

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119419.5 - phase: 0
(735 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
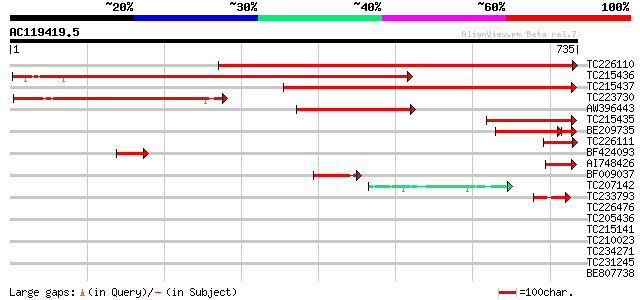
Score E
Sequences producing significant alignments: (bits) Value
TC226110 similar to UP|TKTC_CRAPL (Q42676) Transketolase, chloro... 854 0.0
TC215436 similar to UP|Q9LZY8 (Q9LZY8) Transketolase-like protei... 853 0.0
TC215437 homologue to UP|TKTC_CRAPL (Q42676) Transketolase, chlo... 669 0.0
TC223730 similar to UP|Q7SIC9 (Q7SIC9) Transferase, partial (29%) 341 8e-94
AW396443 similar to SP|Q42676|TKTC Transketolase chloroplast (E... 218 6e-57
TC215435 homologue to UP|TKTC_CRAPL (Q42676) Transketolase, chlo... 207 1e-53
BE209735 similar to GP|25067747|gb| thioredoxin/transketolase fu... 120 3e-31
TC226111 homologue to UP|TKTC_CRAPL (Q42676) Transketolase, chlo... 83 5e-16
BF424093 similar to PIR|T09541|T095 transketolase (EC 2.2.1.1) T... 80 2e-15
AI748426 similar to PIR|T09541|T095 transketolase (EC 2.2.1.1) T... 63 4e-10
BF009037 homologue to PIR|T09541|T095 transketolase (EC 2.2.1.1)... 48 2e-05
TC207142 similar to UP|Q8L692 (Q8L692) 1-deoxy-D-xylulose 5-phos... 45 2e-04
TC233793 similar to UP|TKT_BUCAI (P57195) Transketolase (TK) , ... 42 7e-04
TC226476 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-ph... 37 0.024
TC205436 homologue to UP|PRS6_ARATH (Q9SEI4) 26S protease regula... 37 0.041
TC215141 similar to GB|AAL32703.1|17065098|AY062625 nucleotide s... 35 0.16
TC210023 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 ... 33 0.35
TC234271 homologue to UP|Q25639 (Q25639) Allergen (Fragment), pa... 33 0.35
TC231245 similar to UP|Q9ARE4 (Q9ARE4) ZF-HD homeobox protein, p... 33 0.59
BE807738 weakly similar to GP|4098246|gb|A homeobox 2 protein {L... 32 1.3
>TC226110 similar to UP|TKTC_CRAPL (Q42676) Transketolase, chloroplast (TK)
(Fragment) , partial (90%)
Length = 1719
Score = 854 bits (2206), Expect = 0.0
Identities = 423/465 (90%), Positives = 441/465 (93%)
Frame = +2
Query: 271 ESVEKRFEGLGWHVIWVKNGNTGFDEIRAAIKEAKAVKDRPTLIKFTTTIGYGSPNKSNS 330
ESV+ RFEGLGWHVIWVKNGN G+D+IRAAIKEAKAVKD+PTLIK TTTIGYGSPNK+NS
Sbjct: 2 ESVDSRFEGLGWHVIWVKNGNNGYDDIRAAIKEAKAVKDKPTLIKVTTTIGYGSPNKANS 181
Query: 331 YSVHGSALGAKEVDATRNNLGWPYEPFHVPEDVKKHWSRHIREGAALESEWNAKFADYEK 390
YSVHGSALGAKEVDATR NLGW +EPFHVPEDVKKHWSRH EGAALE+EWNAKFA+YEK
Sbjct: 182 YSVHGSALGAKEVDATRQNLGWSHEPFHVPEDVKKHWSRHTPEGAALEAEWNAKFAEYEK 361
Query: 391 KYKEEAAVLKSIISGDLPAGWEKALPTYTPEIPADATRNLSQQNLNALANVLPGLIGGSA 450
KYKEEAA LKSII+G+ PAGWEKALPTYTPE PADATRNLSQ NLNALA VLPGL+GGSA
Sbjct: 362 KYKEEAAELKSIINGEFPAGWEKALPTYTPESPADATRNLSQTNLNALAKVLPGLLGGSA 541
