

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119414.1 + phase: 0
(489 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
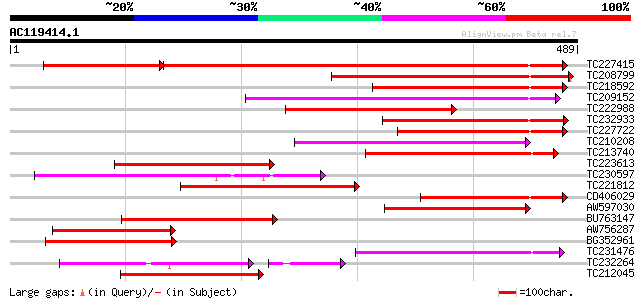
Score E
Sequences producing significant alignments: (bits) Value
TC227415 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (89%) 303 e-108
TC208799 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, pa... 214 9e-56
TC218592 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, pa... 177 1e-44
TC209152 weakly similar to PIR|B96764|B96764 protein integral me... 172 4e-43
TC222988 weakly similar to UP|Q9MAN5 (Q9MAN5) CDS, partial (29%) 161 7e-40
TC232933 similar to GB|AAN73299.1|25141209|BT002302 At3g26590/MF... 157 1e-38
TC227722 147 1e-35
TC210208 146 2e-35
TC213740 weakly similar to UP|TT12_ARATH (Q9LYT3) TRANSPARENT TE... 144 1e-34
TC223613 weakly similar to UP|Q9FNC1 (Q9FNC1) Emb|CAB89401.1, pa... 140 1e-33
TC230597 weakly similar to GB|AAP31960.1|30387589|BT006616 At1g1... 137 1e-32
TC221812 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, pa... 136 2e-32
CD406029 131 6e-31
AW597030 similar to SP|Q9LYT3|TT12 TRANSPARENT TESTA 12 protein.... 130 2e-30
BU763147 weakly similar to GP|15810018|gb AT5g38030/F16F17_30 {A... 127 1e-29
AW756287 122 4e-28
BG352961 121 8e-28
TC231476 weakly similar to GB|AAP31960.1|30387589|BT006616 At1g1... 118 7e-27
TC232264 weakly similar to UP|Q941E1 (Q941E1) At1g66760/F4N21_11... 88 2e-23
TC212045 weakly similar to UP|Q6K702 (Q6K702) MATE efflux protei... 100 1e-21
>TC227415 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (89%)
Length = 1748
Score = 303 bits (775), Expect(2) = e-108
Identities = 154/350 (44%), Positives = 222/350 (63%), Gaps = 1/350 (0%)
Frame = +2
Query: 133 LLPLFIFATPILNFFGQPQEISELAGVISMWLIPTHVTYAFFFPLYFFLQSQLKNNIIGW 192
L +++F+ P+L F G+ I+ A + LIP YA FP+ FLQ+Q +
Sbjct: 548 LTVIYVFSEPMLIFLGESPRIASAAALFVYGLIPQIFAYAANFPIQKFLQAQSIVAPSAY 727
Query: 193 VSLVALLVHVFLCWLVVVKFKLGVIALVASGNVAWIVLVFGFFGYAVLCG-CPLTWTGFS 251
+S L+VH+ + W+ V + LG++ +++W ++V G + Y V C TW GF+
Sbjct: 728 ISAATLVVHLGMSWVAVYEIGLGLLGASLVLSLSWWIMVIGQYVYIVKSERCRRTWQGFT 907
Query: 252 MEAFFDLWEFAKLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTICLTLNIWELM 311
EAF L+ F KLSAAS VMLCLE WY ++L+L+ G L N + +++L+IC T++ W M
Sbjct: 908 WEAFSGLYGFFKLSAASAVMLCLETWYFQILVLLAGLLPNPELALDSLSICTTISGWVFM 1087
Query: 312 FPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTS 371
