

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119412.5 + phase: 0
(221 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
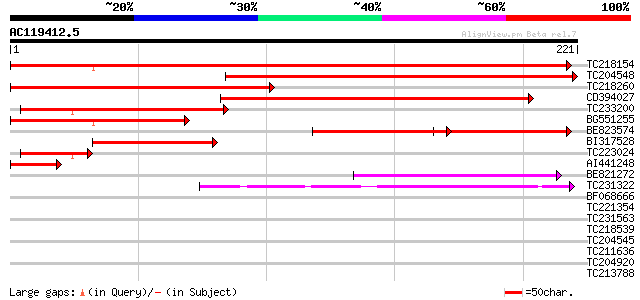
TC218154 similar to GB|AAF87026.1|9295720|AC006535 T24P13.5 {Ara... 291 1e-79
TC204548 similar to UP|VT13_ARATH (Q9LVP9) Vesicle transport v-S... 241 1e-64
TC218260 similar to UP|VT13_ARATH (Q9LVP9) Vesicle transport v-S... 195 2e-50
CD394027 169 1e-42
TC233200 similar to GB|AAF87026.1|9295720|AC006535 T24P13.5 {Ara... 125 2e-29
BG551255 similar to GP|9295720|gb|A T24P13.5 {Arabidopsis thalia... 102 1e-22
BE823574 78 3e-15
BI317528 78 4e-15
TC223024 similar to GB|AAF24061.1|6690274|AF114750 v-SNARE AtVTI... 47 7e-06
AI441248 similar to GP|6690276|gb|A v-SNARE AtVTI1b {Arabidopsis... 44 6e-05
BE821272 weakly similar to SP|Q9SJL6|ME11 Membrin 11 (AtMEMB11) ... 43 1e-04
TC231322 similar to GB|AAC61818.2|20197215|AC004667 expressed pr... 42 2e-04
BF068666 30 0.66
TC221354 similar to UP|SY32_ARATH (Q9LK09) Syntaxin 32 (AtSYP32)... 30 0.66
TC231563 similar to UP|S121_ARATH (Q9ZSD4) Syntaxin 121 (AtSYP12... 30 0.86
TC218539 29 1.5
TC204545 homologue to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, c... 29 1.5
TC211636 28 3.3
TC204920 similar to UP|PPF1_PEA (Q9FY06) Inner membrane protein ... 28 3.3
TC213788 similar to GB|AAN73298.1|25141207|BT002301 At4g16430/dl... 28 4.2
>TC218154 similar to GB|AAF87026.1|9295720|AC006535 T24P13.5 {Arabidopsis
thaliana;} , partial (90%)
Length = 965
Score = 291 bits (746), Expect = 1e-79
Identities = 139/220 (63%), Positives = 188/220 (85%), Gaps = 1/220 (0%)
Frame = +1
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNG-EQKKQKVSEVKSGIDEAEALIRKMDLEA 59
MS VFEGYERQYCELSANLS+KC++A+ ++G EQ++QK+SE+K+G+D+A+ LIRKMDLEA
Sbjct: 166 MSEVFEGYERQYCELSANLSRKCSSASLVSGQEQQQQKLSEIKAGLDDADVLIRKMDLEA 345
Query: 60 RSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQ 119
RS+QP++K +LLAKLREYKSDL NLK E K++ S N + AR+ELLE+GMA+T ASADQ
Sbjct: 346 RSLQPSVKAMLLAKLREYKSDLTNLKKEFKRLTSPNADEVAREELLETGMANTHLASADQ 525
Query: 120 RTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNI 179
R RL S ER+N++GER+++S RT+LETEELG++I+QDLHSQR++LL++H LHG+DD I
Sbjct: 526 RERLTMSVERINQSGERIRESHRTLLETEELGINIIQDLHSQRETLLNSHKRLHGIDDAI 705
Query: 180 GKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
KSKK++T MSRR+ +NKWI+ ++ LV AI+ IL++KL
Sbjct: 706 DKSKKVLTTMSRRITRNKWIVASVIGALVFAIVIILFYKL 825
>TC204548 similar to UP|VT13_ARATH (Q9LVP9) Vesicle transport v-SNARE 13
(AtVTI13) (Vesicle transport v-SNARE protein VTI13)
(Vesicle soluble NSF attachment protein receptor 13),
