

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
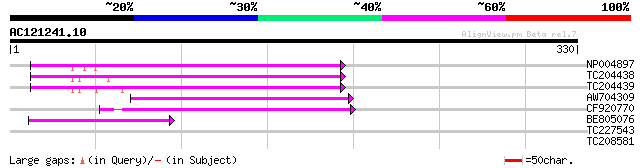
Query= AC121241.10 + phase: 0 /pseudo
(330 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP004897 gag-protease polyprotein 83 1e-16
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 82 3e-16
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 79 4e-15
AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 72 5e-13
CF920770 59 2e-09
BE805076 weakly similar to GP|9294121|dbj copia-like retrotransp... 50 1e-06
TC227543 30 2.0
TC208581 similar to UP|O64851 (O64851) Expressed protein (At2g26... 28 4.4
>NP004897 gag-protease polyprotein
Length = 1923
Score = 83.2 bits (204), Expect = 1e-16
Identities = 62/195 (31%), Positives = 95/195 (47%), Gaps = 12/195 (6%)
Frame = +1
Query: 13 PKFDG-HYDHWAMFMENFLRSKEY--WDLIENE-----ILVVEA----GVEQTEAQRKLI 60
P DG +Y++W M FL+S + W + + +L E G++ E K
Sbjct: 40 PILDGTNYEYWKARMVAFLKSLDSRTWKAVIKDWEHPKMLDTEGKPTDGLKPEEDWTKEE 219
Query: 61 KEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRD 120
E L N K N LF + ++ I K+ W+ +KT G+++V ++LQ L
Sbjct: 220 DELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATK 399
Query: 121 FEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEES 180
FE L MKE E I+ + L IAN A E M+ + KILRS+ + D V +IEE+
Sbjct: 400 FENLKMKEEECIHDFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEA 579
Query: 181 NNLDTMTIDELQSSL 195
++ + +DEL SL
Sbjct: 580 QDICNLRVDELIGSL 624
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 82.0 bits (201), Expect = 3e-16
Identities = 62/195 (31%), Positives = 94/195 (47%), Gaps = 12/195 (6%)
Frame = +1
Query: 13 PKFDG-HYDHWAMFMENFLRSKEY--WDLI----ENEILVVEAGVEQTEAQ-----RKLI 60
P DG +Y++W M FL+S + W + E+ ++ G E + K
Sbjct: 40 PILDGTNYEYWKARMVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKPTNELKPEEDWTKEE 219
Query: 61 KEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRD 120
E L N K N LF + ++ I K+ W+ +KT G+++V ++LQ L
Sbjct: 220 DELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATK 399
Query: 121 FEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEES 180
FE L MKE E I+ + L IAN A E M+ + KILRS+ + D V +IEE+
Sbjct: 400 FENLKMKEEECIHDFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEA 579
Query: 181 NNLDTMTIDELQSSL 195
++ M +DEL SL
Sbjct: 580 QDICNMRVDELIGSL 624
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 78.6 bits (192), Expect = 4e-15
Identities = 59/195 (30%), Positives = 96/195 (48%), Gaps = 12/195 (6%)
Frame = +1
Query: 13 PKFDG-HYDHWAMFMENFLRSKEY--WDLI----ENEILVVEAG--VEQTEAQRKLIKEQ 63
P DG +Y++W M FL+S + W + E+ ++ G ++ + + KE+
Sbjct: 40 PILDGSNYEYWKARMVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKPTDELKPEEDWTKEE 219
Query: 64 K---LMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRD 120
L N K N LF + ++ I K+ W+ +K G+++V ++LQ L
Sbjct: 220 DELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKITHEGTSKVKMSRLQLLATK 399
Query: 121 FEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEES 180
FE L MKE E I+ + L IAN A E ++ + KILRS+ + D V +IEE+
Sbjct: 400 FENLKMKEEECIHDFHMNILEIANACTALGERITDEKLVRKILRSLPKRFDMKVTAIEEA 579
Query: 181 NNLDTMTIDELQSSL 195
++ M +DEL SL