+GF AA VRV+NELGA + ++A F+ V V S IISV L+++ R I Y FT
Sbjct: 1088ISVGFNAAASVRVSNELGARSPKSASFSVVVVTVISFIISVIAALVVLALRDVISYAFTG 1267
Query: 372 SELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLM 431
E V V+ L PLL +++LN +QPVLSGVA+G GWQ +VAY+++GCYY +G+PLG ++
Sbjct: 1268GEEVAAAVSDLCPLLALSLVLNGIQPVLSGVAVGCGWQAFVAYVNVGCYYGVGIPLGAVL 1447
Query: 432 GFVFQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLSKW 481
GF FQFG +G+W G++ GG +QT+IL WVT R DW KE E A L+KW
Sbjct: 1448GFYFQFGAKGIWLGML-GGTVMQTIILLWVTFRTDWTKEVEEAAKRLTKW 1594
Score = 107 bits (267), Expect(2) = e-108
Identities = 52/104 (50%), Positives = 74/104 (71%)
Frame = +1
Query: 30 LGRRVWNESKKLWHIAGPAIFNRVSNYSMLVITQVFAGHLGDMELAATSIAMNLILGLDL 89
+G W E K L+ +A PA+ + NY M + TQ+F+GHLG++ELAA S+ I
Sbjct: 238 VGPATWIELKLLFFLAAPAVIVYLINYLMSMSTQIFSGHLGNLELAAASLGNTGIQMFAY 417
Query: 90 GIMLGMSSALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILL 133
G+MLGM SA+ETLCGQAFGA+++ MLG+YMQRS I+L + G+++
Sbjct: 418 GLMLGMGSAVETLCGQAFGAQKYGMLGVYMQRSTILLSLAGVVV 549
>TC208799 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (46%)
Length = 924
Score = 214 bits (544), Expect = 9e-56
Identities = 101/212 (47%), Positives = 153/212 (71%), Gaps = 3/212 (1%)
Frame = +2
Query: 278 YDKVLMLMTGNLHNAKKFVEALTICLTLNIWELMFPLGFLAATGVRVANELGAGNGQAAK 337
Y+ +L+L+TGN+ NA+ ++AL+ICL +N WE+M LGF+AA VRVANELG G+ +AAK
Sbjct: 2 YNTILVLLTGNMKNAEVEIDALSICLNINGWEMMISLGFMAAASVRVANELGRGSAKAAK 181
Query: 338 FASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTSSELVIEEVNKLSPLLGFTILLNSVQP 397
F+ V+V+TS+ I ++ + FR ++ Y+FTS++ V V LSPLL +ILLNSVQP
Sbjct: 182 FSIIVSVLTSLAIGFLLFIFFLFFRERLAYIFTSNKEVAFAVGDLSPLLSVSILLNSVQP 361
Query: 398 VLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQTLI 457
VLSGVAIG+GWQ VAY+++GCYY IG+P+G ++G V V+G+W G++ G IQT++
Sbjct: 362 VLSGVAIGAGWQSIVAYVNMGCYYAIGIPVGIVLGNVLDLQVKGIWIGMLF-GTLIQTIV 538
Query: 458 LAWVTIRCDWNKEAERAKLHLSKWG---APNH 486
L +T + +W+++ A+ +S+W +P+H
Sbjct: 539 LIVITYKTNWDEQVTIAQKRISRWSKVDSPDH 634
>TC218592 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (37%)
Length = 715
Score = 177 bits (448), Expect = 1e-44
Identities = 83/168 (49%), Positives = 122/168 (72%)
Frame = +3
Query: 314 LGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTSSE 373
LGF+AA VRVANELG G+ +AAKF+ V V+TS+ I +L + R ++ Y+FTS++
Sbjct: 6 LGFMAAASVRVANELGKGSSKAAKFSIVVTVLTSLAIGFVLFLFFLFLRGKLAYIFTSNK 185