partial (61%)
Length = 657
Score = 241 bits (616), Expect = 1e-64
Identities = 124/137 (90%), Positives = 128/137 (92%)
Frame = +2
Query: 85 KSEVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERLNKTGERVKDSRRTM 144
KSEVKKIVSGNLN ARDELLESGMAD MTASADQRTRLM STERLNKT +RVKDSRRTM
Sbjct: 2 KSEVKKIVSGNLNXXARDELLESGMADAMTASADQRTRLMVSTERLNKTSDRVKDSRRTM 181
Query: 145 LETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIV 204
LETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKI+TNMSRRMNKNKW+IG IV
Sbjct: 182 LETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKILTNMSRRMNKNKWVIGGIV 361
Query: 205 LVLVVAIIAILYFKLVK 221
LVLV+AII ILYFK K
Sbjct: 362 LVLVIAIIVILYFKFSK 412
>TC218260 similar to UP|VT13_ARATH (Q9LVP9) Vesicle transport v-SNARE 13
(AtVTI13) (Vesicle transport v-SNARE protein VTI13)
(Vesicle soluble NSF attachment protein receptor 13),
partial (47%)
Length = 523
Score = 195 bits (495), Expect = 2e-50
Identities = 99/103 (96%), Positives = 102/103 (98%)
Frame = +3
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEAR 60
MSNVFEGYERQYCELSANL+KKCTAA ALNGEQKKQKVSEVK+GIDEAEALIRKMDLEAR
Sbjct: 213 MSNVFEGYERQYCELSANLAKKCTAAGALNGEQKKQKVSEVKAGIDEAEALIRKMDLEAR 392
Query: 61 SMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDE 103
S+QPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDE
Sbjct: 393 SLQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDE 521
>CD394027
Length = 366
Score = 169 bits (427), Expect = 1e-42
Identities = 79/122 (64%), Positives = 105/122 (85%)
Frame = -1
Query: 83 NLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERLNKTGERVKDSRR 142
NLKSEVK++ S ++N +ARD+LLESG ADT+ AS DQ+ RL+ STERLN++ +R+K+SR+
Sbjct: 366 NLKSEVKRVTSASVNLTARDDLLESGRADTLAASNDQKGRLLMSTERLNQSSDRIKESRK 187
Query: 143 TMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGC 202
TMLETE+LG IL+DLH QR+SLLHAH T+HGVDDNI KSKKI++ MSRRM++NKWI+
Sbjct: 186 TMLETEDLGEFILRDLHQQRESLLHAHKTIHGVDDNISKSKKILSAMSRRMSRNKWIVSS 7
Query: 203 IV 204
++
Sbjct: 6 LM 1
>TC233200 similar to GB|AAF87026.1|9295720|AC006535 T24P13.5 {Arabidopsis
thaliana;} , partial (31%)
Length = 444
Score = 125 bits (313), Expect = 2e-29
Identities = 62/83 (74%), Positives = 71/83 (84%), Gaps = 2/83 (2%)
Frame = +2
Query: 5 FEGYERQYCELSANLSKKC--TAATALNGEQKKQKVSEVKSGIDEAEALIRKMDLEARSM 62
FEGYERQYCELSANLSK C A LNGE KKQK SE+K GI+E EALIRKMDL+ARS+
Sbjct: 191 FEGYERQYCELSANLSKACIDNVAAPLNGELKKQKKSEIKEGIEEGEALIRKMDLDARSL 370
Query: 63 QPNIKGVLLAKLREYKSDLNNLK 85
QP++K VLLAK+REYK+DLNN+K
Sbjct: 371 QPDLKAVLLAKVREYKADLNNIK 439
