
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33201.1 510 aa
(510 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
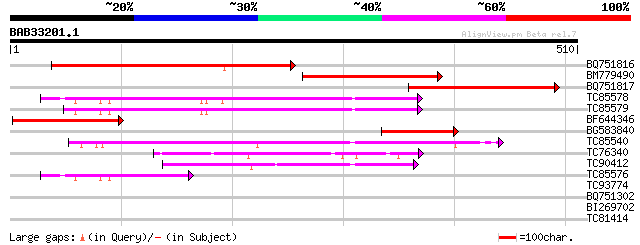
Score E
Sequences producing significant alignments: (bits) Value
BQ751816 similar to SP|P37211|ATPA ATP synthase alpha chain mit... 237 7e-63
BM779490 GP|12659355|gb ATPase alpha subunit {Calycanthus florid... 199 2e-51
BQ751817 similar to SP|P37211|ATPA ATP synthase alpha chain mit... 128 5e-30
TC85578 PIR|T05679|T05679 H+-transporting ATP synthase (EC 3.6.1... 122 3e-28
TC85579 homologue to GP|13366196|dbj|BAB39419. putative H+-trans... 120 1e-27
BF644346 PIR|A26760|A2 H+-transporting ATP synthase (EC 3.6.1.34... 102 3e-22
BG583840 GP|20146556|gb ATP synthase alpha subunit {Clethra barb... 101 8e-22
TC85540 homologue to PIR|T06538|T06538 probable H+-transporting ... 90 2e-18
TC76340 homologue to SP|Q9SM09|VATA_CITUN Vacuolar ATP synthase ... 80 2e-15
TC90412 homologue to SP|P23704|ATPB_NEUCR ATP synthase beta chai... 63 3e-10
TC85576 homologue to GP|6715512|gb|AAF26445.1| vacuolar H+-ATPas... 46 3e-05
TC93774 homologue to SP|Q9TKI7|ATPB_MEDSA ATP synthase beta chai... 40 0.002
BQ751302 similar to SP|P23704|ATPB ATP synthase beta chain mito... 36 0.031
BI269702 29 3.8
TC81414 29 4.9
>BQ751816 similar to SP|P37211|ATPA ATP synthase alpha chain mitochondrial
precursor (EC 3.6.3.14). {Neurospora crassa}, partial
(41%)
Length = 687
Score = 237 bits (605), Expect = 7e-63
Identities = 123/228 (53%), Positives = 160/228 (69%), Gaps = 8/228 (3%)
Frame = +3
Query: 38 GIARIYGLDEVMAGELVEFEEGTIGIALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQ 97
GIAR++GL V A ELVEF G G+ +NLE+ VGVVL G +++EG +VK TG I
Sbjct: 3 GIARVHGLANVQAEELVEFASGVKGMCMNLEAGQVGVVLFGSDRLVKEGEAVKRTGEIVD 182
Query: 98 IPVSEGYLGRVINALAKPIDGRGEISSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSM 157
+PV LGRVI+AL PIDG+G +++ + APGI+ R+SV +P+QTGL ++D+M
Sbjct: 183 VPVGTELLGRVIDALGNPIDGKGPLNTKTRSRAQVKAPGILPRKSVNQPVQTGLKSVDAM 362
Query: 158 IPIGRGQRELIIGDRQTGKTAVATDTILNQQGQN--------VICVYVAVGQKASSVAQV 209
+PIGRGQRELIIGDRQTGKTAVA D +LNQ+ N + CVYVAVGQK S+VAQ+
Sbjct: 363 VPIGRGQRELIIGDRQTGKTAVALDAMLNQKRWNATNDETKKLYCVYVAVGQKRSTVAQL 542
Query: 210 VNTLQERGAMEYTIVVAETADSPATLQYLAPYTGAALAEFFMYRERHT 257
V TL+E AM Y++VVA TA A YLAP+TGA++ E FM +H+
Sbjct: 543 VKTLEENDAMRYSVVVAATASEAAPPAYLAPFTGASVGEHFMQTGKHS 686
>BM779490 GP|12659355|gb ATPase alpha subunit {Calycanthus floridus}, partial
(29%)
Length = 381
Score = 199 bits (506), Expect = 2e-51
Identities = 101/126 (80%), Positives = 112/126 (88%)
Frame = +1
Query: 264 LSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSSQLGEGSMTALPIVETQ 323
L + +AYRQMSLLLRRPPGREA+PGDVFYLHSRLL+RAAK S Q G GS+TALP++ETQ
Sbjct: 4 LVNRFEAYRQMSLLLRRPPGREAFPGDVFYLHSRLLKRAAKRSDQTGAGSLTALPVIETQ 183
Query: 324 SGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAAQIKAMKQVAGKL 383
+GDVSAYIPTNVISITDGQI L +LF GIRPAINVG+SVSRVGSAAQ+KAMKQV G L
Sbjct: 184 AGDVSAYIPTNVISITDGQICLETELFYRGIRPAINVGLSVSRVGSAAQLKAMKQVCGSL 363
Query: 384 KLELAQ 389
KLELAQ
Sbjct: 364 KLELAQ 381
>BQ751817 similar to SP|P37211|ATPA ATP synthase alpha chain mitochondrial
precursor (EC 3.6.3.14). {Neurospora crassa}, partial
(26%)
Length = 669
Score = 128 bits (322), Expect = 5e-30
Identities = 68/136 (50%), Positives = 92/136 (67%)
Frame = -1
Query: 359 NVGISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLARGQRLR 418
NVG+ VS VGSAAQ+KAMKQVAG LKL LAQ+ E+ AFAQF SDLD AT+ L+RG+RL
Sbjct: 669 NVGLPVSGVGSAAQLKAMKQVAGSLKLFLAQYREVAAFAQFGSDLDAATKQTLSRGERLT 490
Query: 419 ELLKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNEIISS 478
ELLKQ Q +P+ V E V I+ G NG+LD++ + ++ ++ + +LKTN+ I
Sbjct: 489 ELLKQKQYSPMAVNEMVPLIFAGVNGHLDNVPVGKIGQWESDFLGHLKTNESDLLATIDK 310
Query: 479 TKTFTGEAEALLKEAI 494
+ + EA LK+ I
Sbjct: 309 EGALSKDLEARLKDVI 262
>TC85578 PIR|T05679|T05679 H+-transporting ATP synthase (EC 3.6.1.34) 54K
chain - Arabidopsis thaliana, partial (96%)
Length = 1994
Score = 122 bits (307), Expect = 3e-28
Identities = 109/381 (28%), Positives = 166/381 (42%), Gaps = 37/381 (9%)
Frame = +1
Query: 28 NTGTVLQVGDGIARIYGLDEVMAGELVEFE-------EGTIGIALNLESKNVGVVLMGDG 80
N L++G + G+ AG LV E + + I L + G VL DG
Sbjct: 97 NDDGTLEIGMEYRTVSGV----AGPLVILEKVKGPKYQEIVNIRLGDGTTRRGQVLEVDG 264
Query: 81 ----LMIQEGSS--------VKATGRIAQIPVSEGYLGRVINALAKPIDGRGEISSSESR 128
+ + EG+S V+ TG + + PVS LGR+ N KPID I
Sbjct: 265 EKAVVQVFEGTSGIDNKFTTVQFTGEVLKTPVSLDMLGRIFNGSGKPIDNGPPILPEAYL 444
Query: 129 LIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGD-------------RQTG 175
I + R E +QTG+ ID M I RGQ+ + RQ G
Sbjct: 445 DISGSSINPSERTYPEEMIQTGISTIDVMNSIARGQKIPLFSAAGLPHNEIAAQICRQAG 624
Query: 176 --KTAVATDTILNQQGQ--NVICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADS 231
K +D +L G+ N V+ A+G + +E G+ME + A+
Sbjct: 625 LVKRLEKSDNLLEGGGEDDNFAIVFAAMGVNMETAQFFKRDFEENGSMERVTLFLNLAND 804
Query: 232 PATLQYLAPYTGAALAEFFMYR-ERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGD 290
P + + P AE+ Y +H L+I D+S A A R++S PGR YPG
Sbjct: 805 PTIERIITPRIALTTAEYLAYECGKHVLVILTDMSSYADALREVSAAREEVPGRRGYPGY 984
Query: 291 VFYLHSRLLERAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLF 350
++ + + ERA ++ + +GS+T +PI+ + D++ P IT+GQI++ L
Sbjct: 985 MYTDLATIYERAGRIEGR--KGSITQIPILTMPNDDITHPTPDLTGYITEGQIYIDRQLH 1158