Query: 374 LVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGF 433
V + V LSPLL +ILLNSVQPVLSGVAIG+GWQ VAY+++GCYY+IG+P+G ++G
Sbjct: 186 DVADAVGDLSPLLAISILLNSVQPVLSGVAIGAGWQSIVAYVNIGCYYIIGIPVGVVLGN 365
Query: 434 VFQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLSKW 481
V V+G+W G++ G IQT++L +T + DW+++ +A+ ++KW
Sbjct: 366 VLNLQVKGIWIGMLF-GTFIQTVVLTVITYKTDWDEQVTKARNRINKW 506
>TC209152 weakly similar to PIR|B96764|B96764 protein integral membrane
protein F25P22.12 [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (41%)
Length = 1166
Score = 172 bits (435), Expect = 4e-43
Identities = 100/273 (36%), Positives = 149/273 (53%), Gaps = 1/273 (0%)
Frame = +1
Query: 204 LCWLVVVKFKLGVI-ALVASGNVAWIVLVFGFFGYAVLCGCPLTWTGFSMEAFFDLWEFA 262
+CW +V K LG + A +A G W+ +V+ C T FS A + EF
Sbjct: 1 ICWGLVFKLGLGHVGAALAIGVSYWLNVVWLAIYMIYSPACQKTKIVFSSNALLSIPEFL 180
Query: 263 KLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTICLTLNIWELMFPLGFLAATGV 322
KL+ SG+M C E W +VL L+ G L N + L+ICL P A+
Sbjct: 181 KLAIPSGLMFCFEWWSFEVLTLLAGILPNPQLETAVLSICLNTTTLHYFIPYAVGASAST 360
Query: 323 RVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTSSELVIEEVNKL 382
RV+NELGAGN + AK A V V+ + + + + R +GY +++ + VI+ V ++
Sbjct: 361 RVSNELGAGNPKTAKGAVRVVVILGVAEAAIVSTVFISCRHVLGYAYSNDKEVIDYVAEM 540
Query: 383 SPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGL 442
+PLL ++ +S+ LSG+A G G+Q+ AY++LG YYL+G+P+G L+GF Q +GL
Sbjct: 541 APLLCVSVTADSLIGALSGIARGGGFQEIGAYVNLGAYYLVGIPMGLLLGFHLQLRAKGL 720
Query: 443 WAGLVCGGPAIQTLILAWVTIRCDWNKEAERAK 475
W G + G Q +ILA VT DW KEA +A+
Sbjct: 721 WMGTL-SGSLTQVIILAIVTALIDWQKEATKAR 816
>TC222988 weakly similar to UP|Q9MAN5 (Q9MAN5) CDS, partial (29%)
Length = 474
Score = 161 bits (407), Expect = 7e-40
Identities = 78/148 (52%), Positives = 103/148 (68%), Gaps = 1/148 (0%)
Frame = +2
Query: 239 VLCG-CPLTWTGFSMEAFFDLWEFAKLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVE 297
+ CG CP TW GFS+ AF DLW AKLS +SG MLCLE WY +L+L+TGN+ NA+ ++
Sbjct: 29 ITCGWCPETWKGFSVLAFKDLWPVAKLSISSGAMLCLEFWYSTILILLTGNMKNAEVQID 208
Query: 298 ALTICLTLNIWELMFPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLL 357
AL+IC+ +N WE+M GF+AA VRVANELG G+ +AAKF+ V V+TS +I +LL
Sbjct: 209 ALSICININGWEMMIAFGFMAAASVRVANELGRGSSKAAKFSIVVTVLTSFVIGFILFLL 388
Query: 358 IMIFRRQIGYLFTSSELVIEEVNKLSPL 385
+ R ++ YL TS+E V V LSPL
Sbjct: 389 FLFVREKVAYLSTSNEDVATAVGDLSPL 472
>TC232933 similar to GB|AAN73299.1|25141209|BT002302 At3g26590/MFE16_11
