

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.9 + phase: 2 /pseudo/partial
(223 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
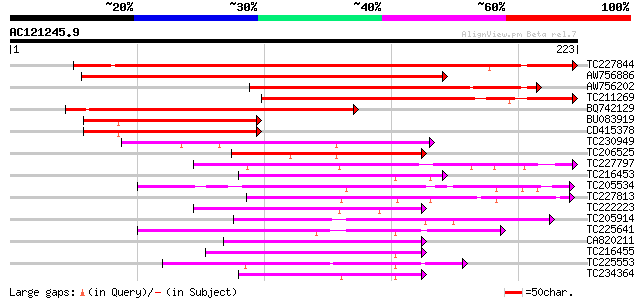
Score E
Sequences producing significant alignments: (bits) Value
TC227844 similar to UP|Q9FJN9 (Q9FJN9) Poly(A)-binding protein I... 350 3e-97
AW756886 similar to GP|10178188|db poly(A)-binding protein II-li... 236 6e-63
AW756202 similar to PIR|T50004|T50 RNA binding protein-like - Ar... 185 2e-47
TC211269 similar to UP|Q8LG48 (Q8LG48) Contains similarity to po... 185 2e-47
BQ742129 homologue to GP|15809838|gb| AT5g51120/MWD22_6 {Arabido... 167 3e-42
BU083919 similar to GP|10178188|dbj poly(A)-binding protein II-l... 103 8e-23
CD415378 similar to PIR|T50004|T500 RNA binding protein-like - A... 96 1e-20
TC230949 similar to UP|Q9LTX1 (Q9LTX1) Similarity to poly(A)-bin... 72 2e-13
TC206525 weakly similar to UP|Q41042 (Q41042) Pisum sativum L. (... 54 6e-08
TC227797 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, parti... 50 1e-06
TC216453 49 1e-06
TC205534 similar to GB|AAL32012.1|16930693|AF436830 AT3g07810/F1... 49 1e-06
TC227813 similar to UP|Q7XXQ1 (Q7XXQ1) Pre-mRNA splicing factor ... 48 3e-06
TC222223 similar to UP|Q8LKG6 (Q8LKG6) Cleavage stimulation fact... 47 5e-06
TC205914 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, ... 47 7e-06
TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding pr... 46 1e-05
CA820211 weakly similar to PIR|T06458|T064 nucleolin homolog - g... 46 1e-05
TC216455 weakly similar to UP|Q83CD7 (Q83CD7) Nucleic acid bindi... 45 3e-05
TC225553 weakly similar to UP|Q06106 (Q06106) Ypr112cp, partial ... 44 4e-05
TC234364 similar to UP|ROC5_NICSY (P19684) 33 kDa ribonucleoprot... 41 4e-04
>TC227844 similar to UP|Q9FJN9 (Q9FJN9) Poly(A)-binding protein II-like,
partial (80%)
Length = 1013
Score = 350 bits (898), Expect = 3e-97
Identities = 172/199 (86%), Positives = 186/199 (93%), Gaps = 1/199 (0%)
Frame = +3
Query: 26 MENDDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQI 85
MENDDVDM AD +E +LDDMKKRLKEMEDEAAAL+EMQAKVEKEMGS QDPANASASQ
Sbjct: 177 MENDDVDMHAADTSE-ELDDMKKRLKEMEDEAAALREMQAKVEKEMGSAQDPANASASQA 353
Query: 86 NREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEA 145
N+EE+D+RS+FVGNVDY+CTPE+VQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEF+EVEA
Sbjct: 354 NKEEIDSRSVFVGNVDYSCTPEEVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFLEVEA 533
Query: 146 VQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYM-GFRGRTPYAPPFAYAPA 204
VQEALLLNESELHGRQLKVTAKRTN+PGMKQ+RPRR NPYM GFRGRTPYAPPF Y +