Query: 451 DLASSNMTLLKKFGDFQSDSPAERNVRFGVREHGMGAICNGIALHSRGLIPYCATFFVFT 510
DLASSNMTLLK FGDFQ D+PAERNVRFGVREHGMGAICNGIALHS GLIPYCATFFVFT
Sbjct: 542 DLASSNMTLLKMFGDFQKDTPAERNVRFGVREHGMGAICNGIALHSPGLIPYCATFFVFT 721
Query: 511 DYMRGAIRLSALSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNE 570
DYMRGAIRLSALSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNE
Sbjct: 722 DYMRGAIRLSALSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNE 901
Query: 571 TAGAYKVAVLNRKRPSILALSRQKLPNLAGTSIEGVEKGGYIVSDNSTGNKPDVILIGTG 630
TAGAYKVAVLNRKRPSILALSRQKLP L GTSIEGVEKGGY +SDNSTGNKPDVILIGTG
Sbjct: 902 TAGAYKVAVLNRKRPSILALSRQKLPQLPGTSIEGVEKGGYTISDNSTGNKPDVILIGTG 1081
Query: 631 SELEIAYKAGEDLRKEGKAVRVVSFVSWELFDEQSQAYKESVLPTAVTARVSIEAGSTFG 690
SELEIA KA +DLRKEGKAVRVVS VSWELFDEQS+AYKESV P AV+ARVSIEAGSTFG
Sbjct: 1082SELEIAAKAADDLRKEGKAVRVVSLVSWELFDEQSEAYKESVFPAAVSARVSIEAGSTFG 1261
Query: 691 WEKIVGSKGKAIGIDRFGASAPAGTIYKEFGITKEAVIAAAKELI 735
WEKIVG+KGKAIGIDRFGASAPAG IYKEFGITKEAV+AAAKELI
Sbjct: 1262WEKIVGAKGKAIGIDRFGASAPAGRIYKEFGITKEAVVAAAKELI 1396
>TC215436 similar to UP|Q9LZY8 (Q9LZY8) Transketolase-like protein, partial
(65%)
Length = 1625
Score = 853 bits (2205), Expect = 0.0
Identities = 421/528 (79%), Positives = 457/528 (85%), Gaps = 9/528 (1%)
Frame = +2
Query: 4 SSSTSLYLTNPHLTR----HTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIH 59
+S++SL+L+ L R H SS LS PS L ++ +P + S R
Sbjct: 47 ASTSSLHLSQALLARAVYLHGSSSDRVSLS-FPSFSGL-KSHSPCKAAATSSRRRGACAS 220
Query: 60 ASVSAPPST-----TTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDE 114
SV + TT+ SLVEKSVNTIRFLA+D+VEKANSGHPGLPMGCAPMGH+LYDE
Sbjct: 221 TSVVRAAAVETLDQTTEVSLVEKSVNTIRFLAIDAVEKANSGHPGLPMGCAPMGHILYDE 400
Query: 115 VMRYNPKNPFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPE 174
VMRYNPKNP WFNRDRF+LSAGHGCMLQYALLHLAGYDSV EDLK+FRQW SRTPGHPE
Sbjct: 401 VMRYNPKNPTWFNRDRFILSAGHGCMLQYALLHLAGYDSVLEEDLKEFRQWGSRTPGHPE 580
Query: 175 NFETPGIEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQMEGI 234
NFET GIEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEI+DHYTY ILGDGCQMEGI
Sbjct: 581 NFETVGIEVTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIVDHYTYVILGDGCQMEGI 760
Query: 235 ANEACSLAGHWGLGKLIAFYDDNHISIDGDTEIAFTESVEKRFEGLGWHVIWVKNGNTGF 294
+NEACSLAGHWGLGKLIA YDDNHISIDGDTEIAFTE+V++RFE LGWHVIWVKNGNTG+
Sbjct: 761 SNEACSLAGHWGLGKLIALYDDNHISIDGDTEIAFTENVDQRFEALGWHVIWVKNGNTGY 940
Query: 295 DEIRAAIKEAKAVKDRPTLIKFTTTIGYGSPNKSNSYSVHGSALGAKEVDATRNNLGWPY 354
DEIRAAIKEAKAVKD+PT+IK TTTIG+GSPNK+NSYSVHGSALG KEVDATR NLGWPY
Sbjct: 941 DEIRAAIKEAKAVKDKPTMIKVTTTIGFGSPNKANSYSVHGSALGTKEVDATRKNLGWPY 1120
Query: 355 EPFHVPEDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKA 414
