

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
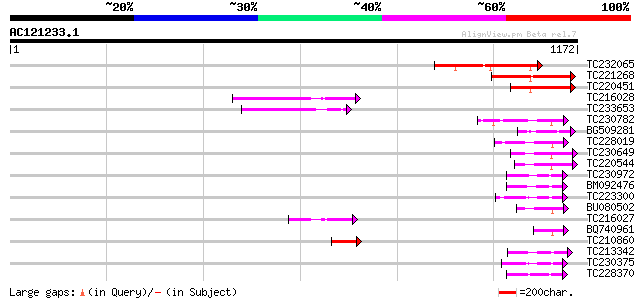
Query= AC121233.1 - phase: 0 /pseudo
(1172 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC232065 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, parti... 271 1e-72
TC221268 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, parti... 249 4e-66
TC220451 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, parti... 232 8e-61
TC216028 homologue to UP|Q9ZSQ0 (Q9ZSQ0) Ethylene response senso... 136 6e-32
TC233653 similar to UP|ETR1_MALDO (O81122) Ethylene receptor , ... 102 1e-21
TC230782 similar to UP|Q9C5U1 (Q9C5U1) Histidine kinase, partial... 86 7e-17
BG509281 similar to GP|14023997|dbj Hybrid sensory histidine kin... 76 1e-13
TC228019 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b,... 73 6e-13
TC230649 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b,... 72 1e-12
TC220544 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b,... 71 2e-12
TC230972 similar to UP|Q9XET9 (Q9XET9) Ethylene receptor homolog... 70 4e-12
BM092476 70 5e-12
TC223300 similar to UP|Q949H5 (Q949H5) Ethylene receptor, partia... 64 5e-10
BU080502 similar to GP|13537196|d histidine kinase {Arabidopsis ... 62 1e-09
TC216027 homologue to UP|Q9ZSQ0 (Q9ZSQ0) Ethylene response senso... 62 1e-09
BQ740961 61 3e-09
TC210860 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b,... 60 7e-09
TC213342 weakly similar to UP|O82445 (O82445) Response regulator... 57 4e-08
TC230375 similar to UP|Q9SSY4 (Q9SSY4) Ethylene receptor CS-ETR2... 57 6e-08
TC228370 similar to UP|Q9AXD7 (Q9AXD7) Response regulator protei... 56 1e-07
>TC232065 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, partial (5%)
Length = 956
Score = 271 bits (693), Expect = 1e-72
Identities = 161/269 (59%), Positives = 187/269 (68%), Gaps = 46/269 (17%)
Frame = -2
Query: 878 KTELRKKGHILTVNKPLYKAKMIHILEDVIKERNVEVQKKNM-------KEDDLHESLEI 930
K +L +KGH LTVNKPLYK KMIHILE +IK+RN E+QKKNM KE +LHESLEI
Sbjct: 955 KMDLCRKGHTLTVNKPLYKTKMIHILESIIKDRNEELQKKNMTTLRATVKEGNLHESLEI 776
Query: 931 EYTHHCDVASSDGSDISEIGSSNLVTANGEKQREEVVRINPSSLYQKSNCLLGLSNGYME 990
+YT CDVASSDGSDIS G SN V+A G+KQRE+VVR +PSS +Q +N L+GL+N ME
Sbjct: 775 DYT-QCDVASSDGSDISXTGGSNPVSARGDKQREKVVRSDPSSQHQINN-LVGLTNECME 602
Query: 991 H-----------------------------KEAFISTRAKGEDSKGGETSRVSGSSKAMN 1021
K+A +T A+ DS+ GET + S SS+A++
Sbjct: 601 DDNHRKEELCQSSLNSNDVTANASPKSSSTKQASFATGARDGDSEYGETRKASSSSRAVS 422
Query: 1022 GKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGV------- 1074
