

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121233.5 - phase: 0
(301 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
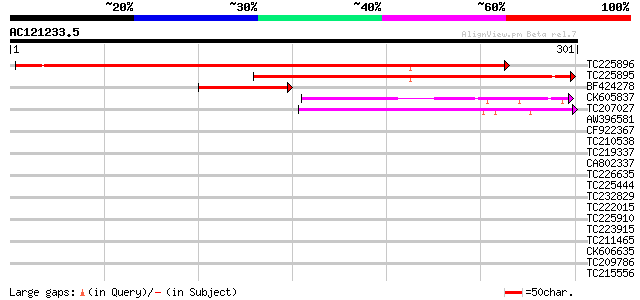
Score E
Sequences producing significant alignments: (bits) Value
TC225896 similar to UP|EBP2_ARATH (Q9LUJ5) Probable rRNA process... 402 e-113
TC225895 similar to UP|EBP2_ARATH (Q9LUJ5) Probable rRNA process... 263 9e-71
BF424278 96 3e-20
CK605837 47 1e-05
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 43 2e-04
AW396581 40 0.002
CF922367 39 0.002
TC210538 similar to UP|H1_MAIZE (P23444) Histone H1, partial (29%) 37 0.008
TC219337 similar to UP|Q9FXI6 (Q9FXI6) F6F9.7 protein, partial (7%) 37 0.014
CA802337 33 0.21
TC226635 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 32 0.27
TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 32 0.35
TC232829 similar to UP|SYN2_HUMAN (Q92777) Synapsin II, partial ... 32 0.35
TC222015 weakly similar to UP|Q9VPS3 (Q9VPS3) CG2839-PA, partial... 32 0.35
TC225910 similar to UP|Q9ASP7 (Q9ASP7) AT3g46780/T6H20_190, part... 32 0.46
TC223915 similar to UP|Q7PZD4 (Q7PZD4) AgCP9387 (Fragment), part... 32 0.46
TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp prote... 31 0.60
CK606635 31 0.60
TC209786 similar to GB|AAS49116.1|44917583|BT011753 At2g28480 {A... 31 0.78
TC215556 weakly similar to GB|AAP37810.1|30725576|BT008451 At3g4... 31 0.78
>TC225896 similar to UP|EBP2_ARATH (Q9LUJ5) Probable rRNA processing protein
EBP2 homolog, partial (59%)
Length = 839
Score = 402 bits (1034), Expect = e-113
Identities = 207/264 (78%), Positives = 226/264 (85%), Gaps = 2/264 (0%)
Frame = +1
Query: 4 SVFFNTSLDLMELPLVNDDTMIDGEAEYLSEEMSPSESESEEDVKLAEPSKTAVNNRDAL 63
+V SL M +P VNDDT+++ E E EEMS SESE +EDVKL EPSK AV NRDAL
Sbjct: 43 AVALTASLG*MAMP-VNDDTLVEAETEDQREEMSSSESEQDEDVKLTEPSKNAVYNRDAL 219
Query: 64 LDKLGDISWPENVEWIHKLSIDIDQEQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIG 123
LDKLGDISWPENVEWIHKLS+D+DQEQEVDVNDDLARELAFYTQALEGT++AFEK QS+G
Sbjct: 220 LDKLGDISWPENVEWIHKLSVDVDQEQEVDVNDDLARELAFYTQALEGTKEAFEKFQSMG 399
Query: 124 LPFLRPADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKE 183
LPFLRPADYYAEMVKTD HM KVK RLLAEK+KM EA+ERRKAREAKRL+KE+Q+QKLKE
Sbjct: 400 LPFLRPADYYAEMVKTDGHMVKVKGRLLAEKRKMEEADERRKAREAKRLAKEIQAQKLKE 579
Query: 184 RAKQKKDDIESVKKWRKQRQQSGFADDG--ADKALDFEDGKVFERSKKKRPGVSPGDRSG 241
RAKQKK+DIESVKKWRKQRQQSG+AD+G AD DFEDGKVFERS KKRP VS DRSG
Sbjct: 580 RAKQKKEDIESVKKWRKQRQQSGYADNGKDADMGFDFEDGKVFERSNKKRPRVSSRDRSG 759
Query: 242 GKAKQAFGKGKKPKKGDVKNSKFG 265
GKAK A GKGKK KK D KNSK G
Sbjct: 760 GKAKLALGKGKKQKKRDGKNSKIG 831
>TC225895 similar to UP|EBP2_ARATH (Q9LUJ5) Probable rRNA processing protein
EBP2 homolog, partial (31%)
Length = 684
