

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122162.15 - phase: 0
(342 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
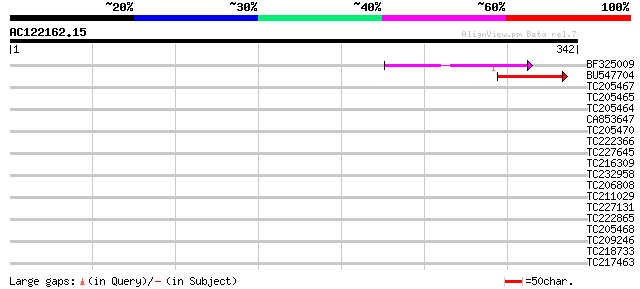
Score E
Sequences producing significant alignments: (bits) Value
BF325009 64 8e-11
BU547704 53 2e-07
TC205467 40 0.001
TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 34 0.11
TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 34 0.11
CA853647 similar to GP|5732065|gb|A contains similarity to Medic... 33 0.24
TC205470 32 0.32
TC222366 31 0.71
TC227645 similar to UP|Q9FZK1 (Q9FZK1) F17L21.13, partial (61%) 31 0.93
TC216309 similar to UP|Q9LNZ8 (Q9LNZ8) F9C16.20, partial (53%) 30 1.2
TC232958 30 1.2
TC206808 similar to UP|P83090 (P83090) 20 kDa protein (Fragment)... 29 2.7
TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%) 28 4.6
TC227131 similar to UP|Q8GSD7 (Q8GSD7) MRNA capping enzyme-like ... 28 4.6
TC222865 28 6.0
TC205468 weakly similar to GB|AAC35522.1|3600034|T12H20 T12H20.2... 28 6.0
TC209246 28 7.8
TC218733 28 7.8
TC217463 similar to PIR|E86378|E86378 protein F21J9.10 [imported... 28 7.8
>BF325009
Length = 382
Score = 64.3 bits (155), Expect = 8e-11
Identities = 39/113 (34%), Positives = 57/113 (49%), Gaps = 24/113 (21%)
Frame = +2
Query: 227 ICLFKGRFYVLDQSGQTVTVGPDSS-TELAAHPLYQRCVRGKNRKLLVESEGELLLLDIH 285
+C+FKG Y D +G+TV V PD LAA P++ G ++K LVESEG LLL+D++
Sbjct: 2 VCVFKGNLYGADSNGRTVRVRPDDGGLTLAAEPVF-----GGDKKFLVESEGALLLVDMY 166
Query: 286 QTFFQ-----------------------FSMKIFKLDENRKKWEKLKKLGDRI 315
+++ +F+LDE KKW +L L DR+
Sbjct: 167 LSYYSCTQGLFHEDFDEEDVAGMGWERTVKFDVFRLDEEGKKWVELTDLRDRV 325
>BU547704
Length = 600
Score = 53.1 bits (126), Expect = 2e-07
Identities = 24/42 (57%), Positives = 31/42 (73%)
Frame = -2
Query: 295 IFKLDENRKKWEKLKKLGDRILFVGSGCSFSASALDLCLPKG 336
+F LDE KKW +L LG+R+LF+G C+FSASA DL L +G
Sbjct: 548 VFXLDEEGKKWVELTDLGERVLFLGDDCAFSASAKDLNLGRG 423
>TC205467
Length = 1504
Score = 40.4 bits (93), Expect = 0.001
Identities = 79/343 (23%), Positives = 124/343 (36%), Gaps = 19/343 (5%)
Frame = +1
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAPDSI 65
W +LP ++L I RL + D IR +VC WHS + +D ++ Q P L Y P
Sbjct: 37 WADLPAELLESILSRL-ILADNIRASAVCRRWHS--VASDVRVVN-QSPWLMYFP---KF 195
Query: 66 NNNNEIIDNINNANTSF----------CYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQI 115
+ E D + +F CY L +P + + P+ ++
Sbjct: 196 GDCYEFYDPVQRKTHTFELPELNGSRVCYTKDGWLLLYRP-----RTHRVFFFNPFTREL 360
Query: 116 TKNSSGKTQLLNPPF-VSLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQHGS 174
K P F ++ + AFS P + + F+ +H+