GKKSLEGLRILLAEDT VIQRVATIMLEKMGA VVAVGDG+QAVDALNGMPGV
Sbjct: 421 GKKSLEGLRILLAEDTPVIQRVATIMLEKMGAIVVAVGDGRQAVDALNGMPGVEDCIRES 242
Query: 1075 ---ERNTITSQTEILSFPSYDLILMDCQM 1100
ERNT +SQTEIL P YDLILMDCQ+
Sbjct: 241 LLKERNTRSSQTEILGCPPYDLILMDCQV 155
>TC221268 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, partial (10%)
Length = 978
Score = 249 bits (637), Expect = 4e-66
Identities = 140/184 (76%), Positives = 152/184 (82%), Gaps = 11/184 (5%)
Frame = +1
Query: 996 ISTRAKGEDSKGGETSRVSGSSKAM-NGKKSLEGLRILLAEDTAVIQRVATIMLEKMGAT 1054
IST + EDS+ G+T+RV+ SSKA+ +GKKSLEGL+ILLAEDT V+QRVATIMLEKMGA
Sbjct: 205 ISTEDQDEDSECGDTNRVTSSSKAVVDGKKSLEGLKILLAEDTPVLQRVATIMLEKMGAD 384
Query: 1055 VVAVGDGQQAVDALNGMPGVE----------RNTITSQTEILSFPSYDLILMDCQMPKMD 1104
VVAVGDGQQAVDALN M E RNT SQTEI + YDLILMDCQMPKMD
Sbjct: 385 VVAVGDGQQAVDALNCMFTAEDCRRESLQKERNT-RSQTEISTCRPYDLILMDCQMPKMD 561
Query: 1105 GYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILS 1164
GYEATK IRKSE+GTS HIPIVALTAHAMSCDEAKC EVGMDAYLTKPIDFK+M STILS
Sbjct: 562 GYEATKAIRKSEVGTSRHIPIVALTAHAMSCDEAKCREVGMDAYLTKPIDFKMMVSTILS 741
Query: 1165 LTKR 1168
LTKR
Sbjct: 742 LTKR 753
>TC220451 similar to UP|Q9SXL4 (Q9SXL4) Histidine kinase 1, partial (9%)
Length = 501
Score = 232 bits (591), Expect = 8e-61
Identities = 121/143 (84%), Positives = 124/143 (86%), Gaps = 10/143 (6%)
Frame = +2
Query: 1036 DTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVE----------RNTITSQTEI 1085
DT VIQRVATIMLEKMGA VVAVGDGQQAVDALNGMPGVE RNT +SQTEI
Sbjct: 2 DTPVIQRVATIMLEKMGAVVVAVGDGQQAVDALNGMPGVEDCIRESLLKERNTRSSQTEI 181
Query: 1086 LSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGM 1145
L P YDLILMDCQMPKMDGYEATK IRKSE+GT HIPIVALTAHAMSCDEAKCLEVGM
Sbjct: 182 LGCPPYDLILMDCQMPKMDGYEATKAIRKSEVGTDLHIPIVALTAHAMSCDEAKCLEVGM 361
Query: 1146 DAYLTKPIDFKLMESTILSLTKR 1168
DAYLTKPIDFKLMESTILSLT+R
Sbjct: 362 DAYLTKPIDFKLMESTILSLTRR 430
>TC216028 homologue to UP|Q9ZSQ0 (Q9ZSQ0) Ethylene response sensor, partial
(78%)
Length = 1745
Score = 136 bits (342), Expect = 6e-32
Identities = 89/264 (33%), Positives = 134/264 (50%)
Frame = +2
Query: 461 ARRKAEASSNYKSQFLANMSHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTA 520
ARR+AE + + ++ FLA M+HE+RTPM A+I L +LL + LT EQ + + K S
Sbjct: 572 ARREAEMAIHARNDFLAVMNHEMRTPMHAIIALSSLLLETE-LTPEQRVMIETVLKSSNV 748
Query: 521 LLHLLNNILDISKVESGKLVLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPK 580
L L+N++LD+S++E G L LE +F+ L +V++ + I L LS D+P
Sbjct: 749 LATLINDVLDLSRLEDGSLELEMGKFNLHGVLGEIVELIKPIASVKKLPITLILSPDLPT 928
