

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121239.4 - phase: 0
(442 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
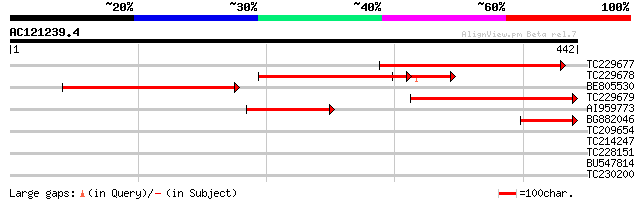
Sequences producing significant alignments: (bits) Value
TC229677 similar to UP|Q8VXD3 (Q8VXD3) N-acetylglucosaminyltrans... 270 7e-73
TC229678 homologue to UP|Q8GUA5 (Q8GUA5) N-acetylglucosaminyltra... 249 2e-66
BE805530 similar to GP|6779209|emb BETA-1 2-N-ACETYLGLUCOSAMINYL... 239 2e-63
TC229679 similar to UP|Q8VXD3 (Q8VXD3) N-acetylglucosaminyltrans... 206 2e-53
AI959773 homologue to GP|4467130|emb| glycosyltransferase like p... 130 2e-30
BG882046 similar to GP|18076146|emb N-acetylglucosaminaltransfer... 71 9e-13
TC209654 similar to UP|Q6NKP7 (Q6NKP7) At1g09800, partial (47%) 33 0.34
TC214247 homologue to UP|Q6I687 (Q6I687) Protein disulfide isome... 30 1.7
TC228151 weakly similar to UP|Q6UQ04 (Q6UQ04) Fatty acid desatur... 30 2.8
BU547814 29 4.8
TC230200 29 4.8
>TC229677 similar to UP|Q8VXD3 (Q8VXD3) N-acetylglucosaminyltransferase I ,
partial (32%)
Length = 438
Score = 270 bits (691), Expect = 7e-73
Identities = 128/145 (88%), Positives = 136/145 (93%)
Frame = +2
Query: 289 WLRLKENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNEVQVDWKSMDLSYLL 348
WLRLKENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLN+V+VDWKSMDLSYLL
Sbjct: 2 WLRLKENHKGRQFIRPEVCRTYNFGEHGSSLGQFFKQYLEPIKLNDVKVDWKSMDLSYLL 181
Query: 349 EDKYVIHFANIIKKAKPVSGADFAQKARHIDGDVRIKYKDQWDFENIAQQFGIFQEWKDG 408
EDKY +HFAN+IKKA PV GAD KA +IDG+VRIKYKDQ DFENIA QFGIFQEWKDG
Sbjct: 182 EDKYSMHFANVIKKATPVYGADMVLKAYNIDGNVRIKYKDQSDFENIAHQFGIFQEWKDG 361
Query: 409 VPRTAYKGVVVFRYQTTKRIFLVGP 433
VPRTAYKGVVVFRYQT++RIFLVGP
Sbjct: 362 VPRTAYKGVVVFRYQTSRRIFLVGP 436
>TC229678 homologue to UP|Q8GUA5 (Q8GUA5) N-acetylglucosaminyltransferase I,
partial (36%)
Length = 454
Score = 249 bits (636), Expect = 2e-66
Identities = 113/119 (94%), Positives = 117/119 (97%)
Frame = +2
Query: 195 QLFDKHNFNRVIILEDDMEIAPDFFDYFEAMATLLDKDKSIMAVSSWNDNGQKQFVHDPY 254
QLF KHNF+RVIILEDDMEIAPDFFDYFEA A+LL+KDKSIMAVSSWNDNGQKQFVHDPY
Sbjct: 2 QLFYKHNFSRVIILEDDMEIAPDFFDYFEAAASLLEKDKSIMAVSSWNDNGQKQFVHDPY 181
Query: 255 ELYRSDFFPGLGWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFG 313
ELYRSDFFPGLGWMLARSTW+ELSPKWPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFG
Sbjct: 182 ELYRSDFFPGLGWMLARSTWNELSPKWPKAYWDDWLRLKENHKGRQFIRPEVCRTYNFG 358
Score = 65.1 bits (157), Expect = 6e-11
Identities = 32/52 (61%), Positives = 39/52 (74%), Gaps = 3/52 (5%)
Frame = +3
