

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122163.4 - phase: 0
(431 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
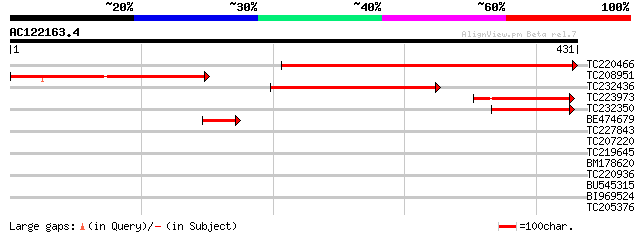
Score E
Sequences producing significant alignments: (bits) Value
TC220466 372 e-103
TC208951 223 1e-58
TC232436 107 1e-23
TC223973 similar to GB|AAP37861.1|30725678|BT008502 At2g21080 {A... 70 1e-12
TC232350 similar to GB|AAP37861.1|30725678|BT008502 At2g21080 {A... 65 8e-11
BE474679 similar to GP|18377809|gb| AT3g20300/MQC12_5 {Arabidops... 57 1e-08
TC227843 weakly similar to GB|AAM70534.1|21700821|AY124825 AT5g6... 30 1.6
TC207220 weakly similar to UP|Q9LSB0 (Q9LSB0) Emb|CAB70981.1, pa... 30 2.1
TC219645 similar to UP|O23207 (O23207) Minor allergen, partial (... 30 2.7
BM178620 29 4.7
TC220936 similar to GB|BAB10901.1|10177507|AB010699 serine prote... 28 8.0
BU545315 28 8.0
BI969524 28 8.0
TC205376 similar to UP|Q9MB45 (Q9MB45) IRE homolog 1 (Fragment),... 28 8.0
>TC220466
Length = 727
Score = 372 bits (956), Expect = e-103
Identities = 191/227 (84%), Positives = 208/227 (91%), Gaps = 2/227 (0%)
Frame = +2
Query: 207 YLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLVTASQLIFL 266
YLQI RLDEFA VFQRETEVG+ILLEHLR+RRNLR+ISHRFR FIL+SLLLVTASQLIFL
Sbjct: 2 YLQILRLDEFARVFQRETEVGSILLEHLRMRRNLRIISHRFRGFILASLLLVTASQLIFL 181
Query: 267 LMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASKWHICATIN 326
LMV KPHADVDILK GEL LVSITLVSGL ILLR ATKITHKA S+TGLA+KWHICATIN
Sbjct: 182 LMVTKPHADVDILKAGELALVSITLVSGLLILLRGATKITHKAHSITGLAAKWHICATIN 361
Query: 327 SFDNIDGETPTALIA-SAQAMAADISWGSSEDESGDEEDELNNTK-MMPIQTRTISFHKR 384
SFDNI+GETPT A SAQA+AA+ISWGS +DE GDEEDEL+NTK ++PI T+TISFHKR
Sbjct: 362 SFDNIEGETPTTANAISAQAIAANISWGSDDDEVGDEEDELDNTKLLLPIYTQTISFHKR 541
Query: 385 QALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVGI 431
QALVTY+ENNRAGITV+GFMLD WLHSIFGIQLALCLWLLNKTVG+
Sbjct: 542 QALVTYLENNRAGITVFGFMLDLFWLHSIFGIQLALCLWLLNKTVGV 682
>TC208951
Length = 653
Score = 223 bits (568), Expect = 1e-58
Identities = 109/154 (70%), Positives = 123/154 (79%), Gaps = 2/154 (1%)
Frame = +2
Query: 1 MEEEREVSFLAMNKENSGIYHDG--KELRRFRAYLRWVYVDQSNLCKAGISWSVFFTLAY 58
ME+ER L N E HD ELR FR+YLRWVYVD SN+CKAG+SW+VFFTLA+
Sbjct: 194 MEDERGSLLLPKNMEEHHPAHDRGTTELRNFRSYLRWVYVDHSNVCKAGLSWTVFFTLAF 373
Query: 59 VVPILSHFLLDCSTTCDADHRRPYHVPVQISLSVFATLSFISLSKWDRKYGFSKFLFLDK 118
VVP LSHFLLDC T CD DH RPYH+PVQISLSVFA LSFIS+S WDR+YGF KFLFLDK
Sbjct: 374 VVPTLSHFLLDCPT-CDRDHSRPYHIPVQISLSVFAALSFISISSWDRRYGFRKFLFLDK 550
Query: 119 VSDESLKIQRGYAIQMKRTMKLILLWGLPCFICQ 152
