
BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33190.1 473 aa
(473 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
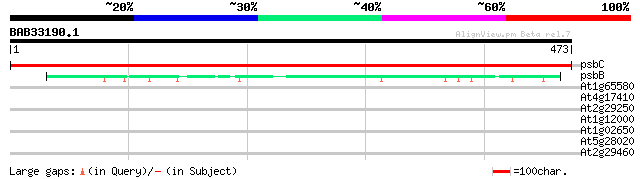
psbC -chloroplast genome- PSII 43 KDa protein 971 0.0
psbB -chloroplast genome- PSII 47KDa protein 92 5e-19
At1g65580 hypothetical protein 30 2.2
At4g17410 hypothetical protein 29 4.9
At2g29250 putative protein kinase 29 4.9
At1g12000 putative pyrophosphate-fructose-6-phosphate 1-phosphot... 29 4.9
At1g02650 hypothetical protein 29 6.4
At5g28020 cysteine synthase 28 8.4
At2g29460 putative glutathione S-transferase 28 8.4
>psbC -chloroplast genome- PSII 43 KDa protein
Length = 473
Score = 971 bits (2510), Expect = 0.0
Identities = 465/473 (98%), Positives = 469/473 (98%)
Query: 1 MKTLYSLRRFYHVETLFNGTLALTGRDQETTGFAWWAGNARLINLSGKLLGAHVAHAGLI 60
MKTLYSLRRFYHVETLFNGTLAL GRDQETTGFAWWAGNARLINLSGKLLGAHVAHAGLI
Sbjct: 1 MKTLYSLRRFYHVETLFNGTLALAGRDQETTGFAWWAGNARLINLSGKLLGAHVAHAGLI 60
Query: 61 VFWAGAMNLFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLI 120
VFWAGAMNLFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLI
Sbjct: 61 VFWAGAMNLFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLI 120
Query: 121 SSAVLGFGGIYHALLGPETLEESFPFFGYVWKDRNKMTTILGIHLILLGIGAFLLVFKAS 180
SSAVLGFGGIYHALLGPETLEESFPFFGYVWKDRNKMTTILGIHLILLG+GAFLLVFKA
Sbjct: 121 SSAVLGFGGIYHALLGPETLEESFPFFGYVWKDRNKMTTILGIHLILLGVGAFLLVFKAL 180
Query: 181 YFGGIYDTWAPGGGDVRKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWL 240
YFGG+YDTWAPGGGDVRKITNLTLSPS+IFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWL
Sbjct: 181 YFGGVYDTWAPGGGDVRKITNLTLSPSVIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWL 240
Query: 241 GSICILGGIWHILTKPFAWARRALVWSGEAYLSYSLAALSVFGFIACCFVWFNNTAYPSE 300
GSICI GGIWHILTKPFAWARRALVWSGEAYLSYSLAALSV GFIACCFVWFNNTAYPSE
Sbjct: 241 GSICIFGGIWHILTKPFAWARRALVWSGEAYLSYSLAALSVCGFIACCFVWFNNTAYPSE 300
Query: 301 FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD 360
FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD
Sbjct: 301 FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD 360
Query: 361 LRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINAVNYV 420
LRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINAVNYV
Sbjct: 361 LRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINAVNYV 420
Query: 421 SPRSWLATSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDFEPVLSMTPLN 473
SPRSWL+TSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDFEPVLSMTPLN
Sbjct: 421 SPRSWLSTSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDFEPVLSMTPLN 473
>psbB -chloroplast genome- PSII 47KDa protein