Query: 581 LVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKT 640
GD R+ Q N++ N++KFT G++ +R P S D
Sbjct: 929 HAIGDEKRLTQTLLNVVGNAVKFTKEGYVSIRVSVAKPESLQD----------------- 1057
Query: 641 KMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGL 700
+ E + ++ D + +V D+GCGI P + +F F Q+ R G GL
Sbjct: 1058-WRPPEFY---PASSDGHFYIRVQVKDSGCGIPPQEIPHLFTKFAQSRSGPARPSSGAGL 1225
Query: 701 GLCIVRSLVNKMGGEIKIVKKEGP 724
GL I + VN MGG I I + EGP
Sbjct: 1226GLAICKRFVNLMGGHIWI-ESEGP 1294
>TC233653 similar to UP|ETR1_MALDO (O81122) Ethylene receptor , partial
(29%)
Length = 663
Score = 102 bits (253), Expect = 1e-21
Identities = 71/227 (31%), Positives = 107/227 (46%)
Frame = +1
Query: 480 SHELRTPMAAVIGLLDILLSDDRLTNEQCATVTQIRKCSTALLHLLNNILDISKVESGKL 539
+HE+RTPM AVI L +L D LT EQ V I K S L L+N++LD+S++E G L
Sbjct: 1 NHEMRTPMHAVIALSSLLQETD-LTAEQRLMVETILKSSNLLATLINDVLDLSRLEDGSL 177
Query: 540 VLEEAEFDFGRELEGLVDMFSVQCINHNVEIILDLSDDMPKLVRGDSARVVQIFANLINN 599
LE A F+ ++++ + + ++ D+P GD R++Q N++ N
Sbjct: 178 QLEAATFNLHSLFREVLNLIKPVASVKKLSLTSHVASDLPMYAIGDEKRLMQTILNVVGN 357
Query: 600 SIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGCSRKTKMKQHENHSKKASNIDNKM 659
++KF+ G I + + P S+ D+ P DN
Sbjct: 358 AVKFSKEGCISITAFVAKPESFRDARIPDFLPVP---------------------SDNHF 474
Query: 660 ILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVR 706
L +V D+G GI+P +F F Q + S TR G+GLGL I R
Sbjct: 475 YLRVQVKDSGSGINPQDIPKLFTKFAQ-NQSLTRNPAGSGLGLXICR 612
>TC230782 similar to UP|Q9C5U1 (Q9C5U1) Histidine kinase, partial (14%)
Length = 869
Score = 86.3 bits (212), Expect = 7e-17
Identities = 63/207 (30%), Positives = 99/207 (47%), Gaps = 19/207 (9%)
Frame = +1
Query: 967 VRINPSSLYQKSNCLLGLSNGYMEHKEAFIST------RAKGEDSKGGETSRVSGSSKAM 1020
+ +N SS + K++ LG+ N + K S RA G +KG + +++
Sbjct: 277 ILVNSSSSF-KASVNLGVHNPTVITKPLRASMLAASLQRAMGVQNKGAPHREL----QSL 441
Query: 1021 NGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTIT 1080
+ + L G +IL+ +D V + VA L+K GA VV V G+ A+ +L
Sbjct: 442 SLRHLLRGRKILIVDDNGVNRAVAAGALKKYGADVVCVSSGKDAISSLKPPH-------- 597
Query: 1081 SQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIG-------------TSFHIPIVA 1127
+D MD QMP+MDG+EATK IR+ E T++H+PI+A
Sbjct: 598 ---------QFDACFMDIQMPEMDGFEATKRIREMEDSVNREVSMDDFENITNWHVPILA 750
Query: 1128 LTAHAMSCDEAKCLEVGMDAYLTKPID 1154
+TA + +CL GMD Y++KP +
Sbjct: 751 MTADVIQATHEECLRCGMDGYVSKPFE 831
>BG509281 similar to GP|14023997|dbj Hybrid sensory histidine kinase
{Mesorhizobium loti}, partial (6%)
Length = 420
Score = 75.9 bits (185), Expect = 1e-13