{Arabidopsis thaliana;} , partial (30%)
Length = 545
Score = 157 bits (396), Expect = 1e-38
Identities = 72/161 (44%), Positives = 112/161 (68%)
Frame = +1
Query: 322 VRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTSSELVIEEVNK 381
VRV+NELGA + + AKF+ VA++TS +I V ++++IFR + F++ V V +
Sbjct: 58 VRVSNELGASHPRTAKFSLLVAMITSTLIGVMLSMVLIIFRNHYPFFFSNDSEVRAMVVE 237
Query: 382 LSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEG 441
L+P+L I++N+VQPVLSGVA+G+GWQ VAY+++ CYYL G+PLG ++G+ GV G
Sbjct: 238 LTPMLALCIVINNVQPVLSGVAVGAGWQALVAYVNIACYYLFGIPLGLVLGYKLDMGVMG 417
Query: 442 LWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHLSKWG 482
+W+G++ G +QT +L ++ R DWNKEA A+ + +WG
Sbjct: 418 IWSGML-SGTILQTCVLFFLVYRTDWNKEASLAEDRIKQWG 537
>TC227722
Length = 676
Score = 147 bits (371), Expect = 1e-35
Identities = 71/147 (48%), Positives = 99/147 (67%)
Frame = +1
Query: 335 AAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTSSELVIEEVNKLSPLLGFTILLNS 394
AAK++ V V S+ + +FF +I+ R +FT+SE++ + V KL LL T++LNS
Sbjct: 4 AAKYSFYVIVFQSLFLGIFFMAIILATRDYYAIIFTNSEVLHKAVAKLGYLLAVTMVLNS 183
Query: 395 VQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQ 454
VQPV+SGVAIG GWQ VAYI++GCYYL G+PLGFL+G+ GVEGLW G++C G IQ
Sbjct: 184 VQPVVSGVAIGGGWQALVAYINIGCYYLFGLPLGFLLGYEANLGVEGLWGGMIC-GIVIQ 360
Query: 455 TLILAWVTIRCDWNKEAERAKLHLSKW 481
TL+L + + +W KE E+ + W
Sbjct: 361 TLLLLLILYKTNWKKEVEQTTERMRIW 441
>TC210208
Length = 743
Score = 146 bits (368), Expect = 2e-35
Identities = 75/204 (36%), Positives = 116/204 (56%)
Frame = +2
Query: 246 TWTGFSMEAFFDLWEFAKLSAASGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTICLTL 305
TW GFS +F ++ +L+ S M+CLE W +VL+ + G + +++ + IC+
Sbjct: 32 TWKGFSTHSFRYVFTNMRLALPSAAMVCLEYWAFEVLVFLAGLMPDSQITTSLIAICINT 211
Query: 306 NIWELMFPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQI 365
M G AA RV+NELGAGN + AK A +V + S+++ + F L +
Sbjct: 212 EFIAYMITYGLSAAASTRVSNELGAGNPERAKHAMSVTLKLSLLLGLCFVLALGFGHNIW 391
Query: 366 GYLFTSSELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGM 425
F+ S + +E ++PLL +ILL+++Q VLSGV+ G GWQ AYI+L +YLIG+
Sbjct: 392 IQFFSDSSTIKKEFASVTPLLAISILLDAIQGVLSGVSRGCGWQHLAAYINLATFYLIGL 571
Query: 426 PLGFLMGFVFQFGVEGLWAGLVCG 449
P+ +GF +GLW GL+CG
Sbjct: 572 PISCFLGFKTNLQYKGLWIGLICG 643
>TC213740 weakly similar to UP|TT12_ARATH (Q9LYT3) TRANSPARENT TESTA 12
protein, partial (20%)
Length = 733