Score = 263 bits (671), Expect = 9e-71
Identities = 133/173 (76%), Positives = 150/173 (85%), Gaps = 2/173 (1%)
Frame = +3
Query: 130 ADYYAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKK 189
A AEMVKTD HMEKV RLL+EK+KM +ERRK REAKRL+KE+Q+QKLKERAKQKK
Sbjct: 3 AXXXAEMVKTDGHMEKVXXRLLSEKRKMEXXDERRKXREAKRLAKEIQAQKLKERAKQKK 182
Query: 190 DDIESVKKWRKQRQQSGFADDG--ADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQA 247
+DIESVKKWRKQRQQSG+AD+G AD DFEDGKVFERS KKRPGVSPGDRSGGKAK A
Sbjct: 183 EDIESVKKWRKQRQQSGYADNGKDADMGFDFEDGKVFERSNKKRPGVSPGDRSGGKAKLA 362
Query: 248 FGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKGAVGGSKKRK 300
FGKGKK KK D KNSKFGFGGKKG+KKQNTADTT+DFG ++ KGAV G+K+++
Sbjct: 363 FGKGKKQKKRDGKNSKFGFGGKKGMKKQNTADTTDDFGKYN-KGAVSGNKRKR 518
>BF424278
Length = 417
Score = 95.5 bits (236), Expect = 3e-20
Identities = 45/50 (90%), Positives = 48/50 (96%)
Frame = +1
Query: 101 ELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKSRL 150
ELAFYTQALEGT++AFEKLQS+GLPFLRPADYYAEMVKTD HMEKVK RL
Sbjct: 4 ELAFYTQALEGTKEAFEKLQSMGLPFLRPADYYAEMVKTDGHMEKVKGRL 153
>CK605837
Length = 498
Score = 47.0 bits (110), Expect = 1e-05
Identities = 46/159 (28%), Positives = 72/159 (44%), Gaps = 15/159 (9%)
Frame = +3
Query: 156 KMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKA 215
K + ++++K ++ K+ K+ + +K K++ K+KK + KK +K+ + G
Sbjct: 48 KFI*TKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKGPRGG--------- 200
Query: 216 LDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGK------KPKKGDVKNSKFGFGGK 269
KKKR +PG GK+ + G+GK PKKG K K G GGK
Sbjct: 201 ----------TPKKKRKFGAPGGAKKGKSPKG-GEGKVGEFKRAPKKGGGKRGKRGSGGK 347
Query: 270 -----KGLKKQNTADTTNDFGGFSKKGA----VGGSKKR 299
K KK+ + + GG K GA GG+K R
Sbjct: 348 MVTRQKFQKKKKRKEKRENRGG-PKGGANPNKWGGAKGR 461
Score = 32.0 bits (71), Expect = 0.35
Identities = 33/145 (22%), Positives = 60/145 (40%), Gaps = 26/145 (17%)
Frame = +1
Query: 153 EKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQR-------QQS 205
+K+K + ++++K ++ K+ K+ + +K K++ K+KK + + +K+ Q+
Sbjct: 70 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKVPGGEPRKKKGNSGPPGGQKK 249
Query: 206 GFADDGADKALDFEDGKV------------------FERSKKKRPGVSPGDRSGG-KAKQ 246
G A G + + G + SKKKR G G GG + Q
Sbjct: 250 GKAPKGGKEKWENSKGPPKKGGENGAKGVPGGKW*PAKNSKKKRSGKKKGKTGGGQRGGQ 429
Query: 247 AFGKGKKPKKGDVKNSKFGFGGKKG 271
G P+ G GGK+G
Sbjct: 430 TQINGGGPRAGFQ-------GGKRG 483
Score = 31.2 bits (69), Expect = 0.60
Identities = 28/89 (31%), Positives = 38/89 (42%)
Frame = +1
Query: 211 GADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKK 270
G D A F+ K ++ KKK+ + K K+ K KK KK K K GG+
Sbjct: 34 GIDMASSFKQKKKKKKKKKKKKKKKKKKKKKKKKKK---KKKKKKKKKKKKKKKVPGGEP 204
Query: 271 GLKKQNTADTTNDFGGFSKKGAVGGSKKR 299