Identities = 48/120 (40%), Positives = 68/120 (56%)
Frame = +1
Query: 1050 KMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFPSYDLILMDCQMPKMDGYEAT 1109
++GA+V+ +G+QAV + G+ RN+ S D ILMDCQMP MDGYEAT
Sbjct: 1 RLGASVMECENGEQAVQTVE--EGLTRNS--------SNRPCDFILMDCQMPVMDGYEAT 150
Query: 1110 KEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPIDFKLMESTILSLTKRE 1169
+ IR+ E HIPI ALTA+ + +E GMD +L KPI+ + + I + +E
Sbjct: 151 RRIREIEKSHGVHIPIFALTANT-GKEAILSIEAGMDDHLIKPINKEALLKAIKRIYTKE 327
>TC228019 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, partial (21%)
Length = 1344
Score = 73.2 bits (178), Expect = 6e-13
Identities = 50/168 (29%), Positives = 81/168 (47%), Gaps = 15/168 (8%)
Frame = +3
Query: 1002 GEDSKGGETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDG 1061
G+ + G+ +GS+ + L G +IL+ +D V +RVA L+ GA V G
Sbjct: 429 GKKRQHGKDMNPNGSTFV---RSLLCGKKILVVDDNVVNRRVAAGALKNFGADVTCAESG 599
Query: 1062 QQAVDALNGMPGVERNTITSQTEILSFP-SYDLILMDCQMPKMDGYEATKEIRKSEIGTS 1120
+ A+ E+L P ++D MD QMP+MDG++AT+ IR E +
Sbjct: 600 KTAL------------------EMLQLPHNFDACFMDIQMPEMDGFQATQRIRMMETKAN 725
Query: 1121 --------------FHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPID 1154
+HIPI+A+TA + +C++ GMD Y++KP +
Sbjct: 726 EQQMNGEGNGWKDKYHIPILAMTADVIHATYDECVKYGMDGYVSKPFE 869
>TC230649 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, partial (12%)
Length = 762
Score = 72.4 bits (176), Expect = 1e-12
Identities = 45/146 (30%), Positives = 71/146 (47%), Gaps = 8/146 (5%)
Frame = +2
Query: 1035 EDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFP-SYDL 1093
+ V RV L+K GA V G+ A+ E+L P ++D
Sbjct: 5 DXNGVXXRVXAGALKKFGADVXCAXXGKVAL------------------EMLQLPHNFDA 130
Query: 1094 ILMDCQMPKMDGYEATKEIRKSEI-------GTSFHIPIVALTAHAMSCDEAKCLEVGMD 1146
MD QMP+MDG+EAT IR E G+ +H+PI+A+TA + KC++ GMD
Sbjct: 131 CFMDIQMPEMDGFEATSRIRMMESKANEEMNGSEWHVPILAMTADVILATYDKCVKCGMD 310
Query: 1147 AYLTKPIDFKLMESTILSLTKRETLT 1172
Y++KP + + + + K +T++
Sbjct: 311 GYVSKPFEEENLYQEVAKFFKSKTIS 388
>TC220544 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, partial (10%)
Length = 772
Score = 71.2 bits (173), Expect = 2e-12
Identities = 44/139 (31%), Positives = 68/139 (48%), Gaps = 9/139 (6%)
Frame = +3
Query: 1043 VATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFP-SYDLILMDCQMP 1101
VA L+K GA V G+ A+ E+L P ++D MD QMP
Sbjct: 3 VAAGALKKFGAXVKCAESGKAAL------------------EMLQLPHNFDACFMDIQMP 128
Query: 1102 KMDGYEATKEIRKSEI--------GTSFHIPIVALTAHAMSCDEAKCLEVGMDAYLTKPI 1153
+MDG+EAT IR E G +H+PI+A+TA + KC++ GMD Y++KP
