

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121245.1 - phase: 0 /pseudo
(660 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
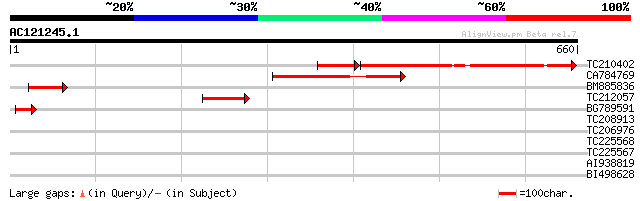
Score E
Sequences producing significant alignments: (bits) Value
TC210402 similar to UP|Q6Z7V1 (Q6Z7V1) LMBR1 integral membrane p... 263 3e-91
CA784769 215 4e-56
BM885836 64 3e-10
TC212057 similar to UP|Q6Z7V1 (Q6Z7V1) LMBR1 integral membrane p... 61 1e-09
BG789591 48 2e-05
TC208913 similar to UP|Q6Z7V1 (Q6Z7V1) LMBR1 integral membrane p... 32 0.90
TC206976 homologue to UP|Q84MM3 (Q84MM3) Lethal leaf spot 1-like... 30 3.4
TC225568 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 30 4.5
TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 30 4.5
AI938819 29 7.6
BI498628 28 10.0
>TC210402 similar to UP|Q6Z7V1 (Q6Z7V1) LMBR1 integral membrane protein-like,
partial (19%)
Length = 1178
Score = 263 bits (672), Expect(2) = 3e-91
Identities = 161/255 (63%), Positives = 182/255 (71%), Gaps = 4/255 (1%)
Frame = +3
Query: 409 KQMFCAENGKH**CCSLFRERIQQNLPSYHGYIHFTNSRQFL*PGHQLLRELENIQV*** 468
K F +NGK * CSLF +RIQQNLP +HG IHFTNS+Q+ *PGHQ+L ELENIQV **
Sbjct: 129 KNNF*TKNGKDQ*RCSLFWKRIQQNLPPHHGCIHFTNSKQYF*PGHQVLWELENIQVQ** 308
Query: 469 CRRYGWI*SIWSNNFAERALSSSARAQSWRTCFPFSQKFQHEYGC*VCQQQGQGSG*ECR 528
CRRYGWI* I SNN AER SS RAQSW TCF F ++FQ +YGC V QQQG GS *EC
Sbjct: 309 CRRYGWI*PIRSNNIAERTFSSPTRAQSW*TCFSFGKEFQCDYGCRV-QQQGHGSE*ECH 485
Query: 529 KRR*NNYHGRDQK*R--S*HEQKNWWQKVFSLKDKPE*RGVK*RFHPRNKFFLFNK*F-- 584
+ H +K*R HEQKNW QKV SLKDK +*RGV *RF P FFL +K*
Sbjct: 486 YQ-----HSGREK*RHSERHEQKNWQQKVCSLKDKLQ*RGV**RFDPGKSFFLLDK*C** 650
Query: 585 SSGHEFSTIFCDNLKMGINDAWI*KFKI*H*LQ*IPST****YLYIISKLNLFI*IS**Y 644
S H+FS+I C LKMGINDAWI*KF+I*H*LQ IP + * + SKL +FI*I **+
Sbjct: 651 FSEHKFSSIICIGLKMGINDAWI*KFEI*H*LQGIPPS--F*CSRVNSKLKIFI*IP**H 824
Query: 645 I*EIETSTIRTQRFW 659
I*EIETS+ R RFW
Sbjct: 825 I*EIETSSFRI*RFW 869
Score = 91.7 bits (226), Expect(2) = 3e-91
Identities = 42/49 (85%), Positives = 48/49 (97%)
Frame = +1
Query: 359 FYSLTPRQTSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTVFEKR 407
FYSLTP+QTSSVSLLMICSM+ARYAAPISYNFLNLIN+GG RKT+FE++
Sbjct: 4 FYSLTPKQTSSVSLLMICSMIARYAAPISYNFLNLINLGGGRKTIFEQK 150
>CA784769
Length = 409
Score = 215 bits (548), Expect = 4e-56
Identities = 111/154 (72%), Positives = 122/154 (79%)
Frame = +1
Query: 307 TILPSGVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSLTPRQ 366
TIL SGVDLSLFS+LVHA G EVLVQLAAF+PLMYMC+CTYYSLFKMGM MFYSLTP+Q
Sbjct: 1 TILTSGVDLSLFSLLVHAGGQQEVLVQLAAFIPLMYMCICTYYSLFKMGMMMFYSLTPKQ 180
Query: 367 TSSVSLLMICSMVARYAAPISYNFLNLINIGGDRKTVFEKRAKQMFCAENGKH**CCSLF 426
TSSVSLLMICSM+ARYAAPISYNFLNL N+ ENGK * CSLF
Sbjct: 181 TSSVSLLMICSMIARYAAPISYNFLNLFNL------------------ENGKDQ*RCSLF 306
Query: 427 RERIQQNLPSYHGYIHFTNSRQFL*PGHQLLREL 460
+RIQQNLP +HG IHFTNS+Q+ *PGHQ+L EL
Sbjct: 307 WKRIQQNLPPHHGCIHFTNSKQYF*PGHQVLWEL 408
>BM885836
Length = 422
Score = 63.5 bits (153), Expect = 3e-10
Identities = 28/46 (60%), Positives = 35/46 (75%)
Frame = +1
Query: 22 FLLTWAVVPLLQGYEDAGDFTVKARLRTSLHGNLVFYLSLGSVALF 67
F+L+ VVPL+QGYED GDF V RL T +H NL+FYL +GS+ LF
Sbjct: 109 FILSRVVVPLIQGYEDVGDFIVSERLETGVHVNLIFYLIIGSIGLF 246
>TC212057 similar to UP|Q6Z7V1 (Q6Z7V1) LMBR1 integral membrane protein-like,
partial (3%)
Length = 725
Score = 61.2 bits (147), Expect = 1e-09
Identities = 27/55 (49%), Positives = 39/55 (70%)
Frame = +2
Query: 225 RSEYTKFVLEALELEDTVKNYDRRDSTGWRYISCLRPERIGKVGAVLDTIEFLWR 279
