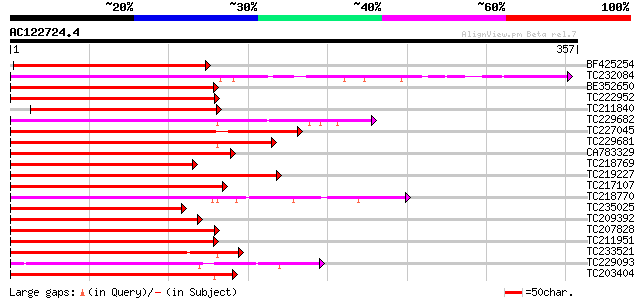
BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - gar... 254 4e-68
TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragmen... 232 2e-61
BE352650 215 2e-56
TC222952 homologue to UP|Q40173 (Q40173) Myb-related transcripti... 209 2e-54
TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4,... 208 3e-54
TC229682 UP|Q9XIU9 (Q9XIU9) GmMYB29A2 protein, complete 206 2e-53
TC227045 homologue to UP|Q43525 (Q43525) Transcription factor, p... 204 6e-53
TC229681 UP|Q9S7E3 (Q9S7E3) GmMYB29A1 protein, complete 202 2e-52
CA783329 homologue to GP|6651292|gb| myb-related transcription f... 201 4e-52
TC218769 UP|Q9XIU8 (Q9XIU8) GmMYB29A2 protein, complete 198 3e-51
TC219227 homologue to UP|Q84U52 (Q84U52) MYB2, partial (36%) 198 4e-51
TC217107 homologue to UP|Q7X9I1 (Q7X9I1) MYB transcription facto... 198 4e-51
TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete 197 6e-51
TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor,... 196 1e-50
TC209392 homologue to UP|Q8S3Y1 (Q8S3Y1) Typical P-type R2R3 Myb... 196 1e-50
TC207828 homologue to UP|Q6SL45 (Q6SL45) MYB6, partial (52%) 195 3e-50
TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcri... 193 9e-50
TC233521 similar to UP|O04140 (O04140) Myb protein, partial (44%) 193 1e-49
TC229093 homologue to PIR|T51621|T51621 myb-like protein [import... 192 1e-49
TC203404 GmMYB16 189 2e-48
>BF425254 homologue to PIR|S26605|S26 myb-related protein 1 - garden petunia,
partial (29%)
Length = 375
Score = 254 bits (649), Expect = 4e-68
Identities = 115/124 (92%), Positives = 119/124 (95%)
Frame = +3
Query: 3 RSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLR 62
RSPCCDKVGLKKGPWTPEEDQKLLAYI+EHGHGSWRALPAKAGLQRCGKSCRLRWTNYLR
Sbjct: 3 RSPCCDKVGLKKGPWTPEEDQKLLAYIEEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLR 182
Query: 63 PDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGID 122
PDIKRGKF +QEEQTIIQLHALLGNRWS+I+THL KRTDNEIKNYWNTHLKK L KM ID
Sbjct: 183 PDIKRGKFXMQEEQTIIQLHALLGNRWSSISTHLPKRTDNEIKNYWNTHLKKXLNKMSID 362
Query: 123 PVTH 126
PVTH
Sbjct: 363 PVTH 374
>TC232084 UP|Q84PP2 (Q84PP2) Transcription factor MYB101 (Fragment), complete
Length = 1383
Score = 232 bits (591), Expect = 2e-61
Identities = 153/380 (40%), Positives = 216/380 (56%), Gaps = 26/380 (6%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR+PC D+ GLKKGPWTPEED+ L+ YI +HGHGSWRALP A L RCGKSCRLRWTNY
Sbjct: 53 MGRTPCSDENGLKKGPWTPEEDRILVDYIQKHGHGSWRALPKHARLNRCGKSCRLRWTNY 232
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPDIKRGKFS +EEQ II LHA+LGN+WSAIA HL RTDNEIKN+WNTHLKK+L +MG
Sbjct: 233 LRPDIKRGKFSEEEEQLIINLHAVLGNKWSAIAGHLPGRTDNEIKNFWNTHLKKKLLQMG 412
Query: 121 IDPVTHKPKND--ALLSSSDN--AHSKTASNLSHMAQWESA-RLEAEARLVRESKIRSHN 175
+DPVTH+P++D LLS+ A + SNL++ +A RL+++A + +K++
Sbjct: 413 LDPVTHRPRSDHLNLLSNLQQLLAATNIVSNLTNTWDTNNALRLQSDA--TQLAKLQLLQ 586