Sbjct: 129 EMDGFEATSRIRMMESKANEEMNNGNEWHVPILAMTADVIHATYDKCMKCGMDGYVSKPF 308
Query: 1154 DFKLMESTILSLTKRETLT 1172
+ + + + K +T++
Sbjct: 309 EEENLYQEVAKFFKSKTMS 365
>TC230972 similar to UP|Q9XET9 (Q9XET9) Ethylene receptor homolog, partial
(21%)
Length = 958
Score = 70.5 bits (171), Expect = 4e-12
Identities = 46/126 (36%), Positives = 66/126 (51%)
Frame = +3
Query: 1028 GLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILS 1087
GL++LLAED V + V +LEK+G V+AV G + + A++G
Sbjct: 363 GLKVLLAEDDGVNRTVTKKLLEKLGCQVIAVSSGFECLSAVSGAGN-------------- 500
Query: 1088 FPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDA 1147
S+ +IL+D MP+MDG+E K IRK S+ + I+AL A KCL GM+
Sbjct: 501 --SFRIILLDLHMPEMDGFELAKRIRKFH-SRSWPL-IIALITSAEEHVREKCLLAGMNG 668
Query: 1148 YLTKPI 1153
+ KPI
Sbjct: 669 LIQKPI 686
>BM092476
Length = 421
Score = 70.1 bits (170), Expect = 5e-12
Identities = 44/127 (34%), Positives = 70/127 (54%), Gaps = 1/127 (0%)
Frame = +1
Query: 1028 GLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILS 1087
GL+++LA+D V + V +LEK+G V AV G + + A++ S
Sbjct: 1 GLKVVLADDDDVNRTVTKKLLEKLGCQVTAVSSGFECLGAISA----------------S 132
Query: 1088 FPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIP-IVALTAHAMSCDEAKCLEVGMD 1146
S+ +I++D MP+MDG+E + IRK + S + P I+A TA A + +CL+VGM+
Sbjct: 133 GNSFKIIMLDLHMPEMDGFEVARRIRKFQ---SHNWPLIIAFTASAEEHIKERCLQVGMN 303
Query: 1147 AYLTKPI 1153
+ KPI
Sbjct: 304 GLIRKPI 324
>TC223300 similar to UP|Q949H5 (Q949H5) Ethylene receptor, partial (25%)
Length = 711
Score = 63.5 bits (153), Expect = 5e-10
Identities = 46/151 (30%), Positives = 77/151 (50%), Gaps = 1/151 (0%)
Frame = +2
Query: 1004 DSKGGETSRVSGSSKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQ 1063
+S+ GE S S N GL++LLA++ V + V +L+K+G V +V G +
Sbjct: 155 NSEPGENSETS------NSNSFFRGLQVLLADNDDVNRAVTQKLLQKLGCVVTSVSSGLE 316
Query: 1064 AVDALNGMPGVERNTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHI 1123
++ + G G S+ +IL+D MP++DG+E IRK S +
Sbjct: 317 CLNVI-GPAG---------------SSFQVILLDLHMPELDGFEVATRIRKFR---SRNW 439
Query: 1124 PIV-ALTAHAMSCDEAKCLEVGMDAYLTKPI 1153
P++ ALTA E +C+++GM+A + KP+
Sbjct: 440 PVIAALTASTEDLWE-RCMQIGMNAVIRKPV 529
Score = 31.6 bits (70), Expect = 2.1
Identities = 35/128 (27%), Positives = 60/128 (46%), Gaps = 10/128 (7%)
Frame = +2
Query: 700 LGLCIVRSLVNKMGGEIKIVKK-EGPGTLMRLYMQL----TAPVDATEQHCQVDFANNN- 753
L I + ++ M G I +V +G +M L+++ + V +E + +N+N
Sbjct: 20 LSFSICKRIIQLMQGNIWLVPNAQGFPQVMALFLRFQLRRSIAVSNSEPGENSETSNSNS 199