Score = 144 bits (362), Expect = 1e-34
Identities = 70/166 (42%), Positives = 104/166 (62%)
Frame = +3
Query: 308 WELMFPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGY 367
W+ M LG A VRV+N LG + +AA ++ V + S+++ + F +I + +
Sbjct: 15 WDDMLRLGINTAISVRVSNTLGMSHPRAAIYSFCVTMFQSLLLGILFMTVIFFSKDEFAK 194
Query: 368 LFTSSELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPL 427
+FT SE +I L+ LLG TI+LNS V+SGVAIGSGWQ V YI+L CYY++G+P+
Sbjct: 195 IFTDSEDMILAAADLAYLLGVTIVLNSASQVMSGVAIGSGWQVMVGYINLACYYIVGLPI 374
Query: 428 GFLMGFVFQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAER 473
G +GF GV+GLW G +CG +QTL+L + + +W+KE E+
Sbjct: 375 GIFLGFKLHLGVKGLWGGTMCGS-ILQTLVLFTIIWKTNWSKEVEQ 509
>TC223613 weakly similar to UP|Q9FNC1 (Q9FNC1) Emb|CAB89401.1, partial (15%)
Length = 460
Score = 140 bits (353), Expect = 1e-33
Identities = 68/139 (48%), Positives = 98/139 (69%), Gaps = 1/139 (0%)
Frame = +1
Query: 91 IMLGMSSALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPLFIFATPILNFFGQP 150
I+LGM+ AL TLCGQA+GAK+++M+G+Y+QRSWIVLF++ I LLPLFIF +PIL GQ
Sbjct: 7 ILLGMAIALSTLCGQAYGAKEYDMMGVYLQRSWIVLFLSAICLLPLFIFTSPILTLLGQD 186
Query: 151 QEISELAGVISMWLIPTHVTYAFFFPLYFFLQSQLKNNIIGWVSLVALLVHVFLCWLVVV 210
+ I+++A IS+W IP Y FLQSQ KN II +++ +++++HV L WL +
Sbjct: 187 ESIAQVARTISIWSIPVLFAYIVSNSCQTFLQSQSKNVIISYLAALSIIIHVSLSWLFTM 366
Query: 211 KFKLGVIALVASGNVA-WI 228
+FK G+ + S +A WI
Sbjct: 367 QFKYGIPGAMISTILAYWI 423
>TC230597 weakly similar to GB|AAP31960.1|30387589|BT006616 At1g15170
{Arabidopsis thaliana;} , partial (51%)
Length = 1061
Score = 137 bits (344), Expect = 1e-32
Identities = 87/256 (33%), Positives = 138/256 (52%), Gaps = 5/256 (1%)
Frame = +3
Query: 22 IDEHENEVLGRRVWNES-KKLWHIAGPAIFNRVSNYSMLVITQVFAGHLGDMELAATSIA 80
+ +HE E + V++E +++ HIAGP + S Y + V++ + GHLG++ L++ ++A
Sbjct: 27 VKKHEQERVTWGVYSEEMRRVCHIAGPMVAVVSSQYLLQVVSTMIVGHLGELYLSSAALA 206
Query: 81 MNLILGLDLGIMLGMSSALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPLFIFA 140
++L +++GM+S LET+CGQA+G +Q+ +GI + L + I + L+I
Sbjct: 207 ISLSGVTGFSLLMGMASGLETICGQAYGGQQYQRIGIQTYTAIFSLILVSIPVSLLWINM 386
Query: 141 TPILNFFGQPQEISELAGVISMWLIPTHVTYAFFFPL--YFFLQSQLKNNIIGWVSLVAL 198
IL F GQ IS AG ++WL+P YA PL YF +QS L S V L
Sbjct: 387 ETILVFIGQDPLISHEAGKFTIWLVPALFAYAILQPLVRYFQVQSLLLPMFAS--SCVTL 560
Query: 199 LVHVFLCWLVVVKFKLGVI--ALVASGNVAWIVLVFGFFGYAVLCGCPLTWTGFSMEAFF 256
++HV LCW +V K +L + AL S ++ W ++F C T SME F