Query: 176 SLLHHNHLKAWNINLESPTSTLSFNENVAPIMNS------GEIGGENSNTINN------- 222
++LH + +PTS L P +S E+ G N + + N
Sbjct: 587 NILH-------VLGDTTPTSNLDLLNPFGPSSSSLPDGFLNEVLGLNQSKLQNLYNGYSA 745
Query: 223 GNEKNNNDDDNNVVSMIEFVGTN-------SSSIVKEEGGDDQDQDQWKGYESSLTFTSN 275
N+ N + ++ G SS +K E GD+Q Y SS F S
Sbjct: 746 QNQPNLQSFEPPHQQQVQPTGNQHMDGGSIDSSCMKPEKGDEQLD---VPYSSSTPFNS- 913
Query: 276 LHHELTMSMDQSVDDVIAEEGFTNLLLKTNSEDLSLSESGGESNNGGDGGSGSGSEFYED 335
L + ++ S + S T++L++ ++ + +E S++ G YE+
Sbjct: 914 LPNLVSASPECS----------THVLMQMENKVMP-NECSDPSSSTSTNFEMWGDFMYEE 1060
Query: 336 -NNNYWNNILNLVNSSPSDS 354
++ YW ++++ + P S
Sbjct: 1061VSDAYWKDLIDKETN*PVTS 1120
>BE352650
Length = 528
Score = 215 bits (548), Expect = 2e-56
Identities = 94/132 (71%), Positives = 111/132 (83%), Gaps = 1/132 (0%)
Frame = +1
Query: 1 MGRSPCCDKVG-LKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTN 59
MGRSPCC++ +KKGPWTPEED+KL+ YI +HGHGSWR LP +AGL RCGKSCRLRWTN
Sbjct: 103 MGRSPCCEESSSVKKGPWTPEEDEKLIDYISKHGHGSWRTLPKRAGLNRCGKSCRLRWTN 282
Query: 60 YLRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKM 119
YLRPDIKRGKFS +E+ II H++LGN+WS IA HL RTDNEIKNYW TH++K+L KM
Sbjct: 283 YLRPDIKRGKFSEDDERIIINFHSVLGNKWSKIAAHLPGRTDNEIKNYWTTHIRKKLLKM 462
Query: 120 GIDPVTHKPKND 131
GIDP T+KP+ D
Sbjct: 463 GIDPETYKPRTD 498
>TC222952 homologue to UP|Q40173 (Q40173) Myb-related transcription factor,
partial (47%)
Length = 612
Score = 209 bits (532), Expect = 2e-54
Identities = 95/132 (71%), Positives = 109/132 (81%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR PCCDKVGLKKGPWT EED+KL+++I +G WRA+P AGL RCGKSCRLRWTNY
Sbjct: 125 MGRQPCCDKVGLKKGPWTAEEDKKLISFILTNGQCCWRAVPKLAGLLRCGKSCRLRWTNY 304
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG S EE+ +I LHA LGNRWS IA+HL RTDNEIKN+WNTH+KK+L KMG
Sbjct: 305 LRPDLKRGLLSEYEEKMVIDLHAQLGNRWSKIASHLPGRTDNEIKNHWNTHIKKKLKKMG 484
Query: 121 IDPVTHKPKNDA 132
IDPVTHKP +A
Sbjct: 485 IDPVTHKPLPNA 520
>TC211840 homologue to UP|Q8LBF0 (Q8LBF0) Myb-related protein M4, partial
(38%)
Length = 516
Score = 208 bits (530), Expect = 3e-54
Identities = 91/120 (75%), Positives = 106/120 (87%)
Frame = +2
Query: 14 KGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNYLRPDIKRGKFSLQ 73
KGPWT EEDQKL+ YI +HG+G+WR LP AGLQRCGKSCRLRWTNYLRPDIKRG+FS +
Sbjct: 2 KGPWTTEEDQKLIDYIQKHGYGNWRTLPKNAGLQRCGKSCRLRWTNYLRPDIKRGRFSFE 181
Query: 74 EEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKNDAL 133
EE+TIIQLH++LGN+WSAIA+ L RTDNEIKNYWNTH++KRL +MGIDPVTH P+ D L
Sbjct: 182 EEETIIQLHSILGNKWSAIASRLPGRTDNEIKNYWNTHIRKRLLRMGIDPVTHSPRLDLL 361
>TC229682 UP|Q9XIU9 (Q9XIU9) GmMYB29A2 protein, complete
Length = 1249
Score = 206 bits (523), Expect = 2e-53
Identities = 111/247 (44%), Positives = 150/247 (59%), Gaps = 16/247 (6%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+PCC+K+GLKKGPWTPEEDQ L++YI +HGHG+WRALP AGL RCGKSCRLRW NY