Sbjct: 361 IK---------LPRFEMTYQIVAFSCAPTSPGCVLFTVKHVS------------------ 459
Query: 175 YLYPQTVLAVTCHGKKPLVLGTLSYCSSKPVLFHDQDKRWTPVSNLSTANGDICLFKGRF 234
P V TC+ T++Y + P +S+ + G F
Sbjct: 460 ---PTVVAISTCY-PGATEWTTVNYQNRLPF--------------VSSIWNKLVFCNGLF 585
Query: 235 YVLDQSGQTVTVGPDSST--ELAAHPLYQRCVR---GKN---RKLLVESEGELLLLDIHQ 286
Y L +G P T LA P +C KN K + E EG++L++
Sbjct: 586 YCLSLTGWLGVFDPVECTWSVLAVPP--PKCPENFFAKNWWKGKFMAEHEGDILVI---Y 750
Query: 287 TFFQFSMKIFKLDENRKKWEKLKKLGDRILFVGSGCSFSASAL 329
T + IFKLD+ KWE++ L LF S S + L
Sbjct: 751 TCCSENPIIFKLDQTLMKWEEMTTLDGVTLFASFLSSHSRTDL 879
>TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (15%)
Length = 490
Score = 33.9 bits (76), Expect = 0.11
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = +3
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHS 39
W++LP ++L LI RL ++ D +R VC WHS
Sbjct: 177 WSDLPTELLELILSRLSLD-DNVRASVVCKRWHS 275
>TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (17%)
Length = 756
Score = 33.9 bits (76), Expect = 0.11
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = +1
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHS 39
W++LP ++L LI RL ++ D +R VC WHS
Sbjct: 181 WSDLPTELLELILSRLSLD-DNVRASVVCKRWHS 279
>CA853647 similar to GP|5732065|gb|A contains similarity to Medicago
truncatula N7 protein (GB:Y17613) {Arabidopsis
thaliana}, partial (13%)
Length = 475
Score = 32.7 bits (73), Expect = 0.24
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = +1
Query: 5 DWTELPRDILFLISQRLDVELDLIRFRSVCSTW 37
+W +LPRD++ I Q+L L R + VCS W
Sbjct: 61 NWLDLPRDVVCTIFQKLGAIEILTRAQRVCSVW 159
>TC205470
Length = 580
Score = 32.3 bits (72), Expect = 0.32
Identities = 17/41 (41%), Positives = 24/41 (58%)
Frame = +2
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDH 46
W +LP ++L I RL + D IR +VC WHS + +DH
Sbjct: 143 WADLPAELLESILSRL-ILADNIRASAVCRRWHS--VASDH 256
>TC222366
Length = 860
Score = 31.2 bits (69), Expect = 0.71
Identities = 25/119 (21%), Positives = 52/119 (43%), Gaps = 15/119 (12%)
Frame = +2
Query: 236 VLDQSGQTVTVGPDSSTELAAHPLYQRCVRGKNRKL---------LVESEGELLLLDIHQ 286
+ ++ GQ + SS + A L RC R L +VE GE++++ + +
Sbjct: 113 ITNKDGQEIVYFLSSSGTVVACNLTSRCFLEYXRLLPVFSEYSIDIVECGGEMVVVLLSE 292
Query: 287 TFFQFSMKIFKLDENRKKWEKLKKLGDRILFVGSG------CSFSASALDLCLPKGKLC 339
S++++K DE + W+++ + + G C ++ + +CL +LC
Sbjct: 293 FLESTSLRVWKYDEANRCWQQIAAMPAAMSQEWYGKKADINCVGASGRIFICLNSPELC 469
>TC227645 similar to UP|Q9FZK1 (Q9FZK1) F17L21.13, partial (61%)
Length = 1671
Score = 30.8 bits (68), Expect = 0.93
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +3
Query: 6 WTELPRDILFLISQRLDVELDLIRFRSVCSTWHS 39