Sbjct: 534 VQEALLLNESELHGRQLKVTAKRTNIPGMKQYRPRRTINPYMGGFRGRTPYAPPFIY--S 707
Query: 205 PYGYGKIPRFRMGMRYSPY 223
PYGYGK+PRFRM MRYSPY
Sbjct: 708 PYGYGKVPRFRMAMRYSPY 764
>AW756886 similar to GP|10178188|db poly(A)-binding protein II-like
{Arabidopsis thaliana}, partial (61%)
Length = 446
Score = 236 bits (602), Expect = 6e-63
Identities = 117/144 (81%), Positives = 129/144 (89%)
Frame = +3
Query: 29 DDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINRE 88
DD + D +LD+MK+RLKEME+EAAAL+EMQAKVEKE+GSVQDPANA+ASQ N+E
Sbjct: 15 DDEAAVDDDAAVKELDEMKRRLKEMEEEAAALREMQAKVEKEIGSVQDPANAAASQANKE 194
Query: 89 EVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQE 148
E DARS+FVGNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVE EAVQE
Sbjct: 195 EADARSVFVGNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEAEAVQE 374
Query: 149 ALLLNESELHGRQLKVTAKRTNVP 172
ALLLNESELHGRQLKV KRTNVP
Sbjct: 375 ALLLNESELHGRQLKVLPKRTNVP 446
>AW756202 similar to PIR|T50004|T50 RNA binding protein-like - Arabidopsis
thaliana, partial (49%)
Length = 439
Score = 185 bits (469), Expect = 2e-47
Identities = 92/115 (80%), Positives = 98/115 (85%)
Frame = +1
Query: 95 IFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNE 154
I V VDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVE EAVQEALLLNE
Sbjct: 103 ICVLQVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEAEAVQEALLLNE 282
Query: 155 SELHGRQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPYAPPFAYAPAPYGYG 209
SELHGRQLKV KRTNVPGMKQ+RPRR NPYM + R PY PP+ Y +PYGYG
Sbjct: 283 SELHGRQLKVLPKRTNVPGMKQYRPRR-FNPYMAYGFRRPYTPPYFY--SPYGYG 438
>TC211269 similar to UP|Q8LG48 (Q8LG48) Contains similarity to
poly(A)-binding protein II, partial (46%)
Length = 824
Score = 185 bits (469), Expect = 2e-47
Identities = 93/126 (73%), Positives = 104/126 (81%), Gaps = 2/126 (1%)
Frame = +1
Query: 100 VDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEALLLNESELHG 159
VDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AYVEFVE++AVQ ALLLNESELHG
Sbjct: 112 VDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAYVEFVEIDAVQNALLLNESELHG 291
Query: 160 RQLKVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPY--APPFAYAPAPYGYGKIPRFRMG 217
RQLKV+AKRTNVPGMKQ+ RRP+ GFR R P+ AP F PYGYG++PR+R
Sbjct: 292 RQLKVSAKRTNVPGMKQYFGRRPA----GFRSRRPFMSAPFF----PPYGYGRVPRYRRP 447
Query: 218 MRYSPY 223
RY PY
Sbjct: 448 TRYRPY 465
>BQ742129 homologue to GP|15809838|gb| AT5g51120/MWD22_6 {Arabidopsis
thaliana}, partial (39%)
Length = 429
Score = 167 bits (424), Expect = 3e-42
Identities = 81/115 (70%), Positives = 98/115 (84%)
Frame = +3
Query: 23 D*DMENDDVDMIGADNNEADLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASA 82