KK N+ GG K A G K++
Sbjct: 205 RKKKGNSGPP----GGQKKGKAPKGGKEK 279
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 42.7 bits (99), Expect = 2e-04
Identities = 40/161 (24%), Positives = 69/161 (42%), Gaps = 13/161 (8%)
Frame = +3
Query: 154 KQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGAD 213
++K + ++ +K +E K + +K+K + +DD E +K +K+++ D
Sbjct: 15 EEKKEKDKKEKKKKENKDKGDKTDVEKVKGKENDGEDDEEKKEKKKKEKKDKDGKTDAEK 194
Query: 214 KALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGK--GKKPKK------GDVKNSKFG 265
+DGK E K+K+ D K K+ GK G K KK GD + K
Sbjct: 195 IKKKEDDGKDDEEKKEKKKKYDKIDAXKVKGKEDDGKDEGNKEKKDKEKGDGDGEEKKEK 374
Query: 266 FGGKKGLKKQ-----NTADTTNDFGGFSKKGAVGGSKKRKR 301
+K KK+ DT + G + G+KK+K+
Sbjct: 375 KDKEKEKKKEKKDKDEETDTLKEKGKNDEGEDDEGNKKKKK 497
Score = 32.0 bits (71), Expect = 0.35
Identities = 56/263 (21%), Positives = 95/263 (35%), Gaps = 27/263 (10%)
Frame = +3
Query: 34 EEMSPSESESEEDVKLAEPSKTAVNNRDALLD-----KLGDISWPENVEWIHKLSIDIDQ 88
E++ E++ E+D + E K ++D D K D + + K D
Sbjct: 90 EKVKGKENDGEDDEEKKEKKKKEKKDKDGKTDAEKIKKKEDDGKDDEEKKEKKKKYDKID 269
Query: 89 EQEVDVNDDLARELAFYTQALEGTRQAFEKLQSIGLPFLRPADYYAEMVKTDSHMEKVKS 148
+V +D ++ EG ++ +K + G D + K D EK K
Sbjct: 270 AXKVKGKEDDGKD--------EGNKEKKDKEKGDG-------DGEEKKEKKDKEKEKKKE 404
Query: 149 R--------LLAEKQKMVEAEE----RRKAREAKRLSKEVQSQKLKERAKQKKD------ 190
+ L EK K E E+ ++K ++ K K+ + +K + +K+D
Sbjct: 405 KKDKDEETDTLKEKGKNDEGEDDEGNKKKKKDKKEKEKDHKKEKKDKEEGEKEDSKVEVS 584
Query: 191 ----DIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQ 246
DIE +KK ++ D G D + ++ K E KK K K+
Sbjct: 585 VRDIDIEEIKKEGEKE------DKGKDGGKEVKEKKKKEDKDKKE-----------KKKK 713
Query: 247 AFGKGKKPKKGDVKNSKFGFGGK 269
GK K +K GK
Sbjct: 714 VTGKDKTKDLSTLKQKLEKINGK 782
>AW396581
Length = 401
Score = 39.7 bits (91), Expect = 0.002
Identities = 31/114 (27%), Positives = 48/114 (41%), Gaps = 4/114 (3%)
Frame = -1
Query: 154 KQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGAD 213
K K + E+ + +E K+ K+ + K + K+D E KK +K + A+
Sbjct: 371 KVKEHDGEDAEEKKEKKKKEKKDKDDKTDVEKVKGKEDKEKKKKEKKVKDDKTDAEKVKG 192
Query: 214 KALDFEDGKVFERSKKKRPGVSPG----DRSGGKAKQAFGKGKKPKKGDVKNSK 263
K D ED + E KKK+ ++ GK +GKK KK K K
Sbjct: 191 KEDDGEDNEEKEEKKKKKKKEKDDKIDVEKVKGKEDDGEDEGKKEKKKKEKKDK 30
>CF922367
Length = 463
Score = 39.3 bits (90), Expect = 0.002
Identities = 32/108 (29%), Positives = 53/108 (48%)
Frame = -2
Query: 168 EAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERS 227
EAKR KE+ +Q++ E K++K + +S K RKQ+++ G + ++G R
Sbjct: 396 EAKRKRKEIPNQRVGESKKKEKGNSQS-KNGRKQKEEKGNSQS--------KNG----RK 256
Query: 228 KKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQ 275
+K++ G S +SG K K+ G + K K K G KK+
Sbjct: 255 QKEKKGNSQ-SKSGRKQKEKKGNSQSKSGRKQKEKKENSQSKNGRKKK 115