Query: 754 ----LMVLLALHGNMSRLITSKWLQKNGVLTMEASEWNGLTQILKELFEAKTSTHNNDFD 809
L VLLA + +++R +T K LQK G + S +GL + L + A +S D
Sbjct: 200 FFRGLQVLLADNDDVNRAVTQKLLQKLGCVVTSVS--SGL-ECLNVIGPAGSSFQVILLD 370
Query: 810 THFPAPEG 817
H P +G
Sbjct: 371 LHMPELDG 394
>BU080502 similar to GP|13537196|d histidine kinase {Arabidopsis thaliana},
partial (10%)
Length = 429
Score = 62.0 bits (149), Expect = 1e-09
Identities = 40/127 (31%), Positives = 65/127 (50%), Gaps = 19/127 (14%)
Frame = +2
Query: 1047 MLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEILSFP-SYDLILMDCQMPKMDG 1105
+L+K GA V AV G+ A+ ++L P ++D MD QMP+MDG
Sbjct: 8 VLQKYGAKVTAVESGRAAL------------------KMLELPHNFDACFMDLQMPEMDG 133
Query: 1106 YEATKEIR--KSEIGTS----------------FHIPIVALTAHAMSCDEAKCLEVGMDA 1147
+EAT++IR +SE+ +HIPI+A+TA + +C++ GM+
Sbjct: 134 FEATRKIRCLESEVNEKIACGQASAEMFGNISYWHIPILAMTADSTQSSNEECIKCGMND 313
Query: 1148 YLTKPID 1154
Y++KP +
Sbjct: 314 YVSKPFE 334
>TC216027 homologue to UP|Q9ZSQ0 (Q9ZSQ0) Ethylene response sensor, partial
(30%)
Length = 869
Score = 62.0 bits (149), Expect = 1e-09
Identities = 42/142 (29%), Positives = 60/142 (41%)
Frame = +3
Query: 577 DMPKLVRGDSARVVQIFANLINNSIKFTLSGHIILRGWCENPNSYGDSENFTLEQKPFGC 636
D+P GD R+ Q N++ N++KFT G++ +R P S D
Sbjct: 6 DLPTHAIGDEKRLTQTLLNVVGNAVKFTKEGYVSVRVSVAKPESSQD------------- 146
Query: 637 SRKTKMKQHENHSKKASNIDNKMILWFEVDDTGCGIDPSKWDSVFESFEQADLSTTRLHG 696
+ E + + D + +V D+GCGI P +F F Q+ R
Sbjct: 147 -----WRPPEFYPASS---DGHFYIRVQVKDSGCGILPQDIPHLFTKFAQSRSGPARPSS 302
Query: 697 GTGLGLCIVRSLVNKMGGEIKI 718
G GLGL I + VN MGG I I
Sbjct: 303 GAGLGLAICKRFVNLMGGHIWI 368
>BQ740961
Length = 428
Score = 61.2 bits (147), Expect = 3e-09
Identities = 32/90 (35%), Positives = 50/90 (55%), Gaps = 19/90 (21%)
Frame = +2
Query: 1084 EILSFP-SYDLILMDCQMPKMDGYEATKEIRKSEIGTS------------------FHIP 1124
E+L P ++D MD QMP+MDG+EAT++IR E + +HIP
Sbjct: 8 EMLQLPHNFDACFMDIQMPEMDGFEATRQIRMMETKANEQQMNGECGEGNGWKDKKYHIP 187
Query: 1125 IVALTAHAMSCDEAKCLEVGMDAYLTKPID 1154
I+A+TA + +C++ GMD Y++KP +
Sbjct: 188 ILAMTADVIHATYDECVKCGMDGYVSKPFE 277
>TC210860 similar to UP|Q9C5T8 (Q9C5T8) Cytokinin receptor CRE1b, partial
(15%)
Length = 661
Score = 59.7 bits (143), Expect = 7e-09
Identities = 30/62 (48%), Positives = 40/62 (64%)
Frame = +2
Query: 665 VDDTGCGIDPSKWDSVFESFEQADLSTTRLHGGTGLGLCIVRSLVNKMGGEIKIVKKEGP 724
V+DTG GI S D +F F QAD ST+R +GGTG+GL I + LV MGG+I + +
Sbjct: 8 VEDTGIGIPFSAQDRIFMPFVQADSSTSRNYGGTGIGLSISKCLVELMGGQINFISRPQV 187
Query: 725 GT 726
G+
Sbjct: 188 GS 193
>TC213342 weakly similar to UP|O82445 (O82445) Response regulator protein,