+ DESLKIQRGY+ QM+ TMKLI+ WGLPCFI +
Sbjct: 551 LGDESLKIQRGYSEQMQGTMKLIMRWGLPCFIAE 652
>TC232436
Length = 741
Score = 107 bits (267), Expect = 1e-23
Identities = 53/129 (41%), Positives = 80/129 (61%)
Frame = +1
Query: 199 CVLFRLIGYLQIQRLDEFAPVFQRETEVGTILLEHLRIRRNLRVISHRFRAFILSSLLLV 258
CVLF L+ LQ+ D++ + QRE +V + EH+R+R +L ISHRFR ++L L+V
Sbjct: 13 CVLFHLVCSLQLIHFDDYGKLLQRENDVLVFMEEHIRLRFHLSKISHRFRIYLLLQFLVV 192
Query: 259 TASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYILLRSATKITHKAQSLTGLASK 318
TASQ + LL V + + GG+ + ++ V G+ I+L +ATKI+H+AQ + LAS+
Sbjct: 193 TASQFVTLLQVTGFGGALTFISGGDFAVSTLVQVVGIIIVLHAATKISHRAQGIVSLASR 372
Query: 319 WHICATINS 327
WH T S
Sbjct: 373 WHALVTCTS 399
>TC223973 similar to GB|AAP37861.1|30725678|BT008502 At2g21080 {Arabidopsis
thaliana;} , partial (13%)
Length = 325
Score = 70.5 bits (171), Expect = 1e-12
Identities = 33/77 (42%), Positives = 49/77 (62%)
Frame = +3
Query: 353 GSSEDESGDEEDELNNTKMMPIQTRTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHS 412
G + D+ D +D +N + I + SF RQ LVTY+++N GITVYG+ LDR LH+
Sbjct: 12 GLASDDDSDSDDS-SNIHVSVIPPQLSSFQTRQTLVTYLQHNHGGITVYGYSLDRGLLHT 188
Query: 413 IFGIQLALCLWLLNKTV 429
+F + +L LW+L+K V
Sbjct: 189 LFAFEFSLVLWILSKVV 239
>TC232350 similar to GB|AAP37861.1|30725678|BT008502 At2g21080 {Arabidopsis
thaliana;} , partial (13%)
Length = 767
Score = 64.7 bits (156), Expect = 8e-11
Identities = 29/63 (46%), Positives = 42/63 (66%)
Frame = +2
Query: 367 NNTKMMPIQTRTISFHKRQALVTYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLN 426
N+ + I + SF RQ LVTY+++N GITVYG+ LDR LH++F + +L LW+L+
Sbjct: 416 NSARGSVIPPQLSSFQTRQTLVTYLQHNHGGITVYGYSLDRGLLHTLFAFEFSLVLWILS 595
Query: 427 KTV 429
K V
Sbjct: 596 KVV 604
>BE474679 similar to GP|18377809|gb| AT3g20300/MQC12_5 {Arabidopsis
thaliana}, partial (6%)
Length = 100
Score = 57.4 bits (137), Expect = 1e-08
Identities = 21/29 (72%), Positives = 24/29 (82%)
Frame = +2
Query: 147 PCFICQCVYKIWWYISGASQIPYYGEVYV 175
PCF+ +C YKIWWY SGASQIP+ G VYV
Sbjct: 14 PCFVAECAYKIWWYASGASQIPFLGNVYV 100
>TC227843 weakly similar to GB|AAM70534.1|21700821|AY124825
AT5g62930/MQB2_230 {Arabidopsis thaliana;} , partial
(64%)
Length = 884
Score = 30.4 bits (67), Expect = 1.6
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = -1
Query: 61 PILSHFLLDCSTTCDADHRRPYHVPVQISLSVFA 94
P L +F+ S +C DHR YH P+ + S A
Sbjct: 317 PSLKNFVCQPSVSCILDHRLIYHTPISFATSTQA 216
>TC207220 weakly similar to UP|Q9LSB0 (Q9LSB0) Emb|CAB70981.1, partial (16%)
Length = 880
Score = 30.0 bits (66), Expect = 2.1