Score = 36.2 bits (82), Expect = 0.019
Identities = 35/118 (29%), Positives = 59/118 (49%), Gaps = 9/118 (7%)
Frame = -3
Query: 155 QKMVEAEERRKARE---AKRLSKEVQSQKLKERAKQKKDDI-----ESVKKWRKQRQQSG 206
+K E E + +E AKR KE+ +Q++ E K++K+ ES KK ++ Q
Sbjct: 398 RKQKERERKFPIKEWEKAKRKRKEIPNQRMGESKKKRKEIPNQRMGESKKKRKETPNQR- 222
Query: 207 FADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKK-PKKGDVKNSK 263
+ K + + +V E SK+KR + P R G + K+ +GKK P +G +N +
Sbjct: 221 -VGESKKKRKEIPNQRVGE-SKRKRKKI-PNQRMGERKKKGRKEGKKAPDQGSKENRR 57
>TC210538 similar to UP|H1_MAIZE (P23444) Histone H1, partial (29%)
Length = 643
Score = 37.4 bits (85), Expect = 0.008
Identities = 33/128 (25%), Positives = 59/128 (45%), Gaps = 7/128 (5%)
Frame = +2
Query: 116 FEKLQSIGLPFLRPADYYAEM-----VKTDSHMEKVKSRLLAEKQKMVEAEERRK--ARE 168
F KL S+ L L ++ ++ + + + E K+ KQK+ + ++++K +
Sbjct: 125 FRKLLSVQLKKLVKSEKLYKVKNSYKLSSATDKETQKNTETEPKQKVEKKKKKKKKVGEK 304
Query: 169 AKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSK 228
KRLS+ + LK++A KKD + K++ S G K K E K
Sbjct: 305 IKRLSQVKTPETLKKKAPTKKDAAAT-----KKKSASAVEGSGGGKVKCLSQVKTPEALK 469
Query: 229 KKRPGVSP 236
KK+P ++P
Sbjct: 470 KKKPNLTP 493
>TC219337 similar to UP|Q9FXI6 (Q9FXI6) F6F9.7 protein, partial (7%)
Length = 740
Score = 36.6 bits (83), Expect = 0.014
Identities = 30/101 (29%), Positives = 48/101 (46%)
Frame = +1
Query: 182 KERAKQKKDDIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSG 241
K+ AK+ ++ KK RKQ + S +D+ A+++ + + ++ ++ KRP R
Sbjct: 187 KQAAKKGAKGADNSKK-RKQSKDSSDSDEDAEESEEDSEDQINGEAEAKRP------RGS 345
Query: 242 GKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTN 282
GK G+GK PKK K G G G + TA N
Sbjct: 346 GK-----GRGKAPKKSGAK----GKGSVSGRGRGGTAANKN 441
>CA802337
Length = 381
Score = 32.7 bits (73), Expect = 0.21
Identities = 37/142 (26%), Positives = 64/142 (45%)
Frame = +2
Query: 158 VEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADDGADKALD 217
V ++++K ++ K+ K+ + +K K++ K+KK + KK +K++++
Sbjct: 32 VTVKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK------------- 172
Query: 218 FEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNT 277
K ++ KKK+ G G K+ F +G+ KK F +GGKK KK
Sbjct: 173 ----KKKKKKKKKKXG-------GV*KKKVFXRGEWKKKF------F*WGGKK--KK--- 286
Query: 278 ADTTNDFGGFSKKGAVGGSKKR 299
FG KG G KK+
Sbjct: 287 ---FFFFGKKXXKGKKMGGKKK 343
>TC226635 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), complete
Length = 2039
Score = 32.3 bits (72), Expect = 0.27
Identities = 35/109 (32%), Positives = 53/109 (48%), Gaps = 7/109 (6%)
Frame = +3
Query: 139 TDSHMEKVKSRL-LAEKQKMVEAEERR------KAREAKRLSKEVQSQKLKERAKQKKDD 191
T+ ME+ K +L + E +K+V + KA EAKR ++ +Q L AKQ K +
Sbjct: 1443 TELKMERQKKKLQIEELEKIVRLKNAEADMFQLKADEAKREAERLQMIAL---AKQDKSE 1613