Query: 299 RQFIRPEVCRTYNFGEH---GSSLGQFFKQYLEPIKLNEVQVDWKSMDLSYL 347
++ I+ + + EH GSSLGQFFKQYLEPIKLN+V+VDWK MDLSYL
Sbjct: 297 KRIIKDDSLSGPKYAEHIILGSSLGQFFKQYLEPIKLNDVKVDWKLMDLSYL 452
>BE805530 similar to GP|6779209|emb BETA-1 2-N-ACETYLGLUCOSAMINYLTRANSFERASE
I {Nicotiana tabacum}, partial (30%)
Length = 419
Score = 239 bits (609), Expect = 2e-63
Identities = 118/138 (85%), Positives = 131/138 (94%)
Frame = +2
Query: 42 EAENHCTAQIRSLIDQISLQQGRILDLQQERNRREQECSQIKSLVQDLERKDVRRLIDKV 101
EAENHCT+Q RSLIDQISLQQGRI+ L++ER RR+QEC Q+KSLVQDLERKD++RLIDK+
Sbjct: 2 EAENHCTSQTRSLIDQISLQQGRIVALEEERKRRDQECGQMKSLVQDLERKDLQRLIDKM 181
Query: 102 QVPVAAVVIMACNRADYLERTINSVLKYQRPISSRFPLFVSQDGSNSDVKRKALSYDELS 161
QVPVAAVVIMACNRADYLERTINSVLKYQRPISSR+PLFVSQDGSN +VK KALSYD+LS
Sbjct: 182 QVPVAAVVIMACNRADYLERTINSVLKYQRPISSRYPLFVSQDGSNPNVKSKALSYDQLS 361
Query: 162 HMQHLDFEPVQTERPGEL 179
+MQHLD EPVQTERPGEL
Sbjct: 362 YMQHLDXEPVQTERPGEL 415
>TC229679 similar to UP|Q8VXD3 (Q8VXD3) N-acetylglucosaminyltransferase I ,
partial (29%)
Length = 671
Score = 206 bits (524), Expect = 2e-53
Identities = 103/130 (79%), Positives = 111/130 (85%)
Frame = +2
Query: 313 GEHGSSLGQFFKQYLEPIKLNEVQVDWKSMDLSYLLEDKYVIHFANIIKKAKPVSGADFA 372
GEH S FFKQYL IKLN+V+VD K MDLSYLLEDKY + FAN+IKKA PV GAD
Sbjct: 2 GEHXXSXXXFFKQYLXXIKLNDVKVDXKXMDLSYLLEDKYSMXFANVIKKATPVYGADMV 181
Query: 373 QKARHIDGDVRIKYKDQWDFENIAQQFGIFQEWKDGVPRTAYKGVVVFRYQTTKRIFLVG 432
KA +IDG+VRIKYKDQ DFENIA QFGIFQEWKDGVPRTAYKGVVVFRYQT++RIFLVG
Sbjct: 182 LKASNIDGNVRIKYKDQSDFENIAHQFGIFQEWKDGVPRTAYKGVVVFRYQTSRRIFLVG 361
Query: 433 PESLKLLQIE 442
PESLKLLQIE
Sbjct: 362 PESLKLLQIE 391
>AI959773 homologue to GP|4467130|emb| glycosyltransferase like protein
{Arabidopsis thaliana}, partial (17%)
Length = 455
Score = 130 bits (326), Expect = 2e-30
Identities = 60/69 (86%), Positives = 65/69 (93%)
Frame = +1
Query: 185 IARHYKWALGQLFDKHNFNRVIILEDDMEIAPDFFDYFEAMATLLDKDKSIMAVSSWNDN 244
++ HYKWAL QLF KHNF+RVIILEDDMEIAPDFFDYFEA A+LL+KDKSIMAVSSWNDN
Sbjct: 1 LSGHYKWALDQLFYKHNFSRVIILEDDMEIAPDFFDYFEAAASLLEKDKSIMAVSSWNDN 180
Query: 245 GQKQFVHDP 253
GQKQFVHDP
Sbjct: 181 GQKQFVHDP 207
Score = 30.0 bits (66), Expect = 2.2
Identities = 12/12 (100%), Positives = 12/12 (100%)
Frame = +2
Query: 255 ELYRSDFFPGLG 266
ELYRSDFFPGLG
Sbjct: 419 ELYRSDFFPGLG 454
>BG882046 similar to GP|18076146|emb N-acetylglucosaminaltransferase I
{Nicotiana tabacum}, partial (6%)
Length = 396
Score = 71.2 bits (173), Expect = 9e-13
Identities = 35/44 (79%), Positives = 38/44 (85%)
Frame = +3
Query: 399 FGIFQEWKDGVPRTAYKGVVVFRYQTTKRIFLVGPESLKLLQIE 442