Sbjct: 84 MVRAPCCEKMGLKKGPWTPEEDQILMSYIQKHGHGNWRALPKLAGLLRCGKSCRLRWINY 263
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPDIKRG FS +EE+ II++H LLGNRWSAIA L RTDNEIKN W+THLKKRL
Sbjct: 264 LRPDIKRGNFSSEEEEIIIKMHELLGNRWSAIAAKLPGRTDNEIKNVWHTHLKKRLLNSD 443
Query: 121 IDPVTHKPK------NDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVRESKIRSH 174
I KP+ N + L+ + S ++LS + + + +++ I S
Sbjct: 444 IQKRVSKPRIKRSDSNSSTLTQLEPTSSACTTSLSDFSSFSEGTKNMD-NMIKSEDIESV 620
Query: 175 NSLLHHNHLKAWN---INLESPT----STLSFNENVAP---IMNSGEIGGENSNTINNGN 224
+++ W+ ++ ES T ++ + + +AP NS E + S N N
Sbjct: 621 ETIMPPIDESFWSEATVDYESSTMMTSNSWTISNELAPPQYQFNSVESFQQQSVDYNGSN 800
Query: 225 EKNNNDD 231
+ ++ D
Sbjct: 801 DDHDGMD 821
>TC227045 homologue to UP|Q43525 (Q43525) Transcription factor, partial (55%)
Length = 560
Score = 204 bits (518), Expect = 6e-53
Identities = 101/185 (54%), Positives = 123/185 (65%), Gaps = 1/185 (0%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCC+K KG WT EED +L++YI HG G WR+LP AGL RCGKSCRLRW NY
Sbjct: 15 MGRSPCCEKAHTNKGAWTKEEDHRLISYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 194
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG FSL+E+Q II+LH+LLGN+WS IA L RTDNEIKNYWNTH++++L G
Sbjct: 195 LRPDLKRGNFSLEEDQLIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKLLSRG 374
Query: 121 IDPVTHKPKNDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVRESKIR-SHNSLLH 179
IDP TH+P N D++H + A+ + ES E L E I H+
Sbjct: 375 IDPATHRPLN-------DSSHQEPAAVSAPPKHQESFHHERCPDLNLELTISPPHHPQPD 533
Query: 180 HNHLK 184
H HLK
Sbjct: 534 HPHLK 548
>TC229681 UP|Q9S7E3 (Q9S7E3) GmMYB29A1 protein, complete
Length = 795
Score = 202 bits (514), Expect = 2e-52
Identities = 97/174 (55%), Positives = 119/174 (67%), Gaps = 6/174 (3%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+PCC+K+GLKKGPWTPEEDQ L++YI +HGHG+WRALP AGL RCGKSCRLRW NY
Sbjct: 1 MVRAPCCEKMGLKKGPWTPEEDQILMSYIQKHGHGNWRALPKLAGLLRCGKSCRLRWINY 180
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPDIKRG FS +EE+ II++H LLGNRWSAIA L RTDNEIKN W+THLKKRL
Sbjct: 181 LRPDIKRGNFSSEEEEIIIKMHELLGNRWSAIAAKLPGRTDNEIKNVWHTHLKKRLMNSD 360
Query: 121 IDPVTHKPK------NDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVRE 168
+ KP+ N + L+ S+ S + S + + + + RE
Sbjct: 361 TNKRVSKPRIKRSDSNSSTLTQSEPTSSSGCTTSSDFSSFSEGTKNMDNMIKRE 522
>CA783329 homologue to GP|6651292|gb| myb-related transcription factor
{Pimpinella brachycarpa}, partial (48%)
Length = 443
Score = 201 bits (511), Expect = 4e-52
Identities = 94/143 (65%), Positives = 112/143 (77%), Gaps = 1/143 (0%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR P CDK+G+KKGPWT EED+KL+ +I +G WRA+P AGL+RCGKSCRLRWTNY
Sbjct: 3 MGRQPRCDKLGVKKGPWTAEEDKKLINFILTNGQCCWRAVPKLAGLKRCGKSCRLRWTNY 182
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG + EEQ +I LHA LGNRWS IA L RTDNEIKN+WNTH+KK+L K+G
Sbjct: 183 LRPDLKRGLLTEAEEQLVIDLHARLGNRWSKIAARLPGRTDNEIKNHWNTHIKKKLLKIG 362