Length = 508
Score = 92.4 bits (228), Expect = 5e-19
Identities = 116/515 (22%), Positives = 187/515 (35%), Gaps = 107/515 (20%)
Query: 32 GFAWWAGNARLINLSGKLLGAHVAHAGLIVFWAGAMNLFEVAHFVPE----KPMYEQGLI 87
G W+ + ++N G+LL H+ H L+ WAG+M L+E+A F P PM+ QG+
Sbjct: 2 GLPWYRVHTVVLNDPGRLLAVHIMHTALVAGWAGSMALYELAVFDPSDPVLDPMWRQGMF 61
Query: 88 LLPHLATL-------GWGVGPGGEVIDTFPYFVSGV--LHLISSAVLGFGGIYHALLGPE 138
++P + L GW + GG + + + GV H++ S + I+H +
Sbjct: 62 VIPFMTRLGITNSWGGWNI-TGGTITNPGLWSYEGVAGAHIVFSGLCFLAAIWHWVYWDL 120
Query: 139 TL--EESFPFFGYVWKDRNKMTTILGIHLILLGIGAFLLVFKASYFGGIYDTWAPG--GG 194
+ +E K + I GIHL L G+ F F A + G+Y PG
Sbjct: 121 EIFCDER------TGKPSLDLPKIFGIHLFLSGVACF--GFGAFHVTGLY---GPGIWVS 169
Query: 195 DVRKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWLGSICILGGIWHILT 254
D +T + +G PF G I H+ G++ IL G++H+
Sbjct: 170 DPYGLTGKVQPVNPAWGVEGFDPFVPGG----------IASHHIAAGTLGILAGLFHLSV 219
Query: 255 KPFAWARRAL-VWSGEAYLSYSLAALSVFGFIACCFVWFNNTAYPSEFYGPTGPEASQA- 312
+P + L + + E LS S+AA+ F+ +W+ + P E +GPT + Q
Sbjct: 220 RPPQRLYKGLRMGNIETVLSSSIAAVFFAAFVVAGTMWYGSATTPIELFGPTRYQWDQGY 279
Query: 313 -------QAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD-LRAP 364
+ L +Q L Y+ +P +F +M D +
Sbjct: 280 FQQEIYRRVSAGLAENQSLSEAWAKIPEKLAFYDYIGNNPAKGGLFRAGSMDNGDGIAVG 339
Query: 365 WL--EPLRGPNGLDL--SRLKKDIQPWQ----------------ERRSAEYMTHAPLGSL 404
WL R G +L R+ + + R ++Y ++
Sbjct: 340 WLGHPVFRNKEGRELFVRRMPTFFETFPVVLVDGDGIVRADVPFRRAESKYSVEQVGVTV 399
Query: 405 NSVGGVATEINAVNYVSP---------------------------------RSWLATSHF 431
GG E+N V+Y P R W H
Sbjct: 400 EFYGG---ELNGVSYSDPATVKKYARRAQLGEIFELDRATLKSDGVFRSSPRGWFTFGHA 456
Query: 432 VLGFFLFVGHLWHAGRA--RAAAAGFEKGIDRDFE 464
F GH+WH R R AG + +D E
Sbjct: 457 SFALLFFFGHIWHGARTLFRDVFAGIDPDLDAQVE 491
>At1g65580 hypothetical protein
Length = 993
Score = 30.4 bits (67), Expect = 2.2
Identities = 17/38 (44%), Positives = 25/38 (65%), Gaps = 1/38 (2%)
Query: 38 GNARLINLSGKLLGAHVAHAGLIVFWA-GAMNLFEVAH 74
G ++ +L GKLLG VAH+G ++ A GA LF +A+
Sbjct: 460 GTVQVWDLEGKLLGGWVAHSGPVIKMAIGAGYLFTLAN 497
>At4g17410 hypothetical protein
Length = 744
Score = 29.3 bits (64), Expect = 4.9
Identities = 18/54 (33%), Positives = 24/54 (44%), Gaps = 2/54 (3%)
Query: 98 GVGPGGEVIDTFPYFVSGVLHLISSAVLGFGGIYHALLG-PETLEESF-PFFGY 149
G GPG + + F +GV H + GF G +H G P + PF GY
Sbjct: 411 GPGPGNQYFNGFQPGFNGVQHGFNGVQPGFNGFHHGFNGFPGPFPGAMPPFVGY 464
>At2g29250 putative protein kinase
Length = 623
Score = 29.3 bits (64), Expect = 4.9
Identities = 18/67 (26%), Positives = 31/67 (45%), Gaps = 13/67 (19%)