EPFHVPEDVKKHWSRH EGA LE+EWNAKFA+YEKKY EEAA LK+II+G+LPAGWEKA
Sbjct: 1121EPFHVPEDVKKHWSRHTPEGAKLEAEWNAKFAEYEKKYSEEAAELKAIITGELPAGWEKA 1300
Query: 415 LPTYTPEIPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAER 474
LPTYTPE PADATRNLSQQNLNAL VLPGL+GGSADLASSNMTLLK +GDFQ ++P ER
Sbjct: 1301LPTYTPESPADATRNLSQQNLNALVKVLPGLLGGSADLASSNMTLLKSYGDFQKNTPEER 1480
Query: 475 NVRFGVREHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSAL 522
NVRFGVREHGMGAICNGIALHS G IPYCATFFVFTDYMR AIR+SAL
Sbjct: 1481NVRFGVREHGMGAICNGIALHSPGFIPYCATFFVFTDYMRAAIRISAL 1624
>TC215437 homologue to UP|TKTC_CRAPL (Q42676) Transketolase, chloroplast
(TK) (Fragment) , partial (73%)
Length = 1475
Score = 669 bits (1726), Expect = 0.0
Identities = 332/379 (87%), Positives = 350/379 (91%)
Frame = +2
Query: 356 PFHVPEDVKKHWSRHIREGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKAL 415
PFHVPEDVKKHWSRH EGA LE+EWNAKF +YEK+Y EEAA LK+II+G+LPAGWEKAL
Sbjct: 2 PFHVPEDVKKHWSRHTPEGAKLEAEWNAKFVEYEKQYSEEAAELKAIITGELPAGWEKAL 181
Query: 416 PTYTPEIPADATRNLSQQNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAERN 475
PTYTPE PADATRNLSQQNLNAL VLPGL+GGSADLASSNMTLLK +GDFQ ++P ERN
Sbjct: 182 PTYTPESPADATRNLSQQNLNALVKVLPGLLGGSADLASSNMTLLKSYGDFQKNTPEERN 361
Query: 476 VRFGVREHGMGAICNGIALHSRGLIPYCATFFVFTDYMRGAIRLSALSEAGVIYVMTHDS 535
VRFGVREHGMGAICNGIALHS G IPYCATFFVFTDYMR AIR+SAL EAGVIYVMTHDS
Sbjct: 362 VRFGVREHGMGAICNGIALHSPGFIPYCATFFVFTDYMRAAIRISALCEAGVIYVMTHDS 541
Query: 536 IGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKL 595
IGLGEDGPTHQPIEHLASFRAMPN LMLRPADGNETAG+YKVAV+NRKRPSILALSRQKL
Sbjct: 542 IGLGEDGPTHQPIEHLASFRAMPNTLMLRPADGNETAGSYKVAVVNRKRPSILALSRQKL 721
Query: 596 PNLAGTSIEGVEKGGYIVSDNSTGNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVSF 655
L GTSIEGVEKGGY +SDNS+GNKPDVILIGTGSELEIA A EDLRKEGKAVRVVSF
Sbjct: 722 TQLPGTSIEGVEKGGYTISDNSSGNKPDVILIGTGSELEIAAAAAEDLRKEGKAVRVVSF 901
Query: 656 VSWELFDEQSQAYKESVLPTAVTARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGT 715
VSWELFDEQS YKESVLP +VT RVSIEAGSTFGW+KIVGS+GKAIGIDRFGASAPAG
Sbjct: 902 VSWELFDEQSDEYKESVLPASVTVRVSIEAGSTFGWQKIVGSQGKAIGIDRFGASAPAGK 1081
Query: 716 IYKEFGITKEAVIAAAKEL 734
IYKEFGITKEAVIAAAKEL
Sbjct: 1082IYKEFGITKEAVIAAAKEL 1138
>TC223730 similar to UP|Q7SIC9 (Q7SIC9) Transferase, partial (29%)
Length = 865
Score = 341 bits (874), Expect = 8e-94
Identities = 188/290 (64%), Positives = 210/290 (71%), Gaps = 12/290 (4%)
Frame = +1
Query: 5 SSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHA--SV 62
S++S+ L+ L T+S ++ SSL T TL TP LS L T+I A SV
Sbjct: 1 STSSISLSQALLRPTTTSLPSSSSSSLRLTPTLRPTPPA---LSLRRRLT-TSIRAAASV 168
