

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.2 - phase: 0 /pseudo
(547 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
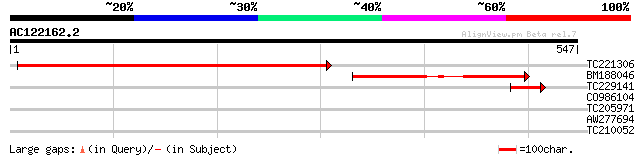
Score E
Sequences producing significant alignments: (bits) Value
TC221306 similar to UP|Q9SKB3 (Q9SKB3) Poly(ADP-ribose) glycohyd... 449 e-126
BM188046 202 4e-52
TC229141 weakly similar to UP|Q9SKB3 (Q9SKB3) Poly(ADP-ribose) g... 57 2e-08
CO986104 30 3.6
TC205971 29 6.2
AW277694 28 8.1
TC210052 similar to UP|Q87C92 (Q87C92) Cation:proton antiporter,... 28 8.1
>TC221306 similar to UP|Q9SKB3 (Q9SKB3) Poly(ADP-ribose) glycohydrolase,
partial (14%)
Length = 921
Score = 449 bits (1155), Expect = e-126
Identities = 226/305 (74%), Positives = 260/305 (85%), Gaps = 2/305 (0%)
Frame = +3
Query: 8 RSILPYLPVVMRSSSLFWPSQVVGALRELGCGRVDSGQLLFIFITELRNSLSLSSEPLAP 67
+SIL YLPVVMR SS FWPSQ V LRELG GRV+SG F I++LRN+LSLSSEPLAP
Sbjct: 6 QSILAYLPVVMRCSSPFWPSQAVETLRELGGGRVNSGHHFFQAISDLRNALSLSSEPLAP 185
Query: 68 SAAHGYALFFDELISREECRKWFEEVLPSLGDLLLRLPSLLEAHYENAD-MVIDGK-GAT 125
SA HGYALFFDE++S EE KWF+EVLP+LG+LLLRLPSLLE+HY+N D M IDG+ GA
Sbjct: 186 SAPHGYALFFDEVMSGEESSKWFQEVLPALGNLLLRLPSLLESHYQNTDNMAIDGEAGAM 365
Query: 126 VRTGLRMLDSQEAGIVFLTQELIAALLACSFLCLFPVHDRYEKQLQPVNFDELFASLYND 185
+ T LR+LDSQ+ GIVFLTQELIAALL+CS CLFPV DR L +NFD LF SLY+D
Sbjct: 366 LTTALRLLDSQQPGIVFLTQELIAALLSCSLFCLFPVSDRPVIHLPMINFDVLFGSLYDD 545
Query: 186 YTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWSTSVKPLC 245
Y+QKQE+KIWCI+HYFQRI+S MPKG+VSFERKVLP+++D IHISYP+A+FWSTS PLC
Sbjct: 546 YSQKQENKIWCIVHYFQRISSEMPKGIVSFERKVLPFKNDSIHISYPDANFWSTSAIPLC 725
Query: 246 RFEVKSSGLIEDHSSETVEVDFANEYLGGGALRRGCVQEEIRFMISPELIAGMIFLPSMA 305
RFEV SSGLIED SS VEVDFAN+YLGGGAL RGCVQEEIRFM+SPEL+ GM+FLP+MA
Sbjct: 726 RFEVHSSGLIEDQSSGAVEVDFANKYLGGGALGRGCVQEEIRFMVSPELMVGMLFLPAMA 905
Query: 306 DNEAI 310
DNEAI
Sbjct: 906 DNEAI 920
>BM188046
Length = 437
Score = 202 bits (513), Expect = 4e-52
Identities = 109/171 (63%), Positives = 117/171 (67%)
Frame = +2
Query: 331 SGDYVDDKEVDPFGRRKTRIVAIDALCGPGMRQYREKFLLREINKAFCGFLQQSQYQRDQ 390
SGDYVD++EVD GRRKTRIVAIDALC PGMRQYR FLLREINKAFCGFL Q +YQ Q
Sbjct: 5 SGDYVDEREVDTLGRRKTRIVAIDALCSPGMRQYRANFLLREINKAFCGFLYQCKYQPYQ 184
Query: 391 KIPQENVCSSLLVVFSMTF*HQYGLLSASLVVQLSVLNIFRFS*FDAMETSEGKYSYQEI 450
KI QEN C+S L Y S S MET EG+ S +I
Sbjct: 185 KILQENGCTSALF---------YAATSTS------------------METDEGEISNHKI 283
Query: 451 RNSQNDYDMMENSNDIGVATGNWGCGAFGGDPEVKTIIQWLAASQAGRPFI 501
NSQNDY M+ N+IGVATGNWGCGAFGGDPEVKTIIQWLAASQA RPFI