Query: 382 KDIQPWQERRSAEYMTHAPLGSLN-------------SVGGVATEINAVNYVSPRSWLAT 428
K I+ W E + E M + L L+ ++ G+ +E N V + + + +
Sbjct: 193 KPIRVWIEYNATETMLNVTLAPLDRPKPKLPLLSRKLNLSGIISEENYVGFSAATGTVTS 252
Query: 429 SHFVLGF 435
SHFVLG+
Sbjct: 253 SHFVLGW 259
>At1g12000 putative pyrophosphate-fructose-6-phosphate
1-phosphotransferase
Length = 566
Score = 29.3 bits (64), Expect = 4.9
Identities = 26/109 (23%), Positives = 44/109 (39%), Gaps = 15/109 (13%)
Query: 323 RLGANVGSAQGP------TGLGKYLMRSPTGEVIFGGE-------TMRFWDLRAPWLEPL 369
++G + Q P +GL YL G +G + ++ +L A +++P
Sbjct: 97 KIGVVLSGGQAPGGHNVISGLFDYLQERAKGSTFYGFKGGPAGIMKCKYVELNAEYIQPY 156
Query: 370 RGPNGLDL--SRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINA 416
R G D+ S K P Q +++ E L L +GG + NA
Sbjct: 157 RNQGGFDMICSGRDKIETPDQFKQAEETAKKLDLDGLVVIGGDDSNTNA 205
>At1g02650 hypothetical protein
Length = 513
Score = 28.9 bits (63), Expect = 6.4
Identities = 17/44 (38%), Positives = 21/44 (47%), Gaps = 4/44 (9%)
Query: 6 SLRRFYHVETLFNGTLALTGRDQETTGFAWWAGNARLINLSGKL 49
SL H++ LFN L RD+ TG W N R + GKL
Sbjct: 307 SLHDLEHLKLLFNSIL----RDRSLTGPVWKRHNVRYREIPGKL 346
>At5g28020 cysteine synthase
Length = 323
Score = 28.5 bits (62), Expect = 8.4
Identities = 19/50 (38%), Positives = 24/50 (48%), Gaps = 4/50 (8%)
Query: 292 FNNTAYPSEFYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYL 341
F N A P Y TGPE + A + L A VG+ TG+GK+L
Sbjct: 150 FENPANPEIHYRTTGPEIWRDSAGKVDI----LVAGVGTGGTATGVGKFL 195
>At2g29460 putative glutathione S-transferase
Length = 224
Score = 28.5 bits (62), Expect = 8.4
Identities = 27/108 (25%), Positives = 46/108 (42%), Gaps = 13/108 (12%)
Query: 290 VWFNNTAYPSE--------FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYL 341
+W NN P + F+ E AF + + ++ G V + L +L
Sbjct: 80 IWKNNPILPQDPYEKAMALFWAKFVDEQVGPVAFMSVAKAEK-GVEVAIKEAQE-LFMFL 137
Query: 342 MRSPTGEVIFGGETMRFWDLRAPWLEPL---RGPNGLDLSRLKKDIQP 386
+ TG+ FGG+T+ F DL A + P RG G+ + + ++ P
Sbjct: 138 EKEVTGKDFFGGKTIGFLDLVAGSMIPFCLARGWEGMGIDMIPEEKFP 185
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.143 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,425,050
Number of Sequences: 26719
Number of extensions: 526037
Number of successful extensions: 876
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 868
Number of HSP's gapped (non-prelim): 10
length of query: 473
length of database: 11,318,596
effective HSP length: 103
effective length of query: 370
effective length of database: 8,566,539
effective search space: 3169619430
effective search space used: 3169619430
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Description of BAB33190.1

