
BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33187.1 734 aa
(734 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
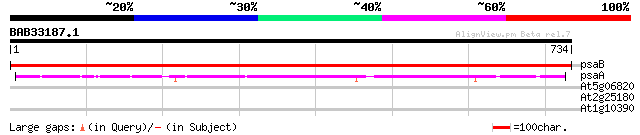
psaB -chloroplast genome- PSI P700 apoprotein A2 1522 0.0
psaA -chloroplast genome- PSI P700 apoprotein A1 628 e-180
At5g06820 SRF2 (LRR receptor protein kinase like) 34 0.26
At2g25180 putative two-component response regulator protein 30 3.7
At1g10390 unknown protein 29 8.2
>psaB -chloroplast genome- PSI P700 apoprotein A2
Length = 734
Score = 1522 bits (3940), Expect = 0.0
Identities = 718/734 (97%), Positives = 728/734 (98%)
Query: 1 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW 60
MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW
Sbjct: 1 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW 60
Query: 61 TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 120
TSGNLFHVAWQGNFE WVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV
Sbjct: 61 TSGNLFHVAWQGNFETWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 120
Query: 121 YQWWYTIGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS 180
YQWWYTIGLRTN DLY+GALFLLFLSA+SL+ GWLHLQPKWKP VSWFKNAESRLNHHLS
Sbjct: 121 YQWWYTIGLRTNEDLYTGALFLLFLSALSLIGGWLHLQPKWKPRVSWFKNAESRLNHHLS 180
Query: 181 GLFGVSSLAWTGHLVHVAIPGSRGEYVRWNNFLSILPHPQGLGPLFTGQWNLYAQNPDSS 240
GLFGVSSLAWTGHLVHVAIP SRGEYVRWNNFL++LPHPQGLGPLFTGQWNLYAQNPDSS
Sbjct: 181 GLFGVSSLAWTGHLVHVAIPASRGEYVRWNNFLNVLPHPQGLGPLFTGQWNLYAQNPDSS 240
Query: 241 SHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLLAGHMYRTNFGIGHS 300
SHLFGTSQG+GTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFL+AGHMYRTNFGIGHS
Sbjct: 241 SHLFGTSQGSGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLIAGHMYRTNFGIGHS 300
Query: 301 IKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAYAF 360
IKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAYAF
Sbjct: 301 IKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAYAF 360
Query: 361 IAQDYTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAIIS 420
IAQD+TTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAIIS
Sbjct: 361 IAQDFTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAIIS 420
Query: 421 HLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSS 480
HLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSS
Sbjct: 421 HLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSS 480
Query: 481 TNSPAFNAGRSIWLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALD 540
T+ PAFNAGRSIWLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALD
Sbjct: 481 TSGPAFNAGRSIWLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALD 540
Query: 541 ARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHIT 600
ARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHIT
Sbjct: 541 ARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHIT 600
Query: 601 LWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATG 660
LWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATG
Sbjct: 601 LWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATG 660
Query: 661 FMFLISWRGYWQELIETLAWAHERTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFT 720
FMFLISWRGYWQELIETLAWAHERTPLANLIRW+DKPVALSIVQARLVGLAHFSVGYIFT
Sbjct: 661 FMFLISWRGYWQELIETLAWAHERTPLANLIRWKDKPVALSIVQARLVGLAHFSVGYIFT 720
Query: 721 YAAFLIASTSGKFG 734
YAAFLIASTSGKFG
Sbjct: 721 YAAFLIASTSGKFG 734
>psaA -chloroplast genome- PSI P700 apoprotein A1
Length = 750
Score = 628 bits (1619), Expect = e-180
Identities = 342/746 (45%), Positives = 449/746 (59%), Gaps = 59/746 (7%)
Query: 8 FSQGLAQDP-TTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLWTSGNLF 66
FS+ +A+ P TT IW A AHDF+SH EE + + +F++HFGQL+IIFLW SG F
Sbjct: 32 FSRTIAKGPDTTTWIWNLHADAHDFDSHTSDLEE-ISRKVFSAHFGQLSIIFLWLSGMYF 90
Query: 67 HVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGVYQWWYT 126
H A N+EAW+ DP H+ P A +W P GQ + GG G + I SG +Q W
Sbjct: 91 HGARFSNYEAWLSDPTHIGPSAQVVW-PIVGQEILNGDVGGGFRG-IQIT-SGFFQIWRA 147