D D E D + + ++ +L+DMKKRLKE+E+EA+AL+EMQAKVEKEMG+VQD + SA
Sbjct: 87 DADAEQQD-EQLAPNHTNKELEDMKKRLKEIEEEASALREMQAKVEKEMGAVQDSSGTSA 263
Query: 83 SQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAY 137
+Q +EEVDARSI+VGNVDYACTPE+VQQHFQSCGTVNRVTI TDKFGQPKG+AY
Sbjct: 264 TQAEKEEVDARSIYVGNVDYACTPEEVQQHFQSCGTVNRVTILTDKFGQPKGFAY 428
>BU083919 similar to GP|10178188|dbj poly(A)-binding protein II-like
{Arabidopsis thaliana}, partial (39%)
Length = 421
Score = 103 bits (256), Expect = 8e-23
Identities = 50/72 (69%), Positives = 65/72 (89%), Gaps = 2/72 (2%)
Frame = +3
Query: 30 DVDMIGADNNEA--DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINR 87
D+DM AD++ A +LD+MK+RLKEME+EAAAL+EMQAKV+KE+GSVQDPAN++ASQ N+
Sbjct: 87 DIDMSAADDDAAVKELDEMKRRLKEMEEEAAALREMQAKVDKEIGSVQDPANSAASQANK 266
Query: 88 EEVDARSIFVGN 99
EE D+RS+FVGN
Sbjct: 267 EEADSRSVFVGN 302
>CD415378 similar to PIR|T50004|T500 RNA binding protein-like - Arabidopsis
thaliana, partial (36%)
Length = 455
Score = 95.9 bits (237), Expect = 1e-20
Identities = 48/78 (61%), Positives = 62/78 (78%), Gaps = 8/78 (10%)
Frame = -1
Query: 30 DVDMIGADNNEA--------DLDDMKKRLKEMEDEAAALKEMQAKVEKEMGSVQDPANAS 81
D+DM AD+ A +LD+MK+RLKEME+E AAL+E+QAK+EKE+GSVQDPANA+
Sbjct: 404 DIDMSAADDEAAVDDDAAVKELDEMKRRLKEMEEETAALREIQAKLEKEIGSVQDPANAA 225
Query: 82 ASQINREEVDARSIFVGN 99
ASQ N+EE DA S+F+GN
Sbjct: 224 ASQANKEEADALSVFLGN 171
>TC230949 similar to UP|Q9LTX1 (Q9LTX1) Similarity to poly(A)-binding
protein, partial (14%)
Length = 1287
Score = 72.0 bits (175), Expect = 2e-13
Identities = 49/131 (37%), Positives = 69/131 (52%), Gaps = 8/131 (6%)
Frame = +2
Query: 45 DMKKRLKEMEDEAAALKEMQAK------VEKEMGSVQDPANAS-ASQINREEVDARSIFV 97
D + + +++ A+ LK V+KE A+ S A+ E+VD+R+IFV
Sbjct: 173 DPRSSTRLVKENASTLKVSNGNANPALDVQKEFSKAHLTASGSNAAGRPSEDVDSRTIFV 352
Query: 98 GNVDYACTPEDVQQHFQSCGTVNRVTIRTD-KFGQPKGYAYVEFVEVEAVQEALLLNESE 156
NV +A T + + +HF G V +V I TD GQPKG AYVEF+ EA AL L+ +
Sbjct: 353 SNVHFAATKDGLSRHFNRFGDVLKVIIVTDAATGQPKGAAYVEFMRKEAADNALSLDNTS 532
Query: 157 LHGRQLKVTAK 167
R LKV K
Sbjct: 533 FMSRILKVIKK 565
>TC206525 weakly similar to UP|Q41042 (Q41042) Pisum sativum L. (clone
na-481-5), partial (20%)
Length = 752
Score = 53.9 bits (128), Expect = 6e-08
Identities = 31/82 (37%), Positives = 51/82 (61%), Gaps = 5/82 (6%)
Frame = +2
Query: 88 EEVDARSIFVGNVDYACTPEDV----QQHFQSCGTVNRVTIRTD-KFGQPKGYAYVEFVE 142
E +++IFV D + +++ Q+HF SCG + RV+I D + G KG+AYV+F +
Sbjct: 2 ERGQSQTIFVRGFDTSLGEDEIRGSLQEHFGSCGDITRVSIPKDYESGAVKGFAYVDFGD 181
Query: 143 VEAVQEALLLNESELHGRQLKV 164
+++ +AL L+E+EL G L V
Sbjct: 182 ADSMGKALELHETELGGYTLTV 247