W + P D+ + RL + RFRSVC W+S
Sbjct: 354 WKDFPEDLFEAVIARLPIST-FFRFRSVCRQWNS 452
>TC216309 similar to UP|Q9LNZ8 (Q9LNZ8) F9C16.20, partial (53%)
Length = 1118
Score = 30.4 bits (67), Expect = 1.2
Identities = 45/193 (23%), Positives = 74/193 (38%), Gaps = 15/193 (7%)
Frame = +1
Query: 83 CYLSKRSFFLVKPPQE-----------QQQQETLIRCRPWLIQITKNSSGKTQLLNPPFV 131
C+ + + F P +E + T RCRP+L+ IT P +
Sbjct: 64 CHCASYNAFTFSPTREFITTTTTNHIMSKTSSTTTRCRPFLLSITN--------ATPSYN 219
Query: 132 SLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQHGSYLYPQTVLAVTCHGKKP 191
+L S A L V +F A L+ + D+ N+H S++ P+T + C
Sbjct: 220 TLVSEAVRLL---VPSARFEASKLKVVLLEDQ-----INKHASFI-PRTYILSHCDFTAN 372
Query: 192 LVLGTLSYCSSKPVL-FHDQD---KRWTPVSNLSTANGDICLFKGRFYVLDQSGQTVTVG 247
L L + + + + ++++D W V N D+CL F G
Sbjct: 373 LTLAVSNVINLEQLRGWYEKDDVVAEWKKVQN------DMCLHVHCF----------VSG 504
Query: 248 PDSSTELAAHPLY 260
P+S +LAA Y
Sbjct: 505 PNSFLDLAAELRY 543
>TC232958
Length = 758
Score = 30.4 bits (67), Expect = 1.2
Identities = 14/29 (48%), Positives = 21/29 (72%)
Frame = +1
Query: 9 LPRDILFLISQRLDVELDLIRFRSVCSTW 37
LP+D++ I RL V+ L+RF+SVC +W
Sbjct: 61 LPQDLITEILLRLPVK-SLVRFKSVCKSW 144
>TC206808 similar to UP|P83090 (P83090) 20 kDa protein (Fragment), complete
Length = 1034
Score = 29.3 bits (64), Expect = 2.7
Identities = 16/43 (37%), Positives = 25/43 (57%)
Frame = +1
Query: 97 QEQQQQETLIRCRPWLIQITKNSSGKTQLLNPPFVSLKSTAFS 139
Q+QQQ+ETL+ C WL+ +T + +L P + LK + S
Sbjct: 712 QKQQQRETLLFCL-WLVPMTSSGKLLRRL*RPCLIPLKFSFLS 837
>TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%)
Length = 858
Score = 28.5 bits (62), Expect = 4.6
Identities = 26/96 (27%), Positives = 38/96 (39%)
Frame = +3
Query: 7 TELPRDILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQFPLLKYVPAPDSIN 66
T L D++ LI L + LIR +VC WHS + + L ++ P
Sbjct: 153 TNLSTDLIELILSLLPIPT-LIRASTVCKLWHSIISSSSFSTLS------NHLNQPWFFL 311
Query: 67 NNNEIIDNINNANTSFCYLSKRSFFLVKPPQEQQQQ 102
+ I + NN + +F S F L P Q Q
Sbjct: 312 HGIHNISSKNNQSFAFDPASNTWFLLPTPQHHHQHQ 419
>TC227131 similar to UP|Q8GSD7 (Q8GSD7) MRNA capping enzyme-like protein,
partial (38%)
Length = 1812
Score = 28.5 bits (62), Expect = 4.6
Identities = 17/42 (40%), Positives = 22/42 (51%)
Frame = -1
Query: 12 DILFLISQRLDVELDLIRFRSVCSTWHSSPIPNDHNILPFQF 53
D++ SQ L V + LIR +CS D +ILPFQF
Sbjct: 894 DVVSYASQHLPVSIKLIRS*FICS---------DSHILPFQF 796
>TC222865
Length = 1064
Score = 28.1 bits (61), Expect = 6.0
Identities = 19/41 (46%), Positives = 23/41 (55%), Gaps = 2/41 (4%)
Frame = +3
Query: 39 SSPIPNDHNILPFQF-PLLKYVPAPDSINNNNEI-IDNINN 77
S P+P NI PF F P + V SIN+N+E I N NN
