

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119419.7 - phase: 0
(208 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
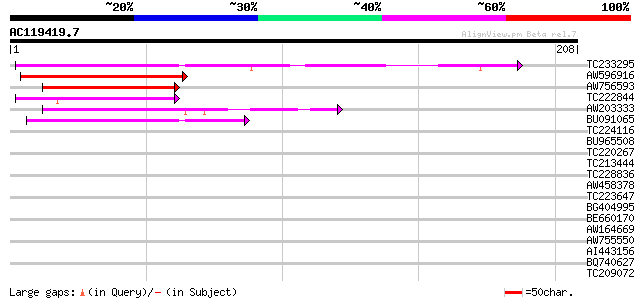
TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%) 77 6e-15
AW596916 52 1e-07
AW756593 49 2e-06
TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat... 45 3e-05
AW203333 42 3e-04
BU091065 40 8e-04
TC224116 39 0.001
BU965508 39 0.002
TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, parti... 37 0.006
TC213444 35 0.032
TC228836 similar to UP|Q6SH18 (Q6SH18) Lipase/esterase, Lip3/Bch... 33 0.092
AW458378 32 0.16
TC223647 30 0.60
BG404995 similar to GP|22831068|dbj OJ1131_E05.32 {Oryza sativa ... 30 1.0
BE660170 30 1.0
AW164669 29 1.3
AW755550 29 1.7
AI443156 28 2.3
BQ740627 28 2.3
TC209072 28 2.3
>TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%)
Length = 557
Score = 77.0 bits (188), Expect = 6e-15
Identities = 64/189 (33%), Positives = 88/189 (45%), Gaps = 3/189 (1%)
Frame = +3
Query: 3 ESNSKRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFN 62
E S RQK + +DDRIS LPD +L ILS LP K V TG LS++W+ LW + VLDF+
Sbjct: 24 EPPSHRQKPTTGDDDRISHLPDVLLLQILSLLPTKQAVITGILSKRWRPLWPAVSVLDFD 203
Query: 63 EYDDYHRSDNRKEQLRRFVVLVNNV-LHRNRHGIRKMRLTCAHSLVDDDNFRSHTVDTWV 121
D+ + L F V +V L + I + RL CA + N+ + + TW+
Sbjct: 204 --DESSPEFHHPGGLTGFAEFVYSVLLLHDAPAIERFRLRCA-----NPNYSARDIATWL 362
Query: 122 RSVIGPQLEELELDLDLFSDDFDEPLDGPDFDEPLDGTDFKLPLSLFTCP--NLVSLRHV 179
V + E +EL L L LP LF C +++ L V
Sbjct: 363 CHVARRRAERVELSLSL-------------------SRYVALPRCLFHCDTVSVMKLNGV 485
Query: 180 LLFFIASFS 188
L +ASFS
Sbjct: 486 FLNALASFS 512
>AW596916
Length = 455
Score = 52.4 bits (124), Expect = 1e-07
Identities = 26/61 (42%), Positives = 37/61 (60%)
Frame = +3
Query: 5 NSKRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEY 64
N + K EE+ DR+S LPD VL HI+ ++ +K V T LS++W+ LWK L L +
Sbjct: 228 NMQSDKDREEDMDRLSDLPDFVLLHIMKFMSMKHAVQTCVLSKRWKELWKRLTNLALHSS 407
Query: 65 D 65
D
Sbjct: 408 D 410
>AW756593
Length = 370
Score = 48.9 bits (115), Expect = 2e-06
Identities = 20/50 (40%), Positives = 34/50 (68%)
Frame = +3
Query: 13 EENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFN 62
+++ D+IS LPD++L H++ ++ + V T LS++W +LWK L L FN
Sbjct: 51 KDDRDKISELPDNILLHMMDFMDTREAVQTCVLSKRWNNLWKRLSTLLFN 200
>TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat shock
transcription factor HSF30, partial (6%)
Length = 746
Score = 44.7 bits (104), Expect = 3e-05
Identities = 25/63 (39%), Positives = 37/63 (58%), Gaps = 3/63 (4%)
Frame = +1
Query: 3 ESNSKRQKASEEND---DRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVL 59
+ N R+ E D DRIS+LPD +L H+++++ K+ V T LS++W L K L L
Sbjct: 445 KKNIAREVDGENEDHRPDRISALPDSLLFHMINFMDTKSAVQTCVLSKRWNDLSKCLTNL 624
Query: 60 DFN 62
FN
Sbjct: 625 TFN 633
>AW203333
Length = 418
Score = 41.6 bits (96), Expect = 3e-04