Sbjct: 561 IIHVPLCWALVFKTRLSNVGGALAVSISI-WSNVIFLGLYMRYFSACAKTRAPISMELFK 737
Query: 257 DLWEFAKLSAASGVML 272
+WEF + + S VM+
Sbjct: 738 GMWEFFRFAIPSAVMV 785
>TC221812 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (37%)
Length = 470
Score = 136 bits (343), Expect = 2e-32
Identities = 66/154 (42%), Positives = 100/154 (64%)
Frame = +3
Query: 148 GQPQEISELAGVISMWLIPTHVTYAFFFPLYFFLQSQLKNNIIGWVSLVALLVHVFLCWL 207
GQ + I+E+AG IS+W IP + F FLQSQ KN II +++ ++++H+FL WL
Sbjct: 9 GQDENIAEVAGNISLWSIPMIFAFIASFTCQNFLQSQSKNTIISFLAAFSIVIHLFLSWL 188
Query: 208 VVVKFKLGVIALVASGNVAWIVLVFGFFGYAVLCGCPLTWTGFSMEAFFDLWEFAKLSAA 267
+ ++FKL + + S N+A+ + G + C TW GFS AF DLW KLS +
Sbjct: 189 LTIQFKLEIPGAMTSTNLAFWIPNIGQLIFITCGWCSDTWKGFSFLAFKDLWPVVKLSLS 368
Query: 268 SGVMLCLEVWYDKVLMLMTGNLHNAKKFVEALTI 301
SG+MLCLE+WY+ +L+L+TGN+ NA+ ++AL+I
Sbjct: 369 SGIMLCLELWYNTILVLLTGNMENAEVQIDALSI 470
>CD406029
Length = 464
Score = 131 bits (330), Expect = 6e-31
Identities = 59/127 (46%), Positives = 92/127 (71%)
Frame = -2
Query: 355 WLLIMIFRRQIGYLFTSSELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAY 414
++L +I R ++ YL TS+E V+ V LSPLL ++LLNS+QPVLSGVA+G GWQ VAY
Sbjct: 463 FVLFLILREKVAYLLTSNEDVVTAVGDLSPLLALSLLLNSIQPVLSGVALGPGWQSTVAY 284
Query: 415 IDLGCYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERA 474
+++GCYYLIG+P+G ++G + V+G+W G++ G IQT+IL +T + +W+++ A
Sbjct: 283 VNIGCYYLIGIPVGIVLGNIIHLQVKGIWIGMLF-GTLIQTIILIIITYKTNWDEQVIIA 107
Query: 475 KLHLSKW 481
+ ++KW
Sbjct: 106 RDRINKW 86
>AW597030 similar to SP|Q9LYT3|TT12 TRANSPARENT TESTA 12 protein. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (25%)
Length = 401
Score = 130 bits (326), Expect = 2e-30
Identities = 62/126 (49%), Positives = 88/126 (69%)
Frame = +1
Query: 324 VANELGAGNGQAAKFASAVAVVTSIIISVFFWLLIMIFRRQIGYLFTSSELVIEEVNKLS 383
V+NELGA + + AKF+ V TSI+ISV F +I+IFR + LFTS VI+ V+ L+
Sbjct: 10 VSNELGASHPRVAKFSVFVVNGTSILISVVFCTIILIFRVSLSKLFTSDSDVIDAVSNLT 189
Query: 384 PLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLGCYYLIGMPLGFLMGFVFQFGVEGLW 443
PLL ++ N +QP+LSGVAIGSGWQ VAY++L YY++G+ +G ++GF GV G+W
Sbjct: 190 PLLAISVFFNGIQPILSGVAIGSGWQALVAYVNLASYYVVGLTVGCVLGFKTSLGVAGIW 369
Query: 444 AGLVCG 449
G++ G
Sbjct: 370 WGMILG 387
>BU763147 weakly similar to GP|15810018|gb AT5g38030/F16F17_30 {Arabidopsis
thaliana}, partial (46%)
Length = 429
Score = 127 bits (318), Expect = 1e-29
Identities = 62/135 (45%), Positives = 89/135 (65%)
Frame = -3