Sbjct: 588 SPPLPPLLNIRPFYFSPFQRIVKNEKSINSNDEYQIINTNN 710
>TC205468 weakly similar to GB|AAC35522.1|3600034|T12H20 T12H20.2
{Arabidopsis thaliana;} , partial (21%)
Length = 867
Score = 28.1 bits (61), Expect = 6.0
Identities = 20/65 (30%), Positives = 33/65 (50%)
Frame = +2
Query: 270 KLLVESEGELLLLDIHQTFFQFSMKIFKLDENRKKWEKLKKLGDRILFVGSGCSFSASAL 329
K + E EG+++++ T + IFKLD+ +WE++ L LF SF +S
Sbjct: 236 KFMTEHEGDIIVI---YTCSSENPIIFKLDQMLMEWEEMTTLDGVTLF----ASFLSSHA 394
Query: 330 DLCLP 334
+ LP
Sbjct: 395 RIDLP 409
>TC209246
Length = 970
Score = 27.7 bits (60), Expect = 7.8
Identities = 18/45 (40%), Positives = 23/45 (51%), Gaps = 2/45 (4%)
Frame = -1
Query: 217 VSNLSTANGDICLFKGR--FYVLDQSGQTVTVGPDSSTELAAHPL 259
V NLST N +CL R F+VL +G+ PD S + PL
Sbjct: 148 VMNLSTENTPVCLLSVRFSFWVLTSTGKP----PDLSVSIIKPPL 26
>TC218733
Length = 1493
Score = 27.7 bits (60), Expect = 7.8
Identities = 28/136 (20%), Positives = 48/136 (34%)
Frame = -3
Query: 71 IIDNINNANTSFCYLSKRSFFLVKPPQEQQQQETLIRCRPWLIQITKNSSGKTQLLNPPF 130
++ N++ TS L V+PP +L RP+ + I+ S + +P
Sbjct: 1317 LVGNVHFCTTSIFKLQPEHK-RVRPPIHSSNVSSLDHHRPFRLPISLGSLVPSYAHSPQH 1141
Query: 131 VSLKSTAFSHLPNAVNFIKFSAQHLRTDFIIDEDDFTFQNQHGSYLYPQTVLAVTCHGKK 190
L + S P D F F Q+ +P+T + +K
Sbjct: 1140 SQLLCFSLSSQPEQCG-----------------DPFLFPRQYCERCHPETTAIYSIKSRK 1012
Query: 191 PLVLGTLSYCSSKPVL 206
L + +SY K +L
Sbjct: 1011 SLSITMISYKKLKTIL 964
>TC217463 similar to PIR|E86378|E86378 protein F21J9.10 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(46%)
Length = 687
Score = 27.7 bits (60), Expect = 7.8
Identities = 28/113 (24%), Positives = 46/113 (39%), Gaps = 17/113 (15%)
Frame = -1
Query: 171 QHGSYLY-------PQTVLAVTCHGKKPLVLGTLSYCSSKPVLFHDQD---KRWTPVSNL 220
QH +L+ PQ L+++ HGKK + L S C S +HD + W ++
Sbjct: 435 QHQQHLHHK*VPIDPQN*LSLSYHGKKGMTLTGCSTCGSI*YTWHDSN*AKPFWCKALSI 256
Query: 221 STANGDICLFKGRFYVLDQSGQTVTVG-------PDSSTELAAHPLYQRCVRG 266
S + LF R+ ++ Q+ + D S + + P Y RG
Sbjct: 255 SHIH---LLFPNRY*LVSHHLQSFSCQNLSHLC*NDLSYKCSTSPSYHEAARG 106
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,412,966
Number of Sequences: 63676
Number of extensions: 289907
Number of successful extensions: 1550
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1536
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1545
length of query: 342
length of database: 12,639,632
effective HSP length: 98
effective length of query: 244
effective length of database: 6,399,384
effective search space: 1561449696
effective search space used: 1561449696
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC122162.15