Query: 192 IESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRS 240
E + KQR + A+K +E K+ E S+ + G S GD S
Sbjct: 1614 EEFTSNYLKQR----LNEAEAEKQYLYEKIKLQESSRASQ-GSSGGDPS 1745
>TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 750
Score = 32.0 bits (71), Expect = 0.35
Identities = 22/74 (29%), Positives = 36/74 (47%), Gaps = 10/74 (13%)
Frame = +2
Query: 133 YAEMVKTDSHMEKVKSRLLAEKQKM----------VEAEERRKAREAKRLSKEVQSQKLK 182
Y + K D E VK R A K+ V ++R + E + ++E Q +++K
Sbjct: 212 YRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDAAREAQLREIK 391
Query: 183 ERAKQKKDDIESVK 196
ER K+ KDD ++ K
Sbjct: 392 ERIKKTKDDKKAKK 433
>TC232829 similar to UP|SYN2_HUMAN (Q92777) Synapsin II, partial (3%)
Length = 425
Score = 32.0 bits (71), Expect = 0.35
Identities = 17/54 (31%), Positives = 25/54 (45%)
Frame = -2
Query: 218 FEDGKVFERSKKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFGFGGKKG 271
F++G + KK + G S K +++FG G + K GFGGK G
Sbjct: 367 FDEGDAGRKRKKGKEGGSGKKEKRMKGEKSFGSGGGKSGSKFGSGKKGFGGKAG 206
>TC222015 weakly similar to UP|Q9VPS3 (Q9VPS3) CG2839-PA, partial (4%)
Length = 796
Score = 32.0 bits (71), Expect = 0.35
Identities = 34/134 (25%), Positives = 60/134 (44%), Gaps = 22/134 (16%)
Frame = +3
Query: 168 EAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQR-QQSGFADDGADKA----------- 215
+ K +KE + ++ +ERAK++KD E ++ KQR ++ G + G D+A
Sbjct: 180 QLKEEAKESERKRKEERAKKEKDREERERRKGKQRKEKEGGRERGKDEAHKKDKADSDSM 359
Query: 216 --LDFEDGKVFERS--------KKKRPGVSPGDRSGGKAKQAFGKGKKPKKGDVKNSKFG 265
+ + K +RS KK++ V D+ K K++ G G KK + G
Sbjct: 360 ELTEIQTSKENKRSEDDNRKQRKKRQSPVYEMDKE--KTKKSHGHGSDSKKS--RRHASG 527
Query: 266 FGGKKGLKKQNTAD 279
+G K++ D
Sbjct: 528 HESDEGRHKRHKRD 569
>TC225910 similar to UP|Q9ASP7 (Q9ASP7) AT3g46780/T6H20_190, partial (32%)
Length = 1272
Score = 31.6 bits (70), Expect = 0.46
Identities = 38/170 (22%), Positives = 69/170 (40%), Gaps = 9/170 (5%)
Frame = +2
Query: 133 YAEMVKTDSHMEKVKSRLLAEKQKMVEAEERRKAREAKRLS-KEVQSQKLKERAKQKKD- 190
YAEM + E+ +R+ AEK + ++ E KRLS +E ++ L + A++K +
Sbjct: 479 YAEMQEKTKAEEE--ARVAAEKAREASESAKKLEEEVKRLSQQETRAASLAQEAQEKAEA 652
Query: 191 ---DIESVKKWRKQRQQSGFADDGADKALDFEDGKVFERSKKKRPGVSPGDRSGGKAKQA 247
+E++ K D GA + + ++ +K P + QA
Sbjct: 653 GGASVENLLNKAK--------DFGAGFSWEKLSSQITTSIQKPDEDEKPKVQLATVRGQA 808
Query: 248 FGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTND----FGGFSKKGAV 293
+ P K VK + K ++K +T + FGG K+ +
Sbjct: 809 KARNLAPNKAVVKQTPQRSAAKPKVEKPKQTETPKEVRKVFGGLFKQETI 958
>TC223915 similar to UP|Q7PZD4 (Q7PZD4) AgCP9387 (Fragment), partial (7%)
Length = 643
Score = 31.6 bits (70), Expect = 0.46
Identities = 19/64 (29%), Positives = 33/64 (50%)
Frame = +3