Query: 63 SAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGHPGLPMGCAPMGHVLYDEVMRYNPKN 122
T D++LVEKSVNTIRFLA+D+VEKANSGHPGLPMGCAPMGHVLYDE M+YNPKN
Sbjct: 169 KTVEKATADAALVEKSVNTIRFLAIDAVEKANSGHPGLPMGCAPMGHVLYDETMKYNPKN 348
Query: 123 PFWFNRDRFVLSAGHGCMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPENFETPGIE 182
PFWFNRDRFVLSAGHGCMLQYALLHLAG+DSVK EDL++FRQW SRTPGHPENFETPGIE
Sbjct: 349 PFWFNRDRFVLSAGHGCMLQYALLHLAGFDSVKEEDLREFRQWGSRTPGHPENFETPGIE 528
Query: 183 VTTGPLGQGIANAVGLALAEKHLAARFNKPDNEIIDHYTYCILGDGCQM-EGIANEACSL 241
VTTGPLGQGIANAVGLALAEKHLAAR+NKPDNE DHYTY ILGDGCQ + + L
Sbjct: 529 VTTGPLGQGIANAVGLALAEKHLAARYNKPDNENGDHYTYAILGDGCQNGXELQMKHAHL 708
Query: 242 AGHWGLGKLIA-------FYDDNHISIDGDTEIAFTESVEKRFEGL--GW 282
GHWGLGKL + FY H + V+ RFEGL GW
Sbjct: 709 VGHWGLGKLDSIL**QSYFY*WXH------RKXV*PRVVDSRFEGLWVGW 840
>AW396443 similar to SP|Q42676|TKTC Transketolase chloroplast (EC 2.2.1.1)
(TK) (Fragment). {Craterostigma plantagineum}, partial
(29%)
Length = 466
Score = 218 bits (556), Expect = 6e-57
Identities = 110/153 (71%), Positives = 125/153 (80%)
Frame = +2
Query: 373 EGAALESEWNAKFADYEKKYKEEAAVLKSIISGDLPAGWEKALPTYTPEIPADATRNLSQ 432
EGA LE+EWNA+FA+ EKKY EEAA LK+II+G+LP GWEKAL TYT + PADATRNL Q
Sbjct: 8 EGAQLEAEWNAQFAESEKKYIEEAAELKAIITGELPPGWEKALATYTSQSPADATRNLPQ 187
Query: 433 QNLNALANVLPGLIGGSADLASSNMTLLKKFGDFQSDSPAERNVRFGVREHGMGAICNGI 492
QNLNAL VLPGL+GGSA LASSNMTLL +GDF +++P ER VR GVREH MGAIC G+
Sbjct: 188 QNLNALVKVLPGLLGGSAYLASSNMTLLTSYGDFHTNTPEERIVRCGVREHVMGAICCGL 367
Query: 493 ALHSRGLIPYCATFFVFTDYMRGAIRLSALSEA 525
ALHS IPYCAT V+TDYMR AIR+ AL EA
Sbjct: 368 ALHSPRFIPYCATHSVYTDYMRAAIRIPALCEA 466
>TC215435 homologue to UP|TKTC_CRAPL (Q42676) Transketolase, chloroplast
(TK) (Fragment) , partial (22%)
Length = 669
Score = 207 bits (527), Expect = 1e-53
Identities = 107/116 (92%), Positives = 108/116 (92%)
Frame = +2
Query: 619 GNKPDVILIGTGSELEIAYKAGEDLRKEGKAVRVVSFVSWELFDEQSQAYKESVLPTAVT 678
GNKPDVILIGTGSELEIA A EDLRKEGKAVRVVSFVSWELFDEQS YKESVLP +VT
Sbjct: 2 GNKPDVILIGTGSELEIAAAAAEDLRKEGKAVRVVSFVSWELFDEQSDEYKESVLPASVT 181
Query: 679 ARVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGTIYKEFGITKEAVIAAAKEL 734
ARVSIEAGSTFGW KIVGSKGKAIGIDRFGASAPAG IYKEFGITKEAVIAAAKEL
Sbjct: 182 ARVSIEAGSTFGWHKIVGSKGKAIGIDRFGASAPAGKIYKEFGITKEAVIAAAKEL 349
>BE209735 similar to GP|25067747|gb| thioredoxin/transketolase fusion protein
{synthetic construct}, partial (10%)
Length = 434
Score = 120 bits (300), Expect(2) = 3e-31
Identities = 65/87 (74%), Positives = 70/87 (79%)
Frame = +2
Query: 630 GSELEIAYKAGEDLRKEGKAVRVVSFVSWELFDEQSQAYKESVLPTAVTARVSIEAGSTF 689
GSELEIA A EDLRKEGKAVRVV+FVSWEL DEQS YKESVLP +VT RVSIEAGST