Query: 127 IGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLSGLFGVS 186
G+ + LY A+ L +A+ L AGW H K P ++WF++ ES LNHHL+GL G+
Sbjct: 148 SGITSELQLYCTAIGALVFAALMLFAGWFHYH-KAAPKLAWFQDVESMLNHHLAGLLGLG 206
Query: 187 SLAWTGHLVHVAIPGSRGEYVRWNNFLSI--------LPHPQGLGPLFTGQWNLYAQNPD 238
SL+W GH VHV++P N FL+ LPH L Q LY +
Sbjct: 207 SLSWAGHQVHVSLP--------INQFLNAGVDPKEIPLPHEFILNRDLLAQ--LYPSFAE 256
Query: 239 SSSHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLLAGHMYRTNFGIG 298
++ F + + LT GG P T LWLTD+AHHHLAIAILFL+AGHMYRTN+GIG
Sbjct: 257 GATPFFTLNWSKYSEFLTFRGGLDPVTGGLWLTDIAHHHLAIAILFLIAGHMYRTNWGIG 316
Query: 299 HSIKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAY 358
H IKD+LEAH G G+GHKGLY+ + S H QL L LA LG +T +VA HMYS+P Y
Sbjct: 317 HGIKDILEAH--KGPFTGQGHKGLYEILTTSWHAQLSLNLAMLGSLTIIVAHHMYSMPPY 374
Query: 359 AFIAQDYTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAI 418
++A DY TQ +L+THH +I GF++ GA AH AIF +RDY+P +++L R+L H++AI
Sbjct: 375 PYLATDYATQLSLFTHHMWIGGFLIVGAAAHAAIFMVRDYDPTNRYNDLLDRVLRHRDAI 434
Query: 419 ISHLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQ-----ILIEPIFAQWIQSAHGKTSYG 473
ISHL+W +FLGFH+ GLY+HND M A G P+ I ++P+FAQWIQ+ H
Sbjct: 435 ISHLNWVCIFLGFHSFGLYIHNDTMSALGRPQDMFSDTAIQLQPVFAQWIQNTH------ 488
Query: 474 FDVLLSSTNSPAFNAGRSI-WLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTL 532
L +P A S+ W G L A+ L + +G DFLVHH A +H T L
Sbjct: 489 --ALAPGVTAPGETASTSLTWGGGELVAVGGKVALLPIPLGTADFLVHHIHAFTIHVTVL 546
Query: 533 ILVKGALDARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTF 592
IL+KG L AR S+L+PDK + G+ FPCDGPGRGGTC +SAWD +L +FWM N I V F
Sbjct: 547 ILLKGVLFARSSRLIPDKANLGFRFPCDGPGRGGTCQVSAWDHVFLGLFWMYNAISVVIF 606
Query: 593 YWHWKHITLWQGNVS-----------QFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMN 641
++ WK + G++S F +SS + GWLRD+LW +SQ+I Y +
Sbjct: 607 HFSWKMQSDVWGSISDQGVVTHITGGNFAQSSITINGWLRDFLWAQASQVIQSYG----S 662
Query: 642 SLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWAHERTPLANLIRWRDKPVALS 701
SLS + FL H VWA MFL S RGYWQELIE++ WAH + +A +P ALS
Sbjct: 663 SLSAYGLFFLGAHFVWAFSLMFLFSGRGYWQELIESIVWAHNKLKVAP----ATQPRALS 718
Query: 702 IVQARLVGLAHFSVGYIFTYAAFLIA 727
I+Q R VG+ H+ +G I T AF +A
Sbjct: 719 IIQGRAVGVTHYLLGGIATTWAFFLA 744
>At5g06820 SRF2 (LRR receptor protein kinase like)
Length = 735
Score = 34.3 bits (77), Expect = 0.26
Identities = 18/49 (36%), Positives = 29/49 (58%), Gaps = 5/49 (10%)
Query: 39 EERLYQNIFASHFGQLAIIFLWTSGNLFHVAWQGNFEAWVQDPLHVRPI 87
E+ + I SHF + +++W GN FHV + N++ W + PL VRP+
Sbjct: 218 EDNQFSGIIPSHFQSIPHLWIW--GNKFHV--EPNYKPW-KFPLDVRPL 261
>At2g25180 putative two-component response regulator protein
Length = 573
Score = 30.4 bits (67), Expect = 3.7
Identities = 25/85 (29%), Positives = 33/85 (38%), Gaps = 5/85 (5%)
Query: 35 DDITEERLYQNIFASHFGQLAIIFLWTSGNLFHVAWQGNFEAWVQ-----DPLHVRPIAH 89
D + E+L + ASH + + SG A N E D H RPI
Sbjct: 227 DLMNVEKLTRENVASHLQKFRLYLKRISGVANQQAIMANSELHFMQMNGLDGFHHRPIPV 286
Query: 90 AIWDPHFGQPAVEAFTRGGALGPVN 114
H G PA+ +F G LG +N
Sbjct: 287 GSGQYHGGAPAMRSFPPNGILGRLN 311
>At1g10390 unknown protein
Length = 1041
Score = 29.3 bits (64), Expect = 8.2
Identities = 17/45 (37%), Positives = 21/45 (45%), Gaps = 2/45 (4%)
Query: 444 LAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSSTNSPAFNA 488
L F TP+ Q Q A G TS+G +TN+PAF A
Sbjct: 114 LGFSTPQSNPFGNS--TQQSQPAFGNTSFGSSTPFGATNTPAFGA 156
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.140 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,985,898
Number of Sequences: 26719
Number of extensions: 811885
Number of successful extensions: 1964
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 1945
Number of HSP's gapped (non-prelim): 8
length of query: 734
length of database: 11,318,596
effective HSP length: 107
effective length of query: 627
effective length of database: 8,459,663
effective search space: 5304208701
effective search space used: 5304208701
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 64 (29.3 bits)
Description of BAB33187.1