Query: 121 IDPVTHKP-KNDALLSSSDNAHS 142
IDPVTH+P K A SS D++ S
Sbjct: 363 IDPVTHEPLKKQAAASSQDSSSS 431
>TC218769 UP|Q9XIU8 (Q9XIU8) GmMYB29A2 protein, complete
Length = 828
Score = 198 bits (504), Expect = 3e-51
Identities = 88/118 (74%), Positives = 103/118 (86%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+PCC+K+GLKKGPW PEEDQ L +YID+HGHG+WRALP +AGL RCGKSCRLRW NY
Sbjct: 1 MVRAPCCEKMGLKKGPWAPEEDQILTSYIDKHGHGNWRALPKQAGLLRCGKSCRLRWINY 180
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTK 118
LRPDIKRG F+++EE+TII+LH +LGNRWSAIA L RTDNEIKN W+T+LKKRL K
Sbjct: 181 LRPDIKRGNFTIEEEETIIKLHDMLGNRWSAIAAKLPGRTDNEIKNVWHTNLKKRLLK 354
>TC219227 homologue to UP|Q84U52 (Q84U52) MYB2, partial (36%)
Length = 1311
Score = 198 bits (503), Expect = 4e-51
Identities = 95/171 (55%), Positives = 119/171 (69%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR CC K L+KG W+PEED+KLL +I ++GHG W ++P +AGLQRCGKSCRLRW NY
Sbjct: 4 MGRHSCCYKQKLRKGLWSPEEDEKLLRHITKYGHGCWSSVPKQAGLQRCGKSCRLRWINY 183
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+ RG FS +EE II+LHA+LGNRWS IA L RTDNEIKN WN+ LKK+L + G
Sbjct: 184 LRPDLNRGTFSQEEETLIIELHAVLGNRWSQIAAQLPGRTDNEIKNLWNSCLKKKLRQRG 363
Query: 121 IDPVTHKPKNDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVRESKI 171
IDPVTHKP ++ D S+ SN ++ ES + + + R S I
Sbjct: 364 IDPVTHKPLSEVENGEEDKTRSQELSNELNLLNSESFKSDEGSYEQRASSI 516
>TC217107 homologue to UP|Q7X9I1 (Q7X9I1) MYB transcription factor R2R3 type,
partial (68%)
Length = 1058
Score = 198 bits (503), Expect = 4e-51
Identities = 85/137 (62%), Positives = 107/137 (78%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCC+K KG WT EED +L++YI HG G WR+LP AGL RCGKSCRLRW NY
Sbjct: 266 MGRSPCCEKAHTNKGAWTKEEDDRLISYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 445
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG F+ +E++ II+LH+LLGN+WS IA L RTDNEIKNYWNTH++++L G
Sbjct: 446 LRPDLKRGNFTEEEDELIIKLHSLLGNKWSLIAGRLPGRTDNEIKNYWNTHIRRKLLNRG 625
Query: 121 IDPVTHKPKNDALLSSS 137
IDP TH+P N+A +++
Sbjct: 626 IDPATHRPLNEAATAAT 676
>TC218770 UP|Q9XIU5 (Q9XIU5) GmMYB29B2 protein, complete
Length = 1316
Score = 197 bits (501), Expect = 6e-51
Identities = 116/275 (42%), Positives = 161/275 (58%), Gaps = 23/275 (8%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+PCC+K+GLKKGPW EEDQ L +YI +HGHG+WRALP +AGL RCGKSCRLRW NY
Sbjct: 112 MVRAPCCEKMGLKKGPWATEEDQILTSYIQKHGHGNWRALPKQAGLLRCGKSCRLRWINY 291
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPDIKRG F+++EE+TII+LH +LGNRWSAIA L RTDNEIKN W+T+LKKRL K
Sbjct: 292 LRPDIKRGNFTIEEEETIIKLHEMLGNRWSAIAAKLPGRTDNEIKNVWHTNLKKRLLKSD 471
Query: 121 IDPVTH----KPK------NDALLSSSDNAH--------SKTASNLSHMAQWESARLEAE 162
+ KPK N ++++ S+ AH + TA N S + + S +
Sbjct: 472 QSKPSSKRATKPKIKRSDSNSSIITQSEPAHLRFREMDTTSTACNTS-SSDFSSVTVGDS 648
Query: 163 ARLVRESKIRSHNSL--LHHNHLKAWNINLESPTSTLSFNENVAPIMNSGEIGGENSN-- 218