Sbjct: 14 GSELEIAAAAAEDLRKEGKAVRVVAFVSWELCDEQSDEYKESVLPASVTVRVSIEAGSTC 193
Query: 690 GWEKIVGSKGKAIGIDRFGASAPAGTI 716
+K+ GS+GKAI IDRFGA A A I
Sbjct: 194 CGQKMGGSQGKAIXIDRFGAGARAKNI 274
Score = 34.3 bits (77), Expect(2) = 3e-31
Identities = 16/19 (84%), Positives = 17/19 (89%)
Frame = +3
Query: 716 IYKEFGITKEAVIAAAKEL 734
IY EFGIT+EAVIA AKEL
Sbjct: 270 IYXEFGITREAVIACAKEL 326
>TC226111 homologue to UP|TKTC_CRAPL (Q42676) Transketolase, chloroplast
(TK) (Fragment) , partial (8%)
Length = 389
Score = 82.8 bits (203), Expect = 5e-16
Identities = 41/44 (93%), Positives = 43/44 (97%)
Frame = +3
Query: 692 EKIVGSKGKAIGIDRFGASAPAGTIYKEFGITKEAVIAAAKELI 735
EKIVG+KGKAIGIDRFGASAPAG IYKEFGITKEAV+AAAKELI
Sbjct: 99 EKIVGAKGKAIGIDRFGASAPAGRIYKEFGITKEAVVAAAKELI 230
>BF424093 similar to PIR|T09541|T095 transketolase (EC 2.2.1.1) TKT1
precursor chloroplast [validated] - pepper, partial
(7%)
Length = 192
Score = 80.5 bits (197), Expect = 2e-15
Identities = 35/42 (83%), Positives = 39/42 (92%)
Frame = +3
Query: 139 CMLQYALLHLAGYDSVKVEDLKQFRQWESRTPGHPENFETPG 180
CMLQYALLHLAGYD+V+ +DLK+FRQW SRTPGHPENFET G
Sbjct: 66 CMLQYALLHLAGYDTVQEQDLKEFRQWGSRTPGHPENFETLG 191
>AI748426 similar to PIR|T09541|T095 transketolase (EC 2.2.1.1) TKT1
precursor chloroplast [validated] - pepper, partial
(4%)
Length = 152
Score = 63.2 bits (152), Expect = 4e-10
Identities = 32/40 (80%), Positives = 33/40 (82%)
Frame = +3
Query: 695 VGSKGKAIGIDRFGASAPAGTIYKEFGITKEAVIAAAKEL 734
VGS GKAIGIDRF A APAG IY EFG+TKEAVIAAA L
Sbjct: 6 VGSHGKAIGIDRFAARAPAGKIYMEFGVTKEAVIAAATYL 125
>BF009037 homologue to PIR|T09541|T095 transketolase (EC 2.2.1.1) TKT1
precursor chloroplast [validated] - pepper, partial
(4%)
Length = 263
Score = 47.8 bits (112), Expect = 2e-05
Identities = 30/62 (48%), Positives = 39/62 (62%)
Frame = +2
Query: 394 EEAAVLKSIISGDLPAGWEKALPTYTPEIPADATRNLSQQNLNALANVLPGLIGGSADLA 453
EEAA LK+II+G+LPAGWEKAL +T + + + QNLNAL V + G +LA
Sbjct: 11 EEAAELKAIITGELPAGWEKAL-RHTHQKALLMLQEICSQNLNALVKVF--WLLGECNLA 181
Query: 454 SS 455
S
Sbjct: 182 CS 187
>TC207142 similar to UP|Q8L692 (Q8L692) 1-deoxy-D-xylulose 5-phosphate
synthase 2 precursor, partial (55%)
Length = 1176
Score = 44.7 bits (104), Expect = 2e-04
Identities = 55/204 (26%), Positives = 82/204 (39%), Gaps = 17/204 (8%)
Frame = +1
Query: 466 FQSDSPAERNVRFGVREHGMGAICNGIALHSRGLIPYCATFFVF-----------TDYMR 514
FQ P ER G+ E G+A + GL P+CA + F D +
Sbjct: 412 FQKKFP-ERCFDVGIAEQHAVTFAAGLA--AEGLKPFCAIYSSFLQRGYDQVVHDVDLQK 582
Query: 515 GAIRLSALSEAGVIYVMTHDSIGLGEDGPTHQPIEHLASFRAMPNILMLRPADGNETAGA 574
+R AL AG++ G DGPTH + +PN++++ P+D E
Sbjct: 583 LPVRF-ALDRAGLV----------GADGPTHCGAFDITYMACLPNMVVMAPSDETELMHM 729