>BG551255 similar to GP|9295720|gb|A T24P13.5 {Arabidopsis thaliana}, partial
(31%)
Length = 372
Score = 102 bits (254), Expect = 1e-22
Identities = 48/71 (67%), Positives = 64/71 (89%), Gaps = 1/71 (1%)
Frame = +1
Query: 1 MSNVFEGYERQYCELSANLSKKCTAATALNG-EQKKQKVSEVKSGIDEAEALIRKMDLEA 59
MS VFEGYERQYCELSANLS+KC++A+ ++ EQK QK+SE+K+G+D+A+ LIRKMDLEA
Sbjct: 160 MSEVFEGYERQYCELSANLSRKCSSASLVSDQEQKPQKLSEIKAGLDDADVLIRKMDLEA 339
Query: 60 RSMQPNIKGVL 70
RS+QP++K +L
Sbjct: 340 RSLQPSVKAML 372
>BE823574
Length = 617
Score = 78.2 bits (191), Expect = 3e-15
Identities = 37/54 (68%), Positives = 44/54 (80%)
Frame = -1
Query: 119 QRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTL 172
+R RLM S RLNK+ +R+ DSR MLETE LG+SILQDLHSQRQSLLH H+T+
Sbjct: 611 ERERLMISXXRLNKSSDRINDSRGXMLETEXLGISILQDLHSQRQSLLHTHDTV 450
Score = 69.7 bits (169), Expect = 1e-12
Identities = 32/54 (59%), Positives = 43/54 (79%)
Frame = -3
Query: 166 LHAHNTLHGVDDNIGKSKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILYFKL 219
L++ LHGVDDN KSKKI++NMSRRM+K+KWI+ I ++L+ II I+YFKL
Sbjct: 180 LYSIMQLHGVDDNTDKSKKILSNMSRRMDKSKWILSTIAVLLIFVIILIVYFKL 19
>BI317528
Length = 384
Score = 77.8 bits (190), Expect = 4e-15
Identities = 35/49 (71%), Positives = 47/49 (95%)
Frame = +3
Query: 33 QKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDL 81
Q++QK+SE+K+G+D+A+ LIRKMDLEARS+QP++K +LLAKLREYKSDL
Sbjct: 225 QQQQKLSEIKAGLDDADVLIRKMDLEARSLQPSVKAMLLAKLREYKSDL 371
>TC223024 similar to GB|AAF24061.1|6690274|AF114750 v-SNARE AtVTI1a
{Arabidopsis thaliana;} , partial (9%)
Length = 156
Score = 47.0 bits (110), Expect = 7e-06
Identities = 23/30 (76%), Positives = 23/30 (76%), Gaps = 2/30 (6%)
Frame = +3
Query: 5 FEGYERQYCELSANLSKKC--TAATALNGE 32
FEGYERQYCELSANLSK C A LNGE
Sbjct: 63 FEGYERQYCELSANLSKACIDNVAAPLNGE 152
>AI441248 similar to GP|6690276|gb|A v-SNARE AtVTI1b {Arabidopsis thaliana},
partial (8%)
Length = 219
Score = 43.9 bits (102), Expect = 6e-05
Identities = 19/20 (95%), Positives = 20/20 (100%)
Frame = +3
Query: 1 MSNVFEGYERQYCELSANLS 20
MSNVFEGYERQYCELSANL+
Sbjct: 159 MSNVFEGYERQYCELSANLA 218
>BE821272 weakly similar to SP|Q9SJL6|ME11 Membrin 11 (AtMEMB11) (Golgi SNAP
receptor complex member 2-1) (27 kDa Golgi SNARE
protein)., partial (68%)
Length = 510
Score = 42.7 bits (99), Expect = 1e-04
Identities = 21/81 (25%), Positives = 42/81 (50%)
Frame = -3
Query: 135 ERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMN 194
+ V+ S + + LG +IL +H QR+ L AH + + +G S ++ + RR
Sbjct: 292 QSVRSSSQELENANALGEAILSSIHGQRERLKSAHRKALDILNTVGISNSVLXLIERRNR 113
Query: 195 KNKWIIGCIVLVLVVAIIAIL 215