+++ I S ++ + + I+ E+PT + ++++ I N + +N
Sbjct: 649 KNIIKSEDIESMETMPVIDESFWSEAAIDDETPTMS---SQSLITISNDMPLQYPFANYE 819
Query: 219 -TINNGNEKNNNDDDNNVVSMIEFVGTNSSSIVKE 252
T + ++N DD F TN S + E
Sbjct: 820 ETFQQSHAYDSNFDDGMDFWYDIFTRTNDSIELSE 924
>TC235025 similar to UP|Q6R077 (Q6R077) MYB transcription factor, partial
(35%)
Length = 434
Score = 196 bits (499), Expect = 1e-50
Identities = 86/112 (76%), Positives = 99/112 (87%), Gaps = 1/112 (0%)
Frame = +1
Query: 1 MGRSPCCDK-VGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTN 59
MGRSPCC++ VG+KKGPWTPEED+KL+ YI +HGHGSWR LP +AGL RCGKSCRLRWTN
Sbjct: 97 MGRSPCCEENVGVKKGPWTPEEDEKLIDYISKHGHGSWRTLPKRAGLNRCGKSCRLRWTN 276
Query: 60 YLRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTH 111
YL PDIKRGKFS ++E+ II LH++LGN+WS IATHL RTDNEIKNYWNTH
Sbjct: 277 YLTPDIKRGKFSEEDERIIINLHSVLGNKWSKIATHLPGRTDNEIKNYWNTH 432
>TC209392 homologue to UP|Q8S3Y1 (Q8S3Y1) Typical P-type R2R3 Myb protein
(Fragment), partial (59%)
Length = 866
Score = 196 bits (498), Expect = 1e-50
Identities = 87/121 (71%), Positives = 102/121 (83%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR PCCDK G+KKGPWTPEED L++YI EHG G+WRA+PAK GL RC KSCRLRWTNY
Sbjct: 429 MGRPPCCDKEGVKKGPWTPEEDIILVSYIQEHGPGNWRAVPAKTGLSRCSKSCRLRWTNY 608
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRP IKRG F+ QEE+ II L LLGNRW+AIA++L +RTDN+IKNYWNTHL+K+L KM
Sbjct: 609 LRPGIKRGNFTEQEEKMIIHLQDLLGNRWAAIASYLPQRTDNDIKNYWNTHLRKKLKKMQ 788
Query: 121 I 121
+
Sbjct: 789 V 791
>TC207828 homologue to UP|Q6SL45 (Q6SL45) MYB6, partial (52%)
Length = 999
Score = 195 bits (495), Expect = 3e-50
Identities = 86/132 (65%), Positives = 102/132 (77%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGRSPCC+K KG WT EED++L+ YI HG G WR+LP AGL RCGKSCRLRW NY
Sbjct: 41 MGRSPCCEKEHTNKGAWTKEEDERLINYIKLHGEGCWRSLPKAAGLLRCGKSCRLRWINY 220
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG F+ +E++ II LH+LLGN+WS IA L RTDNEIKNYWNTH+K++L G
Sbjct: 221 LRPDLKRGNFTEEEDELIINLHSLLGNKWSLIAARLPGRTDNEIKNYWNTHIKRKLYSRG 400
Query: 121 IDPVTHKPKNDA 132
IDP TH+P N A
Sbjct: 401 IDPQTHRPLNAA 436
>TC211951 similar to GB|AAS10022.1|41619094|AY519552 MYB transcription factor
{Arabidopsis thaliana;} , partial (38%)
Length = 776
Score = 193 bits (491), Expect = 9e-50
Identities = 88/131 (67%), Positives = 104/131 (79%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR CC K L+KG W+PEED+KLL YI +HGHG W ++P AGLQRCGKSCRLRW NY
Sbjct: 92 MGRHSCCYKQKLRKGLWSPEEDEKLLNYITKHGHGCWSSVPKLAGLQRCGKSCRLRWINY 271
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPD+KRG FS QEE +II+LHA+LGNRWS IA L RTDNEIKN WN+ LKK+L + G
Sbjct: 272 LRPDLKRGAFSQQEENSIIELHAVLGNRWSQIAAQLPGRTDNEIKNLWNSCLKKKLRQRG 451
Query: 121 IDPVTHKPKND 131
IDP TH+P ++
Sbjct: 452 IDPNTHQPLSE 484
>TC233521 similar to UP|O04140 (O04140) Myb protein, partial (44%)
Length = 886
Score = 193 bits (490), Expect = 1e-49