Query: 97 SALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPLFIFATPILNFFGQPQEISEL 156
SALETLCGQA GA + +MLG+YMQRSW++L T +L PL+IFA +L GQ EISE
Sbjct: 424 SALETLCGQAVGAGKLDMLGVYMQRSWVLLLSTACVLCPLYIFAGQVLKLIGQDTEISEA 245
Query: 157 AGVISMWLIPTHVTYAFFFPLYFFLQSQLKNNIIGWVSLVALLVHVFLCWLVVVKFKLGV 216
AG ++W+IP YA FP+ FLQ+Q K +I ++ +A+++H L WL++VK + G+
Sbjct: 244 AGTFAIWMIPQLFAYALNFPVAKFLQAQSKVMVIAAIAGMAMVLHPVLSWLLMVKLEWGL 65
Query: 217 IALVASGNVAWIVLV 231
+ N +W +V
Sbjct: 64 VGAAVVLNGSWWFVV 20
>AW756287
Length = 319
Score = 122 bits (306), Expect = 4e-28
Identities = 55/106 (51%), Positives = 80/106 (74%)
Frame = +2
Query: 38 SKKLWHIAGPAIFNRVSNYSMLVITQVFAGHLGDMELAATSIAMNLILGLDLGIMLGMSS 97
SK +W +A PAIF + + + + VI+Q F GH+G ELAA ++ +I+ GI+LGMSS
Sbjct: 2 SKVMWIVAAPAIFTKFTTFGLSVISQAFIGHIGSKELAAYALVFTVIIRFANGILLGMSS 181
Query: 98 ALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPLFIFATPI 143
AL TLCGQA+GAK+++M+G+Y+QRS IV+F+T + LLPL I+ +PI
Sbjct: 182 ALSTLCGQAYGAKEYDMMGVYLQRSSIVIFLTALCLLPLIIYTSPI 319
>BG352961
Length = 496
Score = 121 bits (303), Expect = 8e-28
Identities = 60/113 (53%), Positives = 80/113 (70%)
Frame = +2
Query: 32 RRVWNESKKLWHIAGPAIFNRVSNYSMLVITQVFAGHLGDMELAATSIAMNLILGLDLGI 91
R + ESKKLW++ GPAIF V YS+ +TQ+F+ H+ + LA S+ ++I G LGI
Sbjct: 38 REFFAESKKLWYLDGPAIFTCVCQYSLGGVTQLFSRHVNTLTLAVVSVNDSVIAGYSLGI 217
Query: 92 MLGMSSALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPLFIFATPIL 144
GM SALETLCGQ +GA Q +MLG+YM+RSW++L TGILL L+IFA +L
Sbjct: 218 TFGMGSALETLCGQPYGAGQVHMLGVYMRRSWVILSATGILLSLLYIFAGHML 376
>TC231476 weakly similar to GB|AAP31960.1|30387589|BT006616 At1g15170
{Arabidopsis thaliana;} , partial (32%)
Length = 734
Score = 118 bits (295), Expect = 7e-27
Identities = 60/180 (33%), Positives = 104/180 (57%)
Frame = +2
Query: 299 LTICLTLNIWELMFPLGFLAATGVRVANELGAGNGQAAKFASAVAVVTSIIISVFFWLLI 358
L+ICL P G AA R++NELGAGN AA A A+ +I+ + +
Sbjct: 8 LSICLNTISTLFSIPFGIAAAASTRISNELGAGNPHAAHVAVLAAMSFAIMETAIVSGTL 187
Query: 359 MIFRRQIGYLFTSSELVIEEVNKLSPLLGFTILLNSVQPVLSGVAIGSGWQKYVAYIDLG 418
+ R GY+F++ + V++ V ++PL+ +++L+S+Q VL+GVA G GWQ Y++LG
Sbjct: 188 FVCRHDFGYIFSNEKEVVDYVTVMAPLICISVILDSIQGVLAGVARGCGWQHIGVYVNLG 367
Query: 419 CYYLIGMPLGFLMGFVFQFGVEGLWAGLVCGGPAIQTLILAWVTIRCDWNKEAERAKLHL 478
+YL G+P+ + F+ + +GLW G+ G +Q ++ + +T +W ++A +A+ L
Sbjct: 368 AFYLCGIPVAATLAFLAKMRGKGLWIGVQVGA-FVQCILFSTITSCINWEQQAIKARKRL 544