Query: 575 YKVAVLNRKRPSILALSR-----QKLP-NLAGTSIEGVEKGGYIVSDNSTGNKPDVILIG 628
A RPS R LP N GT +E + KG +V + V ++G
Sbjct: 730 VATAAAIDDRPSCFRFPRGNGIGATLPLNNKGTPLE-IGKGRILVEGSR------VAILG 888
Query: 629 TGSELEIAYKAGEDLRKEGKAVRV 652
GS ++ +A L++ G V V
Sbjct: 889 YGSVVQQCRQAS*MLKELGIDVTV 960
>TC233793 similar to UP|TKT_BUCAI (P57195) Transketolase (TK) , partial (3%)
Length = 437
Score = 42.4 bits (98), Expect = 7e-04
Identities = 21/48 (43%), Positives = 30/48 (61%)
Frame = +3
Query: 680 RVSIEAGSTFGWEKIVGSKGKAIGIDRFGASAPAGTIYKEFGITKEAV 727
R+S+EA S GWEK +IG+ FG+SAP ++K+FG T + V
Sbjct: 6 RISVEAASITGWEKYAHF---SIGMVTFGSSAPPDDVFKKFGFTVDQV 140
>TC226476 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-phosphate
synthase 1 precursor, partial (85%)
Length = 2000
Score = 37.4 bits (85), Expect = 0.024
Identities = 40/149 (26%), Positives = 58/149 (38%), Gaps = 5/149 (3%)
Frame = +3
Query: 487 AICNGIALHSRGLIPYCATFFVFTDYMRGA----IRLSALSEAGVIYVMTHDSIGL-GED 541
A+ L GL P+CA +++ +M+ A + L + V + M D GL G D
Sbjct: 1320 AVTFAAGLACEGLKPFCA---IYSSFMQRAYDQVVHDVDLQKLPVRFAM--DRAGLVGAD 1484
Query: 542 GPTHQPIEHLASFRAMPNILMLRPADGNETAGAYKVAVLNRKRPSILALSRQKLPNLAGT 601
GPTH + +PN++++ P+D E A RPS R
Sbjct: 1485 GPTHCGSFDVTFMACLPNMVVMAPSDEAELFHMVATAAAINDRPSCFRYPR--------- 1637
Query: 602 SIEGVEKGGYIVSDNSTGNKPDVILIGTG 630
G I TGNK + IG G
Sbjct: 1638 -------GNGIGVQLPTGNKGTPLEIGKG 1703
>TC205436 homologue to UP|PRS6_ARATH (Q9SEI4) 26S protease regulatory subunit
6B homolog (26S proteasome AAA-ATPase subunit RPT3)
(Regulatory particle triple-A ATPase subunit 3), partial
(32%)
Length = 598
Score = 36.6 bits (83), Expect = 0.041
Identities = 23/58 (39%), Positives = 30/58 (51%)
Frame = +3
Query: 13 NPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSAPPSTTT 70
+P T +S+PST SS P T TPTP+ S H R TA S ++P S T+
Sbjct: 420 DPTTTSESSAPSTVSFSSPPHTSRSTGTPTPS---STFFHRRPTAASRSSASPRSLTS 584
>TC215141 similar to GB|AAL32703.1|17065098|AY062625 nucleotide sugar
epimerase-like protein {Arabidopsis thaliana;} , partial
(89%)
Length = 1743
Score = 34.7 bits (78), Expect = 0.16
Identities = 24/70 (34%), Positives = 33/70 (46%)
Frame = +2
Query: 5 SSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSA 64
+S+S L+ H + +SSPSTT S P+ T TPTPTT + A+
Sbjct: 113 NSSSAPLSWSHSSSSSSSPSTTLPSPPPTPPTATSTPTPTTSSPPPPSAPGRSRSATPPP 292
Query: 65 PPSTTTDSSL 74
P + T SL
Sbjct: 293 PAAPTASPSL 322
>TC210023 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 protein
(Corresponding sequence ZK643.8), partial (12%)
Length = 819
Score = 33.5 bits (75), Expect = 0.35
Identities = 23/97 (23%), Positives = 49/97 (49%), Gaps = 1/97 (1%)
