

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
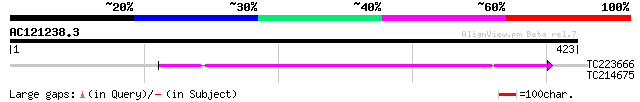
Query= AC121238.3 - phase: 0 /pseudo
(423 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC223666 UP|O81532 (O81532) Mariner transposase, complete 99 5e-21
TC214675 similar to UP|Q6YGT9 (Q6YGT9) Purple acid phosphatase-l... 29 4.6
>TC223666 UP|O81532 (O81532) Mariner transposase, complete
Length = 1278
Score = 98.6 bits (244), Expect = 5e-21
Identities = 101/295 (34%), Positives = 150/295 (50%), Gaps = 1/295 (0%)
Frame = +3
Query: 112 RCFASAFKCTKTTHDR**YESSS*ILFVHVGGK*HSI*AKV*VNAQCYSHR*KVVLCDQK 171
RC ++F+C K T DR + + ILFV ++ *+ V Q YSH *K+VL D++
Sbjct: 342 RCNKASFECHKATTDRGR*KIAVGILFVDA*R--YTA*SNVPKYVQYYSH**KMVLHDEE 515
Query: 172 ISELLPSCR*G*SIPSVQKQKLSNEGHVYSCSG*T*V*LCRS*NMVGKNWSVASS*KSAC 231
I E+L + R G*+ + K K ++ +V++C T +*L + N +NW+
Sbjct: 516 I*EILFAPRRG*TTS*L*K*KFCSKSYVFNCCSSTKI*LGKKCNFFRENWNFPVCHSRTG 695
Query: 232 KKVQCQ*TRRDTRNKTNRFYY*RS*SNVLKRQDFTGRQRKSATRTCIGNNLHSTR*CTVS 291
K +C + + +K+N F *R + + + A R N++HSTR*C S
Sbjct: 696 *KDKC*QSGGNNGDKSNNFDK*RPHKVSFY*ESASSYKGSVAKRRIGVNDIHSTR*CKNS 875
Query: 292 CFC**RRISSSSF*RWI*HPFD*SAT*FS*FKCIGFGIFCSHSSITTKESDKDG**TYSS 351
** + S S RWI*HPF+ S +*FS C+G IF ++ I + S + * T
Sbjct: 876 Y*S**S*VCSGSHTRWI*HPFNVSTS*FSGL*CLGPRIFQCYTVIALQGST*NY*RTCQC 1055
Query: 352 SANFV**IF*CGFKQ-NISYTSVVHD*NHEG*RVQ*I*DSSYEERCHA*TSYTTH 405
S V* + CG Q +IS +V+HD ++E R Q*I S+Y E TTH
Sbjct: 1056SGEVV*KLL-CGEVQFHISLIAVMHDRDNESQRFQ*IYISAYAEGKIRNRRTTTH 1217
>TC214675 similar to UP|Q6YGT9 (Q6YGT9) Purple acid phosphatase-like protein,
partial (34%)
Length = 841
Score = 28.9 bits (63), Expect = 4.6
Identities = 12/24 (50%), Positives = 16/24 (66%)
Frame = +3
Query: 88 T*QSSISSRSIRVYHWTTNEVNKK 111
T*QS I +R+ +YHW N+ KK
Sbjct: 375 T*QSEIKNRTHAIYHWYRNDDGKK 446
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.372 0.162 0.611
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,876,700
Number of Sequences: 63676
Number of extensions: 267094
Number of successful extensions: 2743
Number of sequences better than 10.0: 4
Number of HSP's better than 10.0 without gapping: 1483
Number of HSP's successfully gapped in prelim test: 86
Number of HSP's that attempted gapping in prelim test: 1235
Number of HSP's gapped (non-prelim): 1608
length of query: 423
length of database: 12,639,632
effective HSP length: 100
effective length of query: 323
effective length of database: 6,272,032
effective search space: 2025866336
effective search space used: 2025866336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121238.3