>TC232264 weakly similar to UP|Q941E1 (Q941E1) At1g66760/F4N21_11, partial
(58%)
Length = 701
Score = 88.2 bits (217), Expect(2) = 2e-23
Identities = 55/171 (32%), Positives = 89/171 (51%), Gaps = 4/171 (2%)
Frame = +1
Query: 44 IAGPAIFNRVSNYSMLVITQVFAGHLGDM-ELAATSIAMNLILGLDLGIMLGMSSALETL 102
+A P + VS Y + V++ + GH G + + +IA + ++LGMS ALETL
Sbjct: 10 MAAPMVAVTVSQYLLQVVSLMMVGHFGILVSFSGVAIATSFAEVTGFSVLLGMSGALETL 189
Query: 103 CGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPL---FIFATPILNFFGQPQEISELAGV 159
CGQ +GA+++ G Y +W + ++ LP+ +IF IL F Q EIS A
Sbjct: 190 CGQTYGAEEYRKFGNY---TWCAIVTLTLVCLPISLVWIFTDKILLLFSQDPEISHAARE 360
Query: 160 ISMWLIPTHVTYAFFFPLYFFLQSQLKNNIIGWVSLVALLVHVFLCWLVVV 210
++LIP +A L + Q+Q + + S+ AL +HV +CW +V+
Sbjct: 361 YCIYLIPALFGHAVLQALTRYFQTQSMIFPMVFSSITALCLHVPICWSLVL 513
Score = 39.3 bits (90), Expect(2) = 2e-23
Identities = 23/66 (34%), Positives = 31/66 (46%)
Frame = +2
Query: 224 NVAWIVLVFGFFGYAVLCGCPLTWTGFSMEAFFDLWEFAKLSAASGVMLCLEVWYDKVLM 283
NV W+ + + C T FS A + EF KL+ SG+M C E W +VL
Sbjct: 518 NVVWLAIYMIYSP-----ACQKTKIVFSSNALLSIPEFLKLAIPSGLMFCFEWWSFEVLT 682
Query: 284 LMTGNL 289
L+ G L
Sbjct: 683 LLAGIL 700
>TC212045 weakly similar to UP|Q6K702 (Q6K702) MATE efflux protein-like,
partial (20%)
Length = 415
Score = 100 bits (250), Expect = 1e-21
Identities = 53/124 (42%), Positives = 79/124 (62%)
Frame = +1
Query: 96 SSALETLCGQAFGAKQFNMLGIYMQRSWIVLFITGILLLPLFIFATPILNFFGQPQEISE 155
SSAL TLCGQAFGA + IY+QRSWI+L T I+L P++++ATPIL GQ + I+E
Sbjct: 40 SSALATLCGQAFGAGKMQSTCIYVQRSWIILTATWIILWPIYVYATPILKLLGQDEGIAE 219
Query: 156 LAGVISMWLIPTHVTYAFFFPLYFFLQSQLKNNIIGWVSLVALLVHVFLCWLVVVKFKLG 215
+A S+ +IP ++A FP+ FLQ+Q K +I ++ V LL+ L ++ + + G
Sbjct: 220 VARRYSIQVIPYMFSFAVAFPIQRFLQAQSKVKVIMCIAFVDLLIQNGLLYIFINVYGWG 399
Query: 216 VIAL 219
+ L
Sbjct: 400 ITGL 411
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.142 0.451
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,347,422
Number of Sequences: 63676
Number of extensions: 414071
Number of successful extensions: 3118
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 3060
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3091
length of query: 489
length of database: 12,639,632
effective HSP length: 101
effective length of query: 388
effective length of database: 6,208,356
effective search space: 2408842128
effective search space used: 2408842128
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC119414.1