Identities = 35/125 (28%), Positives = 53/125 (42%), Gaps = 15/125 (12%)
Frame = +1
Query: 13 EENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNE--------- 63
E D IS+LP +L+ ILS +P + V T LS+ W W +L F++
Sbjct: 73 EMERDMISTLPKTILHDILSRMPEEDAVRTSVLSKSWAEAWSTYPILYFSDTMIVGTFPR 252
Query: 64 -YDDYHRS-----DNRKEQLRRFVVLVNNVLHRNRHGIRKMRLTCAHSLVDDDNFRSHTV 117
++D+ R D+ K L RF H I++ RL L + S V
Sbjct: 253 PWEDFLRKRKNFIDHVKRTLLRF--------HTQGLAIKQFRLIINFDL----QYMSLDV 396
Query: 118 DTWVR 122
D W++
Sbjct: 397 DHWLK 411
>BU091065
Length = 460
Score = 40.0 bits (92), Expect = 8e-04
Identities = 23/82 (28%), Positives = 38/82 (46%)
Frame = +3
Query: 7 KRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEYDD 66
K +E+ +RI LP +V+ IL LP + V T L+R W+++W +F DD
Sbjct: 60 KMADCDDEHVNRIGDLPMNVIVSILQRLPFQDLVKTSVLARAWRYMWSSTPRREFR--DD 233
Query: 67 YHRSDNRKEQLRRFVVLVNNVL 88
+ N L ++ +L
Sbjct: 234 FFEKCNHLGYLETSAIITEALL 299
>TC224116
Length = 520
Score = 39.3 bits (90), Expect = 0.001
Identities = 18/47 (38%), Positives = 27/47 (57%)
Frame = +2
Query: 17 DRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNE 63
D IS LP ++ IL LPI+ V T LS +W++ W + L F++
Sbjct: 287 DLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITQLVFDD 427
>BU965508
Length = 447
Score = 38.9 bits (89), Expect = 0.002
Identities = 18/47 (38%), Positives = 27/47 (57%)
Frame = +2
Query: 17 DRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNE 63
D IS LP ++ IL LPI+ V T LS +W++ W + L F++
Sbjct: 296 DLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITRLVFDD 436
>TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, partial (7%)
Length = 586
Score = 37.0 bits (84), Expect = 0.006
Identities = 18/45 (40%), Positives = 25/45 (55%), Gaps = 7/45 (15%)
Frame = +2
Query: 22 LPDDVLNHILSYLPIKTTVATGRLSRQWQHL-------WKHLHVL 59
LPDD+L IL+YLPI + G +S++W + W HVL
Sbjct: 338 LPDDLLERILAYLPIASIFRAGCVSKRWHEIVNSERFVWNLSHVL 472
>TC213444
Length = 530
Score = 34.7 bits (78), Expect = 0.032
Identities = 12/28 (42%), Positives = 22/28 (77%)
Frame = +2
Query: 13 EENDDRISSLPDDVLNHILSYLPIKTTV 40
EE++DR+S +PD +++HILS++ K +
Sbjct: 446 EESEDRLSDMPDCIIHHILSFMETKDAI 529
>TC228836 similar to UP|Q6SH18 (Q6SH18) Lipase/esterase, Lip3/BchO family,
partial (9%)
Length = 702
Score = 33.1 bits (74), Expect = 0.092
Identities = 19/61 (31%), Positives = 31/61 (50%), Gaps = 6/61 (9%)
Frame = +3
Query: 7 KRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQW------QHLWKHLHVLD 60
KR+ + + + LP D+L I S+L +K+ V+ G + R W HLW+ +V
Sbjct: 444 KRRCKAHQVEKGSLPLPQDILVLIFSFLDMKSLVSMGLVCRSWNIAANDNHLWEKQYVAL 623
Query: 61 F 61
F
Sbjct: 624 F 626
>AW458378
Length = 363
Score = 32.3 bits (72), Expect = 0.16
Identities = 17/44 (38%), Positives = 24/44 (53%)
Frame = +2
Query: 9 QKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHL 52
QK ++ +SSLP+DV I S L ++ A G SR W+ L
Sbjct: 89 QKGKADSVSILSSLPEDVALKIASLLQVRDLCALGCCSRFWREL 220
>TC223647
Length = 448