++WI +L+ V+ ++A +
Sbjct: 112 VDQWIKYAGMLLTVIFLLAFI 50
>TC231322 similar to GB|AAC61818.2|20197215|AC004667 expressed protein
{Arabidopsis thaliana;} , partial (58%)
Length = 936
Score = 42.4 bits (98), Expect = 2e-04
Identities = 33/146 (22%), Positives = 72/146 (48%)
Frame = +2
Query: 75 REYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERLNKTG 134
++Y +++ N + E+ + N + + LL S M + D R+M T++ + G
Sbjct: 35 KQYATNIENKRIELFE--GPNEGYTEENGLLASSMTNEQLM--DHGNRMMDETDQAIERG 202
Query: 135 ERVKDSRRTMLETEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIMTNMSRRMN 194
++V + +T +G L +Q + + N L + ++ K+ K++ + R++
Sbjct: 203 KKV------VQDTINVGTDTAAALKAQTEQMSRVVNELDSIHFSMKKASKLVKEIGRQVA 364
Query: 195 KNKWIIGCIVLVLVVAIIAILYFKLV 220
+K I+ + L+ V+ +IAI+ KLV
Sbjct: 365 TDKCIMALLFLI-VIGVIAIIIVKLV 439
>BF068666
Length = 405
Score = 30.4 bits (67), Expect = 0.66
Identities = 17/78 (21%), Positives = 33/78 (41%)
Frame = +2
Query: 109 MADTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLETEELGVSILQDLHSQRQSLLHA 168
+ D+ S+D T + TER+ + + D + + L + I + Q S
Sbjct: 98 LLDSQPTSSDAPTPTQSDTERMEDSANLLPDMKLSTLSKRAASIKIKSSSNHQTFSASTQ 277
Query: 169 HNTLHGVDDNIGKSKKIM 186
HN +D ++ +K+M
Sbjct: 278 HNNKQELDASLVVIRKVM 331
>TC221354 similar to UP|SY32_ARATH (Q9LK09) Syntaxin 32 (AtSYP32), partial
(48%)
Length = 962
Score = 30.4 bits (67), Expect = 0.66
Identities = 40/211 (18%), Positives = 80/211 (36%), Gaps = 5/211 (2%)
Frame = +3
Query: 12 YCELSANLSKKCTAATALNG----EQKKQKVSEVKSGIDEAEALIRKMDLEARSMQPNIK 67
+C L AN C A + E ++Q S S D A +R+ L RS
Sbjct: 162 FCCLFANFRHWCDAIVFVKNLKVHENRRQLFSATASK-DSANPFVRQRPLATRSAASTSN 338
Query: 68 GVLLAKLREYKSDLNNLKSEVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTST 127
A + + ++ + KK V G P + + + + + R + +
Sbjct: 339 ----APAAPWATGSSSSQLFPKKQVDGESQPLLQQQQQQQEVVPLQDSYMQNRAEALQNV 506
Query: 128 ERLNKTGERVKDSRRTMLETE-ELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKKIM 186
E + + T++ + E+ + I +++ +TL V+ G K +
Sbjct: 507 ESTIHELSNIFNQLATLVSQQGEIAIRIDENMD----------DTLANVEGAQGALLKYL 656
Query: 187 TNMSRRMNKNKWIIGCIVLVLVVAIIAILYF 217
N+S N+W++ I +L+ ++ L+F
Sbjct: 657 NNIS----SNRWLMIKIFFILIFFLMVFLFF 737
>TC231563 similar to UP|S121_ARATH (Q9ZSD4) Syntaxin 121 (AtSYP121)
(Syntaxin-related protein At-Syr1), partial (22%)
Length = 681
Score = 30.0 bits (66), Expect = 0.86
Identities = 10/35 (28%), Positives = 21/35 (59%)
Frame = +2
Query: 182 SKKIMTNMSRRMNKNKWIIGCIVLVLVVAIIAILY 216
++++ T + N KW CI+L+LV+ + +L+
Sbjct: 80 AEQLQTARKHQKNTRKWTCYCIILLLVIILFVVLF 184
>TC218539
Length = 1218
Score = 29.3 bits (64), Expect = 1.5