Sbjct: 580 QDICNMRVDELIGSL 624
>AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (23%)
Length = 423
Score = 71.6 bits (174), Expect = 5e-13
Identities = 42/131 (32%), Positives = 72/131 (54%), Gaps = 1/131 (0%)
Frame = -1
Query: 71 KNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRAQLQALRRDFEILHMKEGE 130
++ LF +S + I+ + K IWD +K + G +R+ Q+ LRR+FE+ MKE E
Sbjct: 420 RSCLFTGVSQMIFIRIMTLKSPKAIWDCLKEEYAGDDRIRSMQVLNLRREFEL*RMKESE 241
Query: 131 TINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDYVVCSIEESNNLDTMTIDE 190
TI Y ++ L I NK+K + + I EK L ++ + + + S+E + +L +T+ E
Sbjct: 240 TIKEYSNKLLGIVNKIKLLGSDFADSRIVEKNLVTVPERYEAFIASLENTKDLSKITLAE 61
Query: 191 LQSSL-VHEQR 200
+ +L EQR
Sbjct: 60 VLHALQAQEQR 28
>CF920770
Length = 581
Score = 59.3 bits (142), Expect = 2e-09
Identities = 40/150 (26%), Positives = 70/150 (46%), Gaps = 1/150 (0%)
Frame = -2
Query: 53 TEAQRKLIKEQKLMNLKTKNYLFQAISHDVLETILNKDTYKNIWDSMKTFF*GSNRVIRA 112
+E RK ++ NLK KN + A+ D + N + K +WD+++ G+ V R+
Sbjct: 451 SEEDRKRVQ----YNLKAKNIITSALGMDEYFRVSNCKSAKEMWDTLRLTHEGTTDVKRS 284
Query: 113 QLQALRRDFEILHMKEGETINTYFSRTLTIANKMKAHSEIMSQTIITEKILRSMVSKIDY 172
++ AL ++E+ M E I + R I N + A + + K+LR + +
Sbjct: 283 RINALTHEYELFRMNTNENIQSMQKRFTHIVNHLAALGKEFQNEDLINKVLRCLSREWQP 104
Query: 173 VVCSIEESNNLDTMTIDELQSSL-VHEQRI 201
V +I ES +L M++ L L HE +
Sbjct: 103 KVTAISESRDLSNMSLATLFGKLQEHEMEL 14
>BE805076 weakly similar to GP|9294121|dbj copia-like retrotransposable
element {Arabidopsis thaliana}, partial (2%)
Length = 388
Score = 50.4 bits (119), Expect = 1e-06
Identities = 27/86 (31%), Positives = 43/86 (49%), Gaps = 1/86 (1%)
Frame = +1
Query: 12 IPKFDG-HYDHWAMFMENFLRSKEYWDLIENEILVVEAGVEQTEAQRKLIKEQKLMNLKT 70
+P F+G +YD W + ME +L S++ WD++E + +Q K +K+ K N KT
Sbjct: 130 VPIFNGENYDFWRVKMETYLSSQDLWDIVEEGFTIPADTSALNASQEKELKKNKQKNSKT 309
Query: 71 KNYLFQAISHDVLETILNKDTYKNIW 96
L QA + + I+ T K W
Sbjct: 310 LFTLQQAETDPIFPRIMGAKTAKEAW 387
>TC227543
Length = 1489
Score = 29.6 bits (65), Expect = 2.0
Identities = 15/38 (39%), Positives = 26/38 (67%), Gaps = 1/38 (2%)
Frame = +2
Query: 159 TEKILRSMVS-KIDYVVCSIEESNNLDTMTIDELQSSL 195
TE +LRS+ S KID+ ++E NN+++ I+ + +SL
Sbjct: 221 TEDLLRSLPSSKIDWSAFAMENKNNVNSSAIENIGASL 334
>TC208581 similar to UP|O64851 (O64851) Expressed protein (At2g26190/T1D16.17),
partial (46%)
Length = 1304
Score = 28.5 bits (62), Expect = 4.4
Identities = 16/34 (47%), Positives = 18/34 (52%)
Frame = -2
Query: 243 TMLNLK*KKKKRYYCCPMLKTNMGRKKKCGFLTP 276
T L K KKKK Y+C P K GRK G + P
Sbjct: 1279 TPLTAKKKKKKEYFCIP--KKERGRKMVKGPICP 1184
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.144 0.439
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,870,235
Number of Sequences: 63676
Number of extensions: 142597
Number of successful extensions: 1194
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1187
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1194
length of query: 330
length of database: 12,639,632
effective HSP length: 98
effective length of query: 232
effective length of database: 6,399,384
effective search space: 1484657088
effective search space used: 1484657088
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC121241.10