F F +DGVPRTAYKGVVVFRYQT++RIFLVGPE LKLLQIE
Sbjct: 150 FPAFGNSQDGVPRTAYKGVVVFRYQTSRRIFLVGPEYLKLLQIE 281
>TC209654 similar to UP|Q6NKP7 (Q6NKP7) At1g09800, partial (47%)
Length = 759
Score = 32.7 bits (73), Expect = 0.34
Identities = 26/93 (27%), Positives = 44/93 (46%), Gaps = 1/93 (1%)
Frame = +1
Query: 133 ISSRFPLFVSQDGSNSDVKRKALSYDELSHMQHLDFEPVQTERPGELIAYYKIARHYKWA 192
+ F FV Q S S R LS++ H+D + + RPGE++ ++ A + A
Sbjct: 319 LEEAFSKFVGQPVSVSCSSRTDAGVHALSNVCHVDIQRISKRRPGEVLPPHEPAVVGR-A 495
Query: 193 LGQLFDKHNFNRVIILEDDMEIAP-DFFDYFEA 224
+ KH+ + ++I D+ P DF F+A
Sbjct: 496 VNHFLQKHDSDLMVI---DVRCVPSDFHARFKA 585
>TC214247 homologue to UP|Q6I687 (Q6I687) Protein disulfide isomerase-like
protein, partial (53%)
Length = 991
Score = 30.4 bits (67), Expect = 1.7
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = -2
Query: 260 DFFPGLGWMLARSTWDELSPKWPKA 284
+FFP L M+ R TW SPK+ KA
Sbjct: 687 NFFPSLC*MIIRGTWSSFSPKYWKA 613
>TC228151 weakly similar to UP|Q6UQ04 (Q6UQ04) Fatty acid desaturase, partial
(37%)
Length = 614
Score = 29.6 bits (65), Expect = 2.8
Identities = 19/59 (32%), Positives = 28/59 (47%), Gaps = 2/59 (3%)
Frame = +1
Query: 164 QHLDFEPVQTERPGELIAYYKIARHYKWALGQLFDKHNFNRV--IILEDDMEIAPDFFD 220
+H F P Q E +YY + W +G + H+F R+ L EIAP+F+D
Sbjct: 205 EHYVFNPDQ-----ETYSYYGPLNYLTWHVGYHNEHHDFPRIPGNKLHKVKEIAPEFYD 366
>BU547814
Length = 648
Score = 28.9 bits (63), Expect = 4.8
Identities = 17/59 (28%), Positives = 24/59 (39%), Gaps = 3/59 (5%)
Frame = -1
Query: 251 HDPYELYRSDFFPG---LGWMLARSTWDELSPKWPKAYWDDWLRLKENHKGRQFIRPEV 306
H P E+ +SD FP +GW+L + A + DWL E I P +
Sbjct: 531 HPPNEIIQSDIFPRWALIGWLLTCCRKKHVEESVKLALFYDWLFFDERMDSIMNIEPAI 355
>TC230200
Length = 745
Score = 28.9 bits (63), Expect = 4.8
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = +3
Query: 400 GIFQEWKDGVPRTAYKGVVVFRYQTTKRIFLV 431
G+F+ W + VP T++ VV K++FLV
Sbjct: 417 GLFRPWTEFVPHTSFDRRVVSLLSRVKKVFLV 512
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.139 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,809,594
Number of Sequences: 63676
Number of extensions: 250144
Number of successful extensions: 1100
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1097
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1100
length of query: 442
length of database: 12,639,632
effective HSP length: 100
effective length of query: 342
effective length of database: 6,272,032
effective search space: 2145034944
effective search space used: 2145034944
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121239.4


