

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119414.8 + phase: 0 /pseudo
(427 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
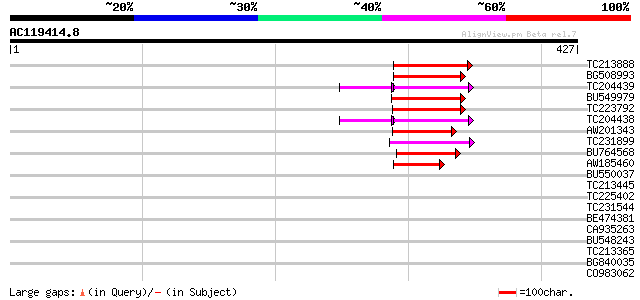
Score E
Sequences producing significant alignments: (bits) Value
TC213888 similar to UP|Q9SFE1 (Q9SFE1) T26F17.17, partial (11%) 67 2e-11
BG508993 62 4e-10
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 48 6e-07
BU549979 50 1e-06
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 47 1e-05
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 45 2e-05
AW201343 47 2e-05
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 45 6e-05
BU764568 44 1e-04
AW185460 44 2e-04
BU550037 41 0.001
TC213445 40 0.002
TC225402 weakly similar to UP|Q6I923 (Q6I923) Copia-like retrotr... 38 0.010
TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, ... 37 0.013
BE474381 similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, pa... 35 0.084
CA935263 32 0.55
BU548243 32 0.71
TC213365 similar to UP|Q6I8L6 (Q6I8L6) Reverse transcriptase (Fr... 31 0.93
BG840035 weakly similar to GP|9294121|dbj copia-like retrotransp... 30 2.1
CO983062 29 3.5
>TC213888 similar to UP|Q9SFE1 (Q9SFE1) T26F17.17, partial (11%)
Length = 493
Score = 66.6 bits (161), Expect = 2e-11
Identities = 31/59 (52%), Positives = 41/59 (68%)
Frame = +3
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
KST+ +V+ +G+ A +W S K+PIVTLST EAE+VAA SC C IWLR + + QE
Sbjct: 18 KSTTGFVFFMGDTAFTWMSKKQPIVTLSTCEAEYVAATSCVCHAIWLRNLLKELKMPQE 194
>BG508993
Length = 374
Score = 62.4 bits (150), Expect = 4e-10
Identities = 28/54 (51%), Positives = 39/54 (71%)
Frame = +1
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
KST+ +V+ +G+ +WSS K+ IVTL T EAE+VAA SC+C IWLRR+ +
Sbjct: 103 KSTTGFVFFMGDCVFTWSSKKQGIVTLFTCEAEYVAATSCTCHAIWLRRLLEEL 264
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 47.8 bits (112), Expect(2) = 6e-07
Identities = 23/60 (38%), Positives = 36/60 (59%)
Frame = +1
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQEI 349
KSTS + +GN ISW S K+ V+LST EAE++AA S +W++++ Q++
Sbjct: 4306 KSTSGGCFYLGNNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDV 4485
Score = 23.5 bits (49), Expect(2) = 6e-07
Identities = 16/43 (37%), Positives = 23/43 (53%)
Frame = +2
Query: 249 *REF*DT*KELSILKFCTKVKRTLA*LDGLTQIIQEIHMTEKS 291
*REF*+ * L + CT + + L + I E+ MTEK+
Sbjct: 4184 *REF*NM*MALVTMGLCTVIVQIQCWLGIVMLIGLEVQMTEKA 4312
>BU549979
Length = 615
Score = 50.4 bits (119), Expect = 1e-06
Identities = 20/56 (35%), Positives = 34/56 (60%)
Frame = -1
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
+ +STS Y++M+ +G +SW S K+ ++ ST E EFV + G+WL+ S +
Sbjct: 450 SRRSTSGYIFMLADGVVSWRSSKQTLIATSTMEVEFVPCFEATSHGVWLKSFMSSL 283
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein,
partial (7%)
Length = 804
Score = 47.4 bits (111), Expect = 1e-05
Identities = 20/55 (36%), Positives = 35/55 (63%)
Frame = +1
Query: 289 EKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
EKST Y++M + ++ SS K+ ++ LST EAE+VAA+ +C +W+ + +
Sbjct: 268 EKSTGGYLFMYNDAPVA*SSKKQDVIALSTCEAEYVAASLGACQAVWMMNLLEEL 432