>TC227797 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, partial (59%)
Length = 1737
Score = 49.7 bits (117), Expect = 1e-06
Identities = 43/168 (25%), Positives = 75/168 (44%), Gaps = 17/168 (10%)
Frame = +2
Query: 73 SVQDPANASASQINREEVDA-------RSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIR 125
++Q P++ S+S + +++ +FVG + + E ++++F G + +
Sbjct: 272 AMQGPSSGSSSSSGFQLLNSPFGDTTYTKVFVGGLAWETQSETMRRYFDQFGEILEAVVI 451
Query: 126 TDK-FGQPKGYAYVEFVEVEAVQEALLLNESELHGRQLKVTAKRTNVPGMKQFRPR---- 180
TDK G+ KGY +V F + EA + A + GR+ N+ + + RP
Sbjct: 452 TDKNTGRSKGYGFVTFRDPEAARRACADPSPVIDGRR-----ANCNLASLGRPRPPLPYG 616
Query: 181 --RPSNPYMGF--RGRTPYAPPFAY-APAPYGYGKIPRFRMGMRYSPY 223
RP++PY+G R Y F Y P Y Y + G+ Y PY
Sbjct: 617 RIRPASPYVGSLQPARGAYVGGFGYQQPVSYSY------QQGLVYPPY 742
>TC216453
Length = 1075
Score = 49.3 bits (116), Expect = 1e-06
Identities = 30/84 (35%), Positives = 44/84 (51%), Gaps = 2/84 (2%)
Frame = +1
Query: 91 DARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL 150
+ + ++V N+ ++ T D+ F CGTV V I K G+ KGYA+V E Q A+
Sbjct: 211 NVKKLYVVNLSWSLTAADITDLFAQCGTVTDVEIIKSKDGRSKGYAFVTMASGEEAQAAV 390
Query: 151 -LLNESELHGRQLKV-TAKRTNVP 172
+ EL GR ++V AKR P
Sbjct: 391 DKFDSYELSGRIIRVELAKRLKKP 462
>TC205534 similar to GB|AAL32012.1|16930693|AF436830 AT3g07810/F17A17_15
{Arabidopsis thaliana;} , partial (51%)
Length = 2354
Score = 49.3 bits (116), Expect = 1e-06
Identities = 49/180 (27%), Positives = 81/180 (44%), Gaps = 8/180 (4%)
Frame = +3
Query: 51 KEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQ 110
+ +E + A ++ Q + ++ GS A+AS + IFVG + T D +
Sbjct: 540 RTVEAKKAVPRDDQQNINRQSGS----AHASPGPGR-----TKKIFVGGLPSTITESDFK 692
Query: 111 QHFQSCGTVNRVTIRTDKFGQ-PKGYAYVEFVEVEAVQEALLLNESELHGRQLKVTAKRT 169
+F GT+ V + D Q P+G+ ++ + EAV L EL+G+ ++V +
Sbjct: 693 TYFDQFGTITDVVVMYDHNTQRPRGFGFITYDSEEAVDRVLYKTFHELNGKMVEV---KR 863
Query: 170 NVPGMKQFRPRRPSNPYMGFR----GRTPYAPPFA--YAPAPY-GYGKIPRFRMGMRYSP 222
VP K+ P +P +G+ + Y FA Y +P GYG RM R+SP
Sbjct: 864 AVP--KELSPGPSRSPLIGYNYGLTRASNYLNSFAQGYNMSPIGGYG----IRMDGRFSP 1025
>TC227813 similar to UP|Q7XXQ1 (Q7XXQ1) Pre-mRNA splicing factor (Fragment),
partial (77%)
Length = 1322
Score = 48.1 bits (113), Expect = 3e-06
Identities = 44/140 (31%), Positives = 65/140 (46%), Gaps = 11/140 (7%)
Frame = +2
Query: 94 SIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEALL- 151
S+ V N+ C PE+++ F+ G V V I D + G+P+G+A+V+FV+ EA
Sbjct: 299 SLLVRNIPLDCRPEELRVPFERFGPVRDVYIPKDYYSGEPRGFAFVQFVDPYDASEAQYH 478
Query: 152 LNESELHGRQLKV-----TAKRTNVPGMKQFRPRRPSNPYMGFR----GRTPYAPPFAYA 202
+N GR++ V T KR + R R P++ Y G R GR+
