

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC119408.1 + phase: 0 /pseudo
(600 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
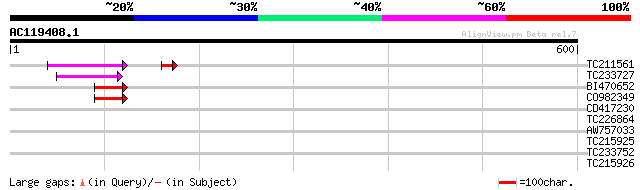
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 73 3e-13
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 59 8e-09
BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase ... 46 5e-05
CO982349 45 9e-05
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 36 0.056
TC226864 similar to UP|Q43382 (Q43382) PSI type II chlorophyll a... 32 0.62
AW757033 32 1.1
TC215925 29 6.8
TC233752 UP|Q8BTF6 (Q8BTF6) Mus musculus 16 days embryo head cDN... 29 6.8
TC215926 28 8.9
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 73.2 bits (178), Expect(2) = 3e-13
Identities = 35/85 (41%), Positives = 50/85 (58%), Gaps = 1/85 (1%)
Frame = +1
Query: 41 IRKTRTNNNGYVGIKLDIAKAYDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILI 100
I + + +N + K+D KAYDS+ WNF+ + M F I I C++ + S+L+
Sbjct: 31 IDEAKRSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEECVKSASISVLV 210
Query: 101 NGQPTEPFSPQRGIRQGDPF-PLIF 124
NG PT F PQRG+RQGDP P +F
Sbjct: 211 NGSPTAEFLPQRGLRQGDPLTPFLF 285
Score = 20.4 bits (41), Expect(2) = 3e-13
Identities = 5/17 (29%), Positives = 12/17 (70%)
Frame = +2
Query: 161 FMLMIVSYFVGPNPRRL 177
+ +++ +F+G PRR+
Sbjct: 389 YSMLMTQFFLGKQPRRM 439
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 58.5 bits (140), Expect = 8e-09
Identities = 28/70 (40%), Positives = 41/70 (58%)
Frame = -3
Query: 50 GYVGIKLDIAKAYDSLKWNFIEDTLTAMGFPINLINTIMLCIRCVTFSILINGQPTEPFS 109
G + +K+DI KA+D+L +F+ L G+ N I + + SI +NG+P FS
Sbjct: 771 GNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIRVILLSAKLSISVNGEPVGFFS 592
Query: 110 PQRGIRQGDP 119
QRG+RQGDP
Sbjct: 591 CQRGVRQGDP 562
>BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase (EC
2.7.7.49) (clone AmLi2) - garden snapdragon (fragment),
partial (22%)
Length = 111
Score = 45.8 bits (107), Expect = 5e-05
Identities = 19/35 (54%), Positives = 23/35 (65%)
Frame = +1
Query: 90 CIRCVTFSILINGQPTEPFSPQRGIRQGDPFPLIF 124
C+ + SIL+NG PTE F P RG+RQG PF F
Sbjct: 7 CLSSASISILVNGSPTEEFKPXRGLRQGGPFGPFF 111
>CO982349
Length = 795
Score = 45.1 bits (105), Expect = 9e-05
Identities = 20/36 (55%), Positives = 25/36 (68%), Gaps = 1/36 (2%)
Frame = +3
Query: 90 CIRCVTFSILINGQPTEPFSPQRGIRQGDPF-PLIF 124
C+ + SIL+N P FSPQRG+RQGDP PL+F
Sbjct: 81 CLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLF 188
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 35.8 bits (81), Expect = 0.056
Identities = 16/43 (37%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Frame = -3
Query: 84 INTIMLCIRCVTFSILINGQPTEPFSPQRGIRQGDPF-PLIFL 125
+ + C+ T S+L NG+ + FSP G+RQ DP P +F+
Sbjct: 686 VQLVWYCMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFV 558
>TC226864 similar to UP|Q43382 (Q43382) PSI type II chlorophyll a/b-binding
protein, partial (42%)
Length = 574
Score = 32.3 bits (72), Expect = 0.62
Identities = 21/53 (39%), Positives = 29/53 (54%)
Frame = +1
Query: 336 SKFLEILIWPCLPNKSRDFRLIKNPLSPEFSKLNISLTKTFSRLI*VIAQVML 388
S+ ++ WP L + S+ F L K+PL + L I +T TFSRL V A L
Sbjct: 211 SRMEDLQCWPLLGSGSKLFTLGKDPLKI*WHTLLILVTATFSRLSRVSAMYSL 369
>AW757033
Length = 441
Score = 31.6 bits (70), Expect = 1.1
Identities = 14/21 (66%), Positives = 16/21 (75%), Gaps = 1/21 (4%)
Frame = -2
Query: 105 TEPFSPQRGIRQGDPF-PLIF 124
T FSP RG+RQGDP PL+F
Sbjct: 440 TREFSPHRGLRQGDPLAPLLF 378
>TC215925
Length = 781
Score = 28.9 bits (63), Expect = 6.8
Identities = 16/58 (27%), Positives = 30/58 (51%), Gaps = 1/58 (1%)
Frame = +1
Query: 299 LRKLFVHFGGVALIVKRRFTGPKEKIFLNPDIR-VEWASKFLEILIWPCLPNKSRDFR 355
+ +L H G + L+ K R + I N D++ ++ + LE+L+ P+K +FR
Sbjct: 46 MARLSRHLGSLPLLAKHRINCIRTAIKRNMDVQNYAYSKQMLELLLSKAPPSKQDEFR 219
>TC233752 UP|Q8BTF6 (Q8BTF6) Mus musculus 16 days embryo head cDNA, RIKEN
full-length enriched library, clone:C130030K03
product:inferred: PROTEASOME SUBUNIT ALPHA TYPE 6
(PROTEASOME IOTA CHAIN) (MACROPAIN IOTA CHAIN) (MULTI,
full insert sequence. (Fra , partial (10%)
Length = 665
Score = 28.9 bits (63), Expect = 6.8
Identities = 17/42 (40%), Positives = 23/42 (54%)
Frame = +2
Query: 145 FMVSPLPLMPQLYLIYFMLMIVSYFVGPNPRRLVLL*TFFRF 186
F +P P Y YF+L S+F P+ R+VL+*TF F
Sbjct: 215 FRTNPSPSKDVRY--YFVLPAFSHF*NPHLTRMVLM*TFILF 334
>TC215926
Length = 1915
Score = 28.5 bits (62), Expect = 8.9
Identities = 16/56 (28%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Frame = +1
Query: 301 KLFVHFGGVALIVKRRFTGPKEKIFLNPDIR-VEWASKFLEILIWPCLPNKSRDFR 355
+L H G + L+ K R + I N D++ ++ + LE+L+ P+K +FR
Sbjct: 1207 RLSRHLGSLPLLAKHRINCIRTAIKRNMDVQNYAYSKQMLELLLSKAPPSKQDEFR 1374
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.344 0.154 0.513
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,871,436
Number of Sequences: 63676
Number of extensions: 439611
Number of successful extensions: 4002
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 2976
Number of HSP's successfully gapped in prelim test: 120
Number of HSP's that attempted gapping in prelim test: 991
Number of HSP's gapped (non-prelim): 3154
length of query: 600
length of database: 12,639,632
effective HSP length: 103
effective length of query: 497
effective length of database: 6,081,004
effective search space: 3022258988
effective search space used: 3022258988
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC119408.1