Identities = 26/101 (25%), Positives = 46/101 (44%), Gaps = 1/101 (0%)
Frame = +2
Query: 35 KQKVSEVKSGIDEAEALIRKMDLEARSMQPNIKGVLLAKLREYKSDLNNLKSEVKKIVSG 94
K + ++KSG +E +A RKMD S P G + +S +NL+ + + +
Sbjct: 212 KDMIEDLKSGNEEGQACWRKMDCYENS--PGSSGYSTGSILICQSASDNLEHSLHQDIGT 385
Query: 95 NLNPSARDELLESGMADTM-TASADQRTRLMTSTERLNKTG 134
+ ++ E G+ M TA+ DQ + E++N G
Sbjct: 386 EMKSKQLED--EKGVKVVMPTANVDQ-DMVPCCMEQINNDG 499
>TC204545 homologue to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, complete
Length = 1924
Score = 29.3 bits (64), Expect = 1.5
Identities = 13/18 (72%), Positives = 16/18 (88%)
Frame = +3
Query: 204 VLVLVVAIIAILYFKLVK 221
+LVL++AII ILYFKL K
Sbjct: 738 LLVLIIAIIVILYFKLSK 791
>TC211636
Length = 749
Score = 28.1 bits (61), Expect = 3.3
Identities = 14/23 (60%), Positives = 17/23 (73%)
Frame = +3
Query: 144 MLETEELGVSILQDLHSQRQSLL 166
MLE EE GV+I+Q+ QR SLL
Sbjct: 168 MLEGEEKGVAIVQEAKKQRVSLL 236
>TC204920 similar to UP|PPF1_PEA (Q9FY06) Inner membrane protein PPF-1,
chloroplast precursor (Post-floral-specific protein 1),
partial (77%)
Length = 1245
Score = 28.1 bits (61), Expect = 3.3
Identities = 16/57 (28%), Positives = 28/57 (49%), Gaps = 12/57 (21%)
Frame = -1
Query: 155 LQDLHSQRQSLLHAHNTLHGVDDNI------------GKSKKIMTNMSRRMNKNKWI 199
++D+H ++ L H N LHG D G S+++ +N RR+ +N+ I
Sbjct: 435 IRDMHGRKAILQHLENELHGFGDEAEPASALLYGSIGGGSRRVRSN--RRVRRNRSI 271
>TC213788 similar to GB|AAN73298.1|25141207|BT002301 At4g16430/dl4240w
{Arabidopsis thaliana;} , partial (19%)
Length = 452
Score = 27.7 bits (60), Expect = 4.2
Identities = 24/98 (24%), Positives = 39/98 (39%)
Frame = +1
Query: 87 EVKKIVSGNLNPSARDELLESGMADTMTASADQRTRLMTSTERLNKTGERVKDSRRTMLE 146
++ ++++GN N AR L+ G D+ + AD+R + K G + + R L
Sbjct: 169 QLNQMMAGNFNAQARVPCLDLGNEDSSSIHADER--------KPKKRGRKPANGREEPLN 324
Query: 147 TEELGVSILQDLHSQRQSLLHAHNTLHGVDDNIGKSKK 184
E +R+ L L V NI K K
Sbjct: 325 HVEAE-------RQRREKLNQRFYALRAVVPNISKMDK 417
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.129 0.342
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,836,155
Number of Sequences: 63676
Number of extensions: 65266
Number of successful extensions: 381
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 378
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 378
length of query: 221
length of database: 12,639,632
effective HSP length: 94
effective length of query: 127
effective length of database: 6,654,088
effective search space: 845069176
effective search space used: 845069176
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC119412.5