Sbjct: 479 MNRQIFAGREISVVVAEETRKRPEEMRHRTSRFRGPAS-YGGRRSSHYGRSRSRSITRSR 655
Query: 203 PAPYGYGKIPRFRMGMRYSP 222
PY G R+R YSP
Sbjct: 656 SPPYHSGSRNRYR-SRSYSP 712
>TC222223 similar to UP|Q8LKG6 (Q8LKG6) Cleavage stimulation factor 64,
partial (22%)
Length = 445
Score = 47.4 bits (111), Expect = 5e-06
Identities = 30/94 (31%), Positives = 48/94 (50%), Gaps = 2/94 (2%)
Frame = +2
Query: 73 SVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDK-FGQ 131
S P++A + + R +FVGN+ Y T E + + Q G V + D+ G+
Sbjct: 98 SSASPSSADVDFMATSQSQHRCVFVGNIPYDATEEQLIEICQEVGPVVSFRLVIDRETGK 277
Query: 132 PKGYAYVEFVEVE-AVQEALLLNESELHGRQLKV 164
PKGY + E+ + E A+ L E++GRQL+V
Sbjct: 278 PKGYGFCEYKDEETALSARRNLQGYEINGRQLRV 379
>TC205914 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, partial (9%)
Length = 1513
Score = 47.0 bits (110), Expect = 7e-06
Identities = 36/134 (26%), Positives = 66/134 (48%), Gaps = 8/134 (5%)
Frame = +3
Query: 89 EVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQE 148
+V+ +IFVGN+D + E+++Q+ G + V I+ KG+ +V+F + +E
Sbjct: 594 DVNNTTIFVGNLDLNVSEEELKQNSLQFGEIVSVKIQPG-----KGFGFVQFGTRASAEE 758
Query: 149 ALLLNESELHGRQL------KVTAKRTNVPG--MKQFRPRRPSNPYMGFRGRTPYAPPFA 200
A+ + ++ G+Q+ + R ++PG Q P + S Y +G YA A
Sbjct: 759 AIQKMQGKMIGQQVVRISWGRTLTARQDLPGGWGPQMDPNQWSAYYGYGQGYEAYAYGPA 938
Query: 201 YAPAPYGYGKIPRF 214
+ P+ Y YG P +
Sbjct: 939 HDPSLYAYGAYPGY 980
Score = 29.3 bits (64), Expect = 1.5
Identities = 13/27 (48%), Positives = 16/27 (59%)
Frame = +3
Query: 124 IRTDKFGQPKGYAYVEFVEVEAVQEAL 150
IR GQP+GY +VEFV A + L
Sbjct: 63 IRNKLTGQPEGYGFVEFVSHAAAERVL 143
>TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding protein 2
{Arabidopsis thaliana;} , partial (59%)
Length = 1223
Score = 46.2 bits (108), Expect = 1e-05
Identities = 40/153 (26%), Positives = 65/153 (42%), Gaps = 8/153 (5%)
Frame = +3
Query: 51 KEMEDEAAALKEMQAKVEKEMGSVQDPANASASQINREEVDARSIFVGNVDYACTPEDVQ 110
KE E+E AA + D + EE + IFVGN+ + E +
Sbjct: 300 KEGEEEEAAWENQGDTAWGTEEGGDDIEDGGEGGFAEEEAEEVKIFVGNLPFDFDSEKLA 479
Query: 111 QHFQSCGTV-------NRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQL 162
F+ GTV NR T R+ +G+ +V +E +++A+ + + EL+GR L
Sbjct: 480 SLFEQAGTVEVAEVIYNRATDRS------RGFGFVTMSTIEELEKAVKMFSGYELNGRVL 641
Query: 163 KVTAKRTNVPGMKQFRPRRPSNPYMGFRGRTPY 195
T + G + RP RP + + G P+
Sbjct: 642 --TVNKAAPKGAQPERPPRPPQSFRVYVGNLPW 734
>CA820211 weakly similar to PIR|T06458|T064 nucleolin homolog - garden pea,
partial (12%)
Length = 327
Score = 46.2 bits (108), Expect = 1e-05
Identities = 23/80 (28%), Positives = 42/80 (51%)
Frame = +1
Query: 85 INREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAYVEFVEVE 144