Score = 30.4 bits (67), Expect = 0.60
Identities = 15/48 (31%), Positives = 27/48 (56%)
Frame = +3
Query: 5 NSKRQKASEENDDRISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHL 52
+S + KAS + + +S L +D+L +IL+ LP + +S+ W L
Sbjct: 72 SSTKVKASTDANTNLSMLNEDLLQNILARLPALHFASAACVSKSWNSL 215
>BG404995 similar to GP|22831068|dbj OJ1131_E05.32 {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 394
Score = 29.6 bits (65), Expect = 1.0
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 7/42 (16%)
Frame = +1
Query: 22 LPDDVLNHILSYLPIKTTVATGRLSRQW------QHL-WKHL 56
LP +VL ILS LP+++ + S+ W QHL W HL
Sbjct: 40 LPREVLTEILSRLPVRSLLRFRSTSKSWKSLIDSQHLNWLHL 165
>BE660170
Length = 512
Score = 29.6 bits (65), Expect = 1.0
Identities = 23/87 (26%), Positives = 36/87 (40%), Gaps = 7/87 (8%)
Frame = -3
Query: 27 LNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFNEYDDYHRSDNRKEQLRRFVVLVNN 86
LNHI S + G L+ W HLW V++ Y D + N + V++ +
Sbjct: 237 LNHIKSSI-------FGFLNEVWPHLWSATRVMNRTRYQDSALAIN-----DQCSVVIRD 94
Query: 87 VLHRNRHGI-------RKMRLTCAHSL 106
+ + R GI + L C+H L
Sbjct: 93 ICSKRRRGINAEEPKQNQQNL*CSHCL 13
>AW164669
Length = 403
Score = 29.3 bits (64), Expect = 1.3
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = +2
Query: 22 LPDDVLNHILSYLPIKTTVATGRLS 46
LPDD+L IL+YLPI + G +S
Sbjct: 329 LPDDLLERILAYLPIASIFRAGCVS 403
>AW755550
Length = 435
Score = 28.9 bits (63), Expect = 1.7
Identities = 12/39 (30%), Positives = 22/39 (55%)
Frame = +3
Query: 48 QWQHLWKHLHVLDFNEYDDYHRSDNRKEQLRRFVVLVNN 86
QW +L ++ LDF+ DD ++ ++ ++ F LV N
Sbjct: 192 QWHNLDRYFSRLDFDVLDDKRYQEDAEKTMQEFTSLVRN 308
>AI443156
Length = 414
Score = 28.5 bits (62), Expect = 2.3
Identities = 12/44 (27%), Positives = 25/44 (56%)
Frame = +3
Query: 19 ISSLPDDVLNHILSYLPIKTTVATGRLSRQWQHLWKHLHVLDFN 62
+++LP +V+ ILS LP+K+ + + W+ + H + F+
Sbjct: 81 MANLPVEVVTEILSRLPVKSVIRLRSTCKWWRSIIDSRHFILFH 212
>BQ740627
Length = 426
Score = 28.5 bits (62), Expect = 2.3
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = +1
Query: 22 LPDDVLNHILSYLPIKTTVATG 43
LPDD+L IL+YLPI + G
Sbjct: 355 LPDDLLERILAYLPIASIFRAG 420
>TC209072
Length = 1312
Score = 28.5 bits (62), Expect = 2.3
Identities = 13/34 (38%), Positives = 19/34 (55%)
Frame = -1
Query: 168 FTCPNLVSLRHVLLFFIASFSFLNVKCHCGKIES 201
F+ +LVS RH++L + S F + CH I S
Sbjct: 652 FSPTHLVSCRHIMLILLRSHHFHRLTCHDTNIPS 551
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,946,183
Number of Sequences: 63676
Number of extensions: 170796
Number of successful extensions: 1108
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 1106
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1108
length of query: 208
length of database: 12,639,632
effective HSP length: 93
effective length of query: 115
effective length of database: 6,717,764
effective search space: 772542860
effective search space used: 772542860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC119419.7