Sbjct: 284 TNSQNDYHGMDQGNNIGVATGNWGCGAFGGDPEVKTIIQWLAASQALRPFI 436
>TC229141 weakly similar to UP|Q9SKB3 (Q9SKB3) Poly(ADP-ribose)
glycohydrolase, partial (13%)
Length = 717
Score = 57.4 bits (137), Expect = 2e-08
Identities = 27/34 (79%), Positives = 31/34 (90%)
Frame = +1
Query: 484 VKTIIQWLAASQAGRPFIAYYTFGSGALQNLDKI 517
VKTIIQWLA+SQA RPFIAYYTFG ALQ+LD++
Sbjct: 1 VKTIIQWLASSQALRPFIAYYTFGLEALQSLDEV 102
Score = 40.0 bits (92), Expect = 0.003
Identities = 23/45 (51%), Positives = 28/45 (62%), Gaps = 4/45 (8%)
Frame = +3
Query: 507 GSGALQNLDKILVGVVA----GLVLDFVAGMDSWRPMEHASGVLN 547
GS +L I +G +A G LDFV+ MDSW PMEHA+ VLN
Sbjct: 36 GSKTFHSLLHIWLGGIAEPRRGCSLDFVSEMDSWGPMEHAN*VLN 170
>CO986104
Length = 797
Score = 29.6 bits (65), Expect = 3.6
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = +1
Query: 175 FDELFASLYNDYTQKQEDKIWCI 197
F+EL + DY + + DK+WCI
Sbjct: 712 FEELXXAYVTDYKKVRNDKVWCI 780
>TC205971
Length = 673
Score = 28.9 bits (63), Expect = 6.2
Identities = 21/75 (28%), Positives = 35/75 (46%)
Frame = +2
Query: 180 ASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHISYPNASFWST 239
+SL++ C++ FQR S +P G+ F +K+LP + + YP+ S
Sbjct: 305 SSLFSRCQSSSTRTCHCLVR-FQRAISTLPLGLQEFGKKMLPIPRLLLLMVYPSHSTTLH 481
Query: 240 SVKPLCRFEVKSSGL 254
KP F + S+ L
Sbjct: 482 LRKPR*FFMIMSNKL 526
>AW277694
Length = 439
Score = 28.5 bits (62), Expect = 8.1
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = -1
Query: 223 EDDCIHISYPNASFWSTSVKPLCRFEVKSSG 253
+++CI I+ S WSTS+ PLC + G
Sbjct: 334 KENCITINNSRNSAWSTSLFPLCSSSMTKEG 242
>TC210052 similar to UP|Q87C92 (Q87C92) Cation:proton antiporter, partial
(3%)
Length = 668
Score = 28.5 bits (62), Expect = 8.1
Identities = 18/57 (31%), Positives = 30/57 (52%)
Frame = -1
Query: 173 VNFDELFASLYNDYTQKQEDKIWCIIHYFQRITSNMPKGVVSFERKVLPWEDDCIHI 229
+NF LF+ YN+ +Q + DK + I S++PK + E K+ + CIH+
Sbjct: 668 INF--LFSVHYNEASQLKIDKY*AHFNEAIAIFSSLPKKNIRSEDKITQQKIKCIHL 504
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.140 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,362,853
Number of Sequences: 63676
Number of extensions: 333981
Number of successful extensions: 1981
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1972
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1979
length of query: 547
length of database: 12,639,632
effective HSP length: 102
effective length of query: 445
effective length of database: 6,144,680
effective search space: 2734382600
effective search space used: 2734382600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC122162.2