+ + ++I+V N+ Y D++ F+ CG + V + + G+ G+ VEF E
Sbjct: 37 LKEQNAPPKTIYVRNLSYRVEHADMENLFKECGKIVDVRLHRNHNGRLNGFGQVEFATAE 216
Query: 145 AVQEALLLNESELHGRQLKV 164
A ++AL L+ +EL R + V
Sbjct: 217 AAKKALELHNTELLRRPIGV 276
>TC216455 weakly similar to UP|Q83CD7 (Q83CD7) Nucleic acid binding domain
protein, partial (35%)
Length = 491
Score = 44.7 bits (104), Expect = 3e-05
Identities = 26/88 (29%), Positives = 43/88 (48%), Gaps = 1/88 (1%)
Frame = +2
Query: 78 ANASASQINREEVDARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKFGQPKGYAY 137
A ++ + + + ++V N+ ++ T D+ F GTV V I K G+ KGYA+
Sbjct: 191 ATEETPELTQPTDNVKKLYVVNLSWSLTAADINDLFAQSGTVTDVEIIKSKDGRSKGYAF 370
Query: 138 VEFVEVEAVQEAL-LLNESELHGRQLKV 164
V E Q A+ + EL GR ++V
Sbjct: 371 VTMASGEEAQAAVDKFDSYELSGRIIRV 454
>TC225553 weakly similar to UP|Q06106 (Q06106) Ypr112cp, partial (6%)
Length = 1371
Score = 44.3 bits (103), Expect = 4e-05
Identities = 30/125 (24%), Positives = 60/125 (48%), Gaps = 5/125 (4%)
Frame = +1
Query: 61 KEMQAKVEKEMGSVQDPANASASQINREEVD----ARSIFVGNVDYACTPEDVQQHFQSC 116
K++Q V + P N ++ ++ + + + NV + T +D+++ F
Sbjct: 688 KDLQGTVLDSHALILQPCNVKNDGQKQKTLEKDRSSTKLLIKNVAFEATEKDLRRLFSPF 867
Query: 117 GTVNRVTIRTDKFGQPKGYAYVEFVEVEAVQEAL-LLNESELHGRQLKVTAKRTNVPGMK 175
G + + + KFG +G+A+VE+V + Q AL L+ + L+GR L V + ++
Sbjct: 868 GQIKSLRLPM-KFGNHRGFAFVEYVTQQEAQNALKALSSTHLYGRHL-VIERAKEAESLE 1041
Query: 176 QFRPR 180
+ R R
Sbjct: 1042ELRAR 1056
>TC234364 similar to UP|ROC5_NICSY (P19684) 33 kDa ribonucleoprotein,
chloroplast precursor, partial (32%)
Length = 710
Score = 41.2 bits (95), Expect = 4e-04
Identities = 25/76 (32%), Positives = 44/76 (57%), Gaps = 2/76 (2%)
Frame = +2
Query: 91 DARSIFVGNVDYACTPEDVQQHFQSCGTVNRVTIRTDKF-GQPKGYAYVEFVEVEAVQEA 149
DA ++VGN+ Y+ T ++ + F GTV V I D+ + +G+A+V VE +EA
Sbjct: 371 DAGRLYVGNLPYSITNSELGELFGEAGTVASVEIVYDRVTDRSRGFAFVTMGSVEDAKEA 550
Query: 150 L-LLNESELHGRQLKV 164
+ + + S++ GR + V
Sbjct: 551 IRMFDGSQVGGRTVXV 598
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.137 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,199,068
Number of Sequences: 63676
Number of extensions: 103225
Number of successful extensions: 667
Number of sequences better than 10.0: 133
Number of HSP's better than 10.0 without gapping: 643
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 657
length of query: 223
length of database: 12,639,632
effective HSP length: 94
effective length of query: 129
effective length of database: 6,654,088
effective search space: 858377352
effective search space used: 858377352
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC121245.9