Frame = -2
Query: 2 SSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHAS 61
+++S+T+ T H + +S+ STT ++ +T TLN+ T +T + + TA +
Sbjct: 401 TTTSTTTATTTLQHRSTPSSTASTTTRTTTATTTTLNQISTASTTTTTTSTTTSTATTLN 222
Query: 62 -VSAPPSTTTDSSLVEKSVNTIRFLAVDSVEKANSGH 97
+S +TT+ ++ ++N I + + A + H
Sbjct: 221 QISTASTTTSTTTAAATTLNQISTASTTTSTTATTFH 111
Score = 28.9 bits (63), Expect = 8.6
Identities = 20/88 (22%), Positives = 42/88 (47%)
Frame = -2
Query: 2 SSSSSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHAS 61
+S+++T ++ P T T++ +TT ++L T + T + TT + + T
Sbjct: 455 TSTATTLQPISTPSSTASTTTSTTTATTTLQHRSTPSSTASTTTRTTTAT----TTTLNQ 288
Query: 62 VSAPPSTTTDSSLVEKSVNTIRFLAVDS 89
+S +TTT +S + T+ ++ S
Sbjct: 287 ISTASTTTTTTSTTTSTATTLNQISTAS 204
>TC234271 homologue to UP|Q25639 (Q25639) Allergen (Fragment), partial (7%)
Length = 568
Score = 33.5 bits (75), Expect = 0.35
Identities = 22/67 (32%), Positives = 34/67 (49%), Gaps = 1/67 (1%)
Frame = +3
Query: 5 SSTSLYLTNPHLTR-HTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVS 63
SST++ +P R TS+ ST + ++ + RTP PTT S S T+ S
Sbjct: 342 SSTTVPSASPSTARSRTSAASTCPTCTSRTSSSSTRTPPPTTTTSSSRRTSTTSTTFCPS 521
Query: 64 APPSTTT 70
PP++T+
Sbjct: 522 RPPTSTS 542
>TC231245 similar to UP|Q9ARE4 (Q9ARE4) ZF-HD homeobox protein, partial (32%)
Length = 739
Score = 32.7 bits (73), Expect = 0.59
Identities = 24/84 (28%), Positives = 36/84 (42%)
Frame = +3
Query: 5 SSTSLYLTNPHLTRHTSSPSTTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAIHASVSA 64
S+T+ L T T+SP TT +S TP P + + HLR + S S
Sbjct: 48 STTARNLHRRRCTTTTNSPRTTTTAS-------RSTPPPPATSTTTSHLRCRSTARSPSR 206
Query: 65 PPSTTTDSSLVEKSVNTIRFLAVD 88
PP S+ +K+ T+ A +
Sbjct: 207 PPHPAAASAAKKKTCQTLAAAAAE 278
>BE807738 weakly similar to GP|4098246|gb|A homeobox 2 protein {Lycopersicon
esculentum}, partial (16%)
Length = 238
Score = 31.6 bits (70), Expect = 1.3
Identities = 17/43 (39%), Positives = 24/43 (55%), Gaps = 1/43 (2%)
Frame = +2
Query: 25 TTRLSSLPSTITLNRTPTPTTHLSQSHHLRHTAI-HASVSAPP 66
T ++ L T +N+ P T LS SHHL H+ I H ++S P
Sbjct: 59 TXHITYLHITTPINQLPLINTQLSXSHHLLHSYISHNTLSLSP 187
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,966,408
Number of Sequences: 63676
Number of extensions: 440003
Number of successful extensions: 3373
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 3245
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3352
length of query: 735
length of database: 12,639,632
effective HSP length: 104
effective length of query: 631
effective length of database: 6,017,328
effective search space: 3796933968
effective search space used: 3796933968
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC119419.5