Identities = 22/54 (40%), Positives = 30/54 (54%), Gaps = 7/54 (12%)
Frame = +2
Query: 245 HRFRAFILS---SLLLVTASQLIFLLMVIKPHADVDILKGGE----LGLVSITL 291
H F F +S +L L TAS LIFL ++I +A+ D L+ GLVS+ L
Sbjct: 464 HLFTVFAISDALALTLSTASTLIFLSILISRYAEEDFLRSLPFKLIFGLVSLFL 625
>TC219645 similar to UP|O23207 (O23207) Minor allergen, partial (73%)
Length = 1162
Score = 29.6 bits (65), Expect = 2.7
Identities = 8/26 (30%), Positives = 18/26 (68%)
Frame = +2
Query: 137 TMKLILLWGLPCFICQCVYKIWWYIS 162
T+ ++L WG PC++ C +K+ +++
Sbjct: 1055 TLSILLGWGFPCWVLLCFWKV*MFVN 1132
>BM178620
Length = 424
Score = 28.9 bits (63), Expect = 4.7
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +2
Query: 158 WWYISGASQIPYYGEVYVSGII 179
W Y+SG ++PYY V+ S II
Sbjct: 287 WNYVSGKIKLPYYTSVFPSSII 352
>TC220936 similar to GB|BAB10901.1|10177507|AB010699 serine protease-like
protein {Arabidopsis thaliana;} , partial (27%)
Length = 854
Score = 28.1 bits (61), Expect = 8.0
Identities = 19/63 (30%), Positives = 36/63 (56%)
Frame = +1
Query: 239 NLRVISHRFRAFILSSLLLVTASQLIFLLMVIKPHADVDILKGGELGLVSITLVSGLYIL 298
NL++IS F+ + LLL+T+ +FLL+ + P + ++K L ++ ++ +YIL
Sbjct: 13 NLQLISGLFQHTSKAGLLLITSLLDLFLLLFLSPIFVLSMVKIMNL-MLRLSFWKNIYIL 189
Query: 299 LRS 301
S
Sbjct: 190 WHS 198
>BU545315
Length = 577
Score = 28.1 bits (61), Expect = 8.0
Identities = 11/18 (61%), Positives = 12/18 (66%), Gaps = 1/18 (5%)
Frame = -3
Query: 143 LWGLPCFIC-QCVYKIWW 159
LW P +IC CV KIWW
Sbjct: 185 LWQSPYWIC*SCVVKIWW 132
>BI969524
Length = 252
Score = 28.1 bits (61), Expect = 8.0
Identities = 11/16 (68%), Positives = 14/16 (86%)
Frame = -1
Query: 416 IQLALCLWLLNKTVGI 431
+QLAL LW+L KTVG+
Sbjct: 231 VQLALXLWVLTKTVGV 184
>TC205376 similar to UP|Q9MB45 (Q9MB45) IRE homolog 1 (Fragment), partial (45%)
Length = 2377
Score = 28.1 bits (61), Expect = 8.0
Identities = 15/43 (34%), Positives = 21/43 (47%)
Frame = +3
Query: 389 TYMENNRAGITVYGFMLDRTWLHSIFGIQLALCLWLLNKTVGI 431
TY +N GI+ +G + T+L G LCL TVG+
Sbjct: 1569 TYDPSNFGGISTFGICIPCTYLEVQSGSGCTLCLLGYRATVGV 1697
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,946,873
Number of Sequences: 63676
Number of extensions: 351226
Number of successful extensions: 2667
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 2632
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2661
length of query: 431
length of database: 12,639,632
effective HSP length: 100
effective length of query: 331
effective length of database: 6,272,032
effective search space: 2076042592
effective search space used: 2076042592
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC122163.4