+S Y +VLEA ELEDT+KN+DR +ST W S +RP + GK+G++ DT+ + R
Sbjct: 80 KSGYMTYVLEAFELEDTIKNFDRHNSTKWECNSSIRPAQTGKLGSLFDTLGVIQR 244
>BG789591
Length = 309
Score = 47.8 bits (112), Expect = 2e-05
Identities = 21/25 (84%), Positives = 21/25 (84%)
Frame = +2
Query: 7 NVTISFLWSLSYWSTFLLTWAVVPL 31
N ISF WS SYWSTFLLTWAVVPL
Sbjct: 233 NGGISFFWSWSYWSTFLLTWAVVPL 307
>TC208913 similar to UP|Q6Z7V1 (Q6Z7V1) LMBR1 integral membrane protein-like,
partial (7%)
Length = 846
Score = 32.0 bits (71), Expect = 0.90
Identities = 18/38 (47%), Positives = 22/38 (57%)
Frame = +1
Query: 471 RYGWI*SIWSNNFAERALSSSARAQSWRTCFPFSQKFQ 508
RYGW *SI N F +RA+ + AR QS +KFQ
Sbjct: 31 RYGWF*SIRINYFTKRAVLA*ARMQSRGASCSTCKKFQ 144
>TC206976 homologue to UP|Q84MM3 (Q84MM3) Lethal leaf spot 1-like protein,
partial (79%)
Length = 1469
Score = 30.0 bits (66), Expect = 3.4
Identities = 29/95 (30%), Positives = 40/95 (41%), Gaps = 13/95 (13%)
Frame = +3
Query: 312 GVDLSLFSILVHAAGHTEVLVQLAAFVPL--MYMC--VCTYYSLFKMG---------MFM 358
GVDL L + +H VL+ L VPL + C VC+ Y L +M
Sbjct: 195 GVDLVLRFLRLHLKAPKHVLLDLLKHVPLGSLPWCPRVCSLYGLMRMVGRKQRPPSLQCC 374
Query: 359 FYSLTPRQTSSVSLLMICSMVARYAAPISYNFLNL 393
+LT R + +ICSMV + +S L L
Sbjct: 375 LMTLTNRSLPRSTFSVICSMVTILSWRMSLILLTL 479
>TC225568 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (36%)
Length = 1287
Score = 29.6 bits (65), Expect = 4.5
Identities = 19/87 (21%), Positives = 37/87 (41%)
Frame = +2
Query: 137 KLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGES 196
+++ AH D N ++ + + + N + L+ N S +GGR
Sbjct: 392 EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTL---LIMTSNVGSSVIEKGGRKIGF 562
Query: 197 DMDYDTDEKSMASLRRRLRRAREQYYR 223
D+DYD + S ++ + +QY+R
Sbjct: 563 DLDYDEKDSSYNRIKSLVTEELKQYFR 643
>TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor ,
partial (61%)
Length = 2077
Score = 29.6 bits (65), Expect = 4.5
Identities = 19/87 (21%), Positives = 37/87 (41%)
Frame = +3
Query: 137 KLDDAHQDFSNAIVITQATSKQMSKRDSLRPYMNIIDKMLVQMLNEDPSFKPQGGRLGES 196
+++ AH D N ++ + + + N + L+ N S +GGR
Sbjct: 1092 EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTL---LIMTSNVGSSVIEKGGRKIGF 1262
Query: 197 DMDYDTDEKSMASLRRRLRRAREQYYR 223
D+DYD + S ++ + +QY+R
Sbjct: 1263 DLDYDEKDSSYNRIKSLVTEELKQYFR 1343
>AI938819
Length = 411
Score = 28.9 bits (63), Expect = 7.6
Identities = 12/38 (31%), Positives = 22/38 (57%)
Frame = +2
Query: 330 VLVQLAAFVPLMYMCVCTYYSLFKMGMFMFYSLTPRQT 367
+L+ +A ++ +Y+CVC Y F + +F+ LT T
Sbjct: 287 ILILIACYI-YVYVCVCAIYIAFSTRILIFFILTYSMT 397
>BI498628
Length = 308
Score = 28.5 bits (62), Expect = 10.0
Identities = 15/35 (42%), Positives = 18/35 (50%)
Frame = +1
Query: 312 GVDLSLFSILVHAAGHTEVLVQLAAFVPLMYMCVC 346
G DLSLF IL HT +L L V ++ C C
Sbjct: 181 GFDLSLFRILHSGTTHTVILPSLTLQVTVIDACHC 285
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.348 0.152 0.521
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,604,529
Number of Sequences: 63676
Number of extensions: 451503
Number of successful extensions: 4642
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 3421
Number of HSP's successfully gapped in prelim test: 141
Number of HSP's that attempted gapping in prelim test: 1166
Number of HSP's gapped (non-prelim): 3644
length of query: 660
length of database: 12,639,632
effective HSP length: 103
effective length of query: 557
effective length of database: 6,081,004
effective search space: 3387119228
effective search space used: 3387119228
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC121245.1