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 45.4 bits (106), Expect(2) = 2e-05
Identities = 22/60 (36%), Positives = 35/60 (57%)
Frame = +1
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQEI 349
KSTS + +G ISW S K+ V+LST EAE++AA S +W++++ Q++
Sbjct: 4309 KSTSGGCFYLGTNLISWFSKKQNCVSLSTAEAEYIAAGSSCSQLVWMKQMLKEYNVEQDV 4488
Score = 20.8 bits (42), Expect(2) = 2e-05
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = +2
Query: 249 *REF*DT*KELSILKFCTKVKRTLA*LDGLTQIIQEIHMTEKS 291
*REF*+ * + CT + + L + I E+ MTEK+
Sbjct: 4187 *REF*NM*MAPVTMGLCTVIVQIQCWLGIVMLIGLEVQMTEKA 4315
>AW201343
Length = 396
Score = 46.6 bits (109), Expect = 2e-05
Identities = 21/48 (43%), Positives = 31/48 (63%)
Frame = -3
Query: 289 EKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWL 336
+ +TS Y+ ISW S K+P+V LST EA+++A + +C* IWL
Sbjct: 166 KNNTSGYILKFLEAPISWCSKKQPVVALSTCEAKYIARSFATC*YIWL 23
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment),
partial (30%)
Length = 687
Score = 45.1 bits (105), Expect = 6e-05
Identities = 19/64 (29%), Positives = 36/64 (55%)
Frame = +2
Query: 287 MTEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHIGEG 346
M +STS Y +G +SW S K+ +V S+ EAE+ + A +C +W+++ +
Sbjct: 56 MDRRSTSGYCVFIGGNLVSWKSKKQTVVARSSAEAEYRSMAMVTCELMWIKQFLQELRFC 235
Query: 347 QEIE 350
+E++
Sbjct: 236 EELQ 247
>BU764568
Length = 420
Score = 44.3 bits (103), Expect = 1e-04
Identities = 20/48 (41%), Positives = 30/48 (61%)
Frame = +3
Query: 292 TSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRI 339
TS Y ++G ISW S K+ +V S+ EAE+ A A +C IWL+++
Sbjct: 165 TSGYCVLIGGNLISWKSKKQSVVAKSSAEAEYRAMALVTCELIWLKQL 308
>AW185460
Length = 411
Score = 43.5 bits (101), Expect = 2e-04
Identities = 20/38 (52%), Positives = 26/38 (67%)
Frame = +2
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAA 327
KSTS Y + +G+G SW+S K+ V ST EAE+VA A
Sbjct: 293 KSTSGYAFSLGSGMFSWASKKQATVAQSTAEAEYVAVA 406
>BU550037
Length = 728
Score = 41.2 bits (95), Expect = 0.001
Identities = 26/83 (31%), Positives = 40/83 (47%)
Frame = -2
Query: 141 DRFGKYEIFSWSRS*AR*SRNFHRSKQVCY*NIDKAWYGKLQYGV*PYSHW*KTSEG*N* 200
D FG E+ SW+ N H +K++C +++K +GK+Q *K +G*
Sbjct: 343 D*FGSNEVLSWNVDFPIRRWNLHFTKEICLGDLEKIPHGKMQAYCYCIGCE*KILQG*RG 164
Query: 201 EGCGCNQV*ADGWMLDVITCYKT 223
+ GC +* W V CY+T
Sbjct: 163 QPRGCICL*KSDWKFVVFVCYQT 95
>TC213445
Length = 705
Score = 40.0 bits (92), Expect = 0.002
Identities = 19/49 (38%), Positives = 30/49 (60%)
Frame = +1
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRR 338
+STS + +G+ +SW S K+ V LST EAE+++A S W+R+
Sbjct: 412 ESTSDTCHFIGSALVSWHSKKQNSVVLSTAEAEYISARSYYAQIFWMRQ 558
>TC225402 weakly similar to UP|Q6I923 (Q6I923) Copia-like retrotransposon
Hopscotch polyprotein, partial (7%)
Length = 1446
Score = 37.7 bits (86), Expect = 0.010
Identities = 19/65 (29%), Positives = 33/65 (50%), Gaps = 2/65 (3%)
Frame = +2
Query: 291 STSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI--GEGQE 348
ST Y +G + W S K +V S+ EAE+ A +C IW++++ + G Q+
Sbjct: 44 STLGYCVSIGENLVLWKSNK*NVVARSSAEAEYKAMTVATCELIWIKQLLQELKFGSTQQ 223
Query: 349 IEHSC 353