Identities = 91/149 (61%), Positives = 114/149 (76%), Gaps = 2/149 (1%)
Frame = +2
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
M R+PCC+++GLKKGPWT EEDQ L+++I +GHG+WRALP +AGL RCGKSCRLRW NY
Sbjct: 20 MTRTPCCERMGLKKGPWTAEEDQILVSHIQRYGHGNWRALPKQAGLLRCGKSCRLRWINY 199
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPDIKRGKFS +EE TI++LH +LGNRWSAIA L RTDNEIKN+W+THL KR+ K G
Sbjct: 200 LRPDIKRGKFSKEEEDTILKLHGILGNRWSAIAASLPGRTDNEIKNFWHTHL-KRIQKSG 376
Query: 121 IDPVTHKPK--NDALLSSSDNAHSKTASN 147
+ + +A ++S NA S +N
Sbjct: 377 VHNGNPSSRILQEAQANTSSNASSVMIAN 463
>TC229093 homologue to PIR|T51621|T51621 myb-like protein [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(44%)
Length = 964
Score = 192 bits (489), Expect = 1e-49
Identities = 104/217 (47%), Positives = 132/217 (59%), Gaps = 19/217 (8%)
Frame = +3
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR PCCDK G+KKGPWTPEED L++YI EHG G+W+A+PA GL RC KSCRLRWTNY
Sbjct: 153 MGRPPCCDK-GVKKGPWTPEEDIILVSYIQEHGPGNWKAVPANTGLSRCSKSCRLRWTNY 329
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTK-- 118
LRP IKRG F+ QEE+ II L ALLGNRW+AIA +L +RTDN+IKNYWNT+LKK+L K
Sbjct: 330 LRPGIKRGNFTDQEEKMIIHLQALLGNRWAAIAAYLPQRTDNDIKNYWNTYLKKKLNKKL 509
Query: 119 -MGIDPVTHKPKNDALLSSSDNAHSKTASNLSHMAQWESARLEAEARLVRE--------- 168
G D + L +N + S QWE RL+ + + ++
Sbjct: 510 EAGSD------QGSLLGGGHNNIGFSSVSQPMSRGQWE-RRLQTDIHMAKKALSEALSPN 668
Query: 169 -------SKIRSHNSLLHHNHLKAWNINLESPTSTLS 198
+ + NS L + + +N N PT+T S
Sbjct: 669 KASSSSSTLLAESNSNLSNENSTFYNSNTNKPTTTQS 779
>TC203404 GmMYB16
Length = 1138
Score = 189 bits (480), Expect = 2e-48
Identities = 90/148 (60%), Positives = 104/148 (69%), Gaps = 5/148 (3%)
Frame = +1
Query: 1 MGRSPCCDKVGLKKGPWTPEEDQKLLAYIDEHGHGSWRALPAKAGLQRCGKSCRLRWTNY 60
MGR+PCC KVGL KGPWTP+ED L YI HG G W++LP KAGL RCGKSCRLRW NY
Sbjct: 106 MGRAPCCSKVGLHKGPWTPKEDALLTKYIQAHGEGQWKSLPKKAGLLRCGKSCRLRWMNY 285
Query: 61 LRPDIKRGKFSLQEEQTIIQLHALLGNRWSAIATHLAKRTDNEIKNYWNTHLKKRLTKMG 120
LRPDIKRG + +E+ II++H+LLGNRWS IA L RTDNEIKNYWNTHL K+L G
Sbjct: 286 LRPDIKRGNIAPEEDDLIIRMHSLLGNRWSLIAGRLPGRTDNEIKNYWNTHLSKKLKIQG 465
Query: 121 I-DPVTHK----PKNDALLSSSDNAHSK 143
D THK P+ +A +N K
Sbjct: 466 TEDTDTHKMLENPQEEAASDGGNNNKKK 549
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.309 0.128 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,513,454
Number of Sequences: 63676
Number of extensions: 249828
Number of successful extensions: 2126
Number of sequences better than 10.0: 233
Number of HSP's better than 10.0 without gapping: 1767
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1966
length of query: 357
length of database: 12,639,632
effective HSP length: 98
effective length of query: 259
effective length of database: 6,399,384
effective search space: 1657440456
effective search space used: 1657440456
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC122724.4