Query: 151 LAEKQKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADD 210
L ++++ AEER + AKR QK K R K+KK + + ++ +++ SG D
Sbjct: 246 LRREERLKAAEERTAKKRAKR-------QKKKLRKKEKKSKLNAGEQLQEKEDSSGDEDS 404
Query: 211 GADK 214
D+
Sbjct: 405 DNDE 416
>TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (15%)
Length = 593
Score = 31.2 bits (69), Expect = 0.60
Identities = 13/26 (50%), Positives = 21/26 (80%)
Frame = -2
Query: 179 QKLKERAKQKKDDIESVKKWRKQRQQ 204
QK KER K KKD++ S+K+ +K+R++
Sbjct: 109 QKGKERKKDKKDELISIKERKKERER 32
>CK606635
Length = 528
Score = 31.2 bits (69), Expect = 0.60
Identities = 23/65 (35%), Positives = 30/65 (45%), Gaps = 6/65 (9%)
Frame = +1
Query: 243 KAKQAFGKGKKPKKGDVKNSKFGFGGKKGLKKQNTADTTNDFGGFSKKG------AVGGS 296
K K+ G G+KPKK KNS+ G KK K +F G + G V G
Sbjct: 34 KKKKRGGGGEKPKKKKKKNSQRGVEKKKKKKNFFPPKKKKNFRGKPRGGEPKREKPVFGK 213
Query: 297 KKRKR 301
KK+K+
Sbjct: 214 KKKKK 228
>TC209786 similar to GB|AAS49116.1|44917583|BT011753 At2g28480 {Arabidopsis
thaliana;} , partial (64%)
Length = 1160
Score = 30.8 bits (68), Expect = 0.78
Identities = 26/96 (27%), Positives = 44/96 (45%), Gaps = 9/96 (9%)
Frame = +2
Query: 155 QKMVEAEERRKAREAKRLSKEVQSQKLKERAKQKKDDIESVKKWRKQRQQSGFADD---- 210
Q+ E ++ K KRL + + +K K RA +KD K +K++Q+ A++
Sbjct: 200 QEEEEPPKKDKWTTKKRLKMQRKREKEKRRAANRKDPRRLGVKGKKKKQRFASAEERIKY 379
Query: 211 -----GADKALDFEDGKVFERSKKKRPGVSPGDRSG 241
+AL E K +E K + P V P + +G
Sbjct: 380 KIEKARIKEALLIERLK*YEVPKAQGPVVKPDELTG 487
>TC215556 weakly similar to GB|AAP37810.1|30725576|BT008451 At3g44190
{Arabidopsis thaliana;} , partial (89%)
Length = 1238
Score = 30.8 bits (68), Expect = 0.78
Identities = 29/122 (23%), Positives = 52/122 (41%), Gaps = 9/122 (7%)
Frame = +2
Query: 40 ESESEEDVKLAEPSKTAVNNRDALLDKLGDISWPENVEWIHKLSIDIDQEQEVDVNDDLA 99
E +E V + T V+ LL+ +G + + ++W+ ID+ EQ VD++
Sbjct: 494 ELAAEIAVDFPDKKVTIVHKGTRLLEYIGTKASSKTLKWLKSKKIDVKLEQSVDLSSSSE 673
Query: 100 RELAFYTQALEGTRQAFEKL---QSIGLPFLRPA----DYYAE-MVKTDSHME-KVKSRL 150
+ T E + L + +G ++R D A+ +K D H+ K KS +
Sbjct: 674 ENKTYQTSNGETIKADLHFLCTGKPLGSTWIRETLLKNDLDADGRIKVDEHLRVKGKSNI 853
Query: 151 LA 152
A
Sbjct: 854 FA 859
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.130 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,727,723
Number of Sequences: 63676
Number of extensions: 85351
Number of successful extensions: 664
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 642
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 653
length of query: 301
length of database: 12,639,632
effective HSP length: 97
effective length of query: 204
effective length of database: 6,463,060
effective search space: 1318464240
effective search space used: 1318464240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC121233.5