++ C
Sbjct: 224 MKLCC 238
>TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, partial
(16%)
Length = 662
Score = 37.4 bits (85), Expect = 0.013
Identities = 18/51 (35%), Positives = 31/51 (60%), Gaps = 2/51 (3%)
Frame = +3
Query: 288 TEKSTSWYVYMVGNGAISWSSMKKPIV--TLSTTEAEFVAAASCSC*GIWL 336
+ KS +WY + +G+ ISW + K+ V + S++EA++ A S +C WL
Sbjct: 138 SSKSITWYCFFLGSSLISWKAKKQNTVSRSSSSSEAKYRALTSTTCELQWL 290
>BE474381 similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial (20%)
Length = 406
Score = 34.7 bits (78), Expect = 0.084
Identities = 19/54 (35%), Positives = 30/54 (55%)
Frame = +3
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSHI 343
+STS Y VG +S S K+ +V S+ EAEF A A C +W++++ +
Sbjct: 195 RSTSGYCTFVGGNLVS*SK-KQSVVARSSAEAEFRALAHGICETLWVKKLLQEL 353
>CA935263
Length = 446
Score = 32.0 bits (71), Expect = 0.55
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = +3
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCS 330
+STS Y ++G ISW S K+ S+ EA +SC+
Sbjct: 27 RSTSGYCVLIGENLISWRSKKQNTFARSSAEAVLCHDSSCN 149
>BU548243
Length = 599
Score = 31.6 bits (70), Expect = 0.71
Identities = 16/50 (32%), Positives = 26/50 (52%)
Frame = -1
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRI 339
+ST +G ISW S K+ + S+TEAE+ + A S W++ +
Sbjct: 554 RSTLGSAIFLGPNLISWWSRKQQVTAQSSTEAEYRSIAQTSAELTWIQAL 405
>TC213365 similar to UP|Q6I8L6 (Q6I8L6) Reverse transcriptase (Fragment),
partial (10%)
Length = 440
Score = 31.2 bits (69), Expect = 0.93
Identities = 15/35 (42%), Positives = 22/35 (62%)
Frame = +3
Query: 314 VTLSTTEAEFVAAASCSC*GIWLRRITSHIGEGQE 348
V LSTTEA+++A + GIWLR + + + QE
Sbjct: 9 VALSTTEAKYMALTEAAKEGIWLRGLINDLRINQE 113
>BG840035 weakly similar to GP|9294121|dbj copia-like retrotransposable
element {Arabidopsis thaliana}, partial (3%)
Length = 804
Score = 30.0 bits (66), Expect = 2.1
Identities = 19/61 (31%), Positives = 28/61 (45%)
Frame = +2
Query: 283 QEIHMTEKSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEFVAAASCSC*GIWLRRITSH 342
+E H + T + G WSS K+ EAE+VA + IWLRRI ++
Sbjct: 134 REEHDIKSXTGLLFSALVRGRFFWSSKKQD------XEAEYVATTTAVNQAIWLRRILAN 295
Query: 343 I 343
+
Sbjct: 296 L 298
>CO983062
Length = 390
Score = 29.3 bits (64), Expect = 3.5
Identities = 14/34 (41%), Positives = 19/34 (55%)
Frame = +3
Query: 290 KSTSWYVYMVGNGAISWSSMKKPIVTLSTTEAEF 323
+STS G ISW S K+ + S+TEAE+
Sbjct: 252 RSTSRAAIFPGRNLISWWSKKQKVTARSSTEAEY 353
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.372 0.166 0.671
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,442,947
Number of Sequences: 63676
Number of extensions: 319859
Number of successful extensions: 2940
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 1536
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 1388
Number of HSP's gapped (non-prelim): 1648
length of query: 427
length of database: 12,639,632
effective HSP length: 100
effective length of query: 327
effective length of database: 6,272,032
effective search space: 2050954464
effective search space used: 2050954464
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC119414.8