partial (41%)
Length = 599
Score = 57.4 bits (137), Expect = 4e-08
Identities = 40/135 (29%), Positives = 66/135 (48%), Gaps = 1/135 (0%)
Frame = +2
Query: 1029 LRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDA-LNGMPGVERNTITSQTEILS 1087
L L+ +D + +++ +LE +G V +GQ+AVD +G
Sbjct: 29 LTALVVDDNKINRKIHQKLLESVGMKNQGVENGQEAVDIHCHGQ---------------- 160
Query: 1088 FPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMDA 1147
+DLILMD MP M+G EATKE+R IG+ IV +++ + K +E G++
Sbjct: 161 --RFDLILMDMDMPIMNGIEATKELRSMGIGSM----IVGVSSRCTEAEIRKFMEAGLND 322
Query: 1148 YLTKPIDFKLMESTI 1162
Y KP++ + S +
Sbjct: 323 YHEKPLNNSKLSSLL 367
>TC230375 similar to UP|Q9SSY4 (Q9SSY4) Ethylene receptor CS-ETR2, partial
(22%)
Length = 736
Score = 56.6 bits (135), Expect = 6e-08
Identities = 41/137 (29%), Positives = 64/137 (45%)
Frame = +3
Query: 1017 SKAMNGKKSLEGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVER 1076
S+ + L L++LL E+ V + V +L+K+G V V G + + + G G
Sbjct: 129 SEHTDSNSMLRNLQVLLVENDDVNRAVTQRLLQKLGCVVTPVASGFECLTVI-GPAG--- 296
Query: 1077 NTITSQTEILSFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCD 1136
S +IL+D MP +DG+E IRK G IVALTA A
Sbjct: 297 ------------SSIQVILLDLHMPDIDGFEVATRIRKFRSGN--RPMIVALTASAEEDL 434
Query: 1137 EAKCLEVGMDAYLTKPI 1153
+C++VG++ + KP+
Sbjct: 435 WDRCMQVGINGVIRKPV 485
>TC228370 similar to UP|Q9AXD7 (Q9AXD7) Response regulator protein, partial
(48%)
Length = 877
Score = 55.8 bits (133), Expect = 1e-07
Identities = 37/127 (29%), Positives = 64/127 (50%)
Frame = +2
Query: 1027 EGLRILLAEDTAVIQRVATIMLEKMGATVVAVGDGQQAVDALNGMPGVERNTITSQTEIL 1086
E + +L +D+ V ++V +L+ + V AV G +A+ L + + I S+T
Sbjct: 125 EEVHVLAVDDSTVDRKVIEHLLKVLACKVTAVDSGLRALQLLGLL---DEQKIPSETNGF 295
Query: 1087 SFPSYDLILMDCQMPKMDGYEATKEIRKSEIGTSFHIPIVALTAHAMSCDEAKCLEVGMD 1146
DLI+ D MP M GYE K I++S T P+V +++ + +CLE G +
Sbjct: 296 VGLKVDLIITDYCMPGMTGYELLKRIKES--STFKETPVVIMSSENVLPRIDRCLEEGAE 469
Query: 1147 AYLTKPI 1153
++ KP+
Sbjct: 470 DFIVKPV 490
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,307,340
Number of Sequences: 63676
Number of extensions: 634584
Number of successful extensions: 3507
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 3398
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3469
length of query: 1172
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1064
effective length of database: 5,762,624
effective search space: 6131431936
effective search space used: 6131431936
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC121233.1