Query: 351 NAGIRPAINVGISVSRVGSAA 371
N I P INV S+SR+ +A
Sbjct: 1159NRQIYPPINVLPSLSRLMKSA 1221
>TC85579 homologue to GP|13366196|dbj|BAB39419. putative H+-transporting ATP
synthase {Oryza sativa (japonica cultivar-group)},
partial (97%)
Length = 1934
Score = 120 bits (302), Expect = 1e-27
Identities = 104/359 (28%), Positives = 158/359 (43%), Gaps = 36/359 (10%)
Frame = +1
Query: 49 MAGELVEFE-------EGTIGIALNLESKNVGVVLMGDG----LMIQEGSS--------V 89
+AG LV E + + I L + G VL DG + + EG+S V
Sbjct: 241 VAGPLVILEKVKGPKYQEIVNIRLGDGTTRRGQVLEVDGEKAVVQVFEGTSGIDNKFTTV 420
Query: 90 KATGRIAQIPVSEGYLGRVINALAKPIDGRGEISSSESRLIESPAPGIISRRSVYEPLQT 149
+ TG + + PVS LGR+ N KPID I I + R E +QT
Sbjct: 421 QFTGEVLKTPVSLDMLGRIFNGSGKPIDNGPPILPEAYLDISGSSINPSERTYPEEMIQT 600
Query: 150 GLIAIDSMIPIGRGQRELIIGD-------------RQTG--KTAVATDTILNQ-QGQNVI 193
G+ ID M I RGQ+ + RQ G K +D +L + N
Sbjct: 601 GISTIDVMNSIARGQKIPLFSAAGLPHNEIAAQICRQAGLVKRLEKSDNLLEGGEDDNFA 780
Query: 194 CVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAPYTGAALAEFFMYR 253
V+ A+G + +E G+ME + A+ P + + P AE+ Y
Sbjct: 781 IVFAAMGVNMETAQFFKRDFEENGSMERVTLFLNLANDPTIERIITPRIALTTAEYLAYE 960
Query: 254 -ERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSSQLGEG 312
+H L+I D+S A A R++S PGR YPG ++ + + ERA ++ + +G
Sbjct: 961 CGKHVLVILTDMSSYADALREVSAAREEVPGRRGYPGYMYTDLATIYERAGRIEGR--KG 1134
Query: 313 SMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAA 371
S+T +PI+ + D++ P IT+GQI++ L N I P INV S+SR+ +A
Sbjct: 1135SITQIPILTMPNDDITHPTPDLTGYITEGQIYIDRQLHNRQIYPPINVLPSLSRLMKSA 1311
>BF644346 PIR|A26760|A2 H+-transporting ATP synthase (EC 3.6.1.34) alpha
chain - garden pea mitochondrion, partial (20%)
Length = 402
Score = 102 bits (255), Expect = 3e-22
Identities = 48/100 (48%), Positives = 68/100 (68%)
Frame = +3
Query: 3 TIRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGTIG 62
++RA E++ ++ RI + T ++ G V+ VGDGIAR+YGL+E+ AGELVEF G G
Sbjct: 102 SVRAAELTTLLESRITNFYTNFQVDEIGRVVSVGDGIARVYGLNEIQAGELVEFASGVKG 281
Query: 63 IALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSE 102
IALNLE++NVG+V+ G I+EG VK TG I +P +
Sbjct: 282 IALNLENENVGIVVFGSDTSIKEGDLVKRTGSIVDVPAGK 401
>BG583840 GP|20146556|gb ATP synthase alpha subunit {Clethra barbinervis},
partial (17%)
Length = 585
Score = 101 bits (251), Expect = 8e-22
Identities = 52/69 (75%), Positives = 59/69 (85%)
Frame = +3
Query: 335 VISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAAQIKAMKQVAGKLKLELAQFAELE 394
VISITDGQI L +LF GIRPAINVG+SVSRVGSAAQ+KAMKQV G LKLELAQ+ E+
Sbjct: 237 VISITDGQICLETELFYRGIRPAINVGLSVSRVGSAAQLKAMKQVCGSLKLELAQYREVA 416
Query: 395 AFAQFASDL 403
AFAQF S++
Sbjct: 417 AFAQFGSNI 443
>TC85540 homologue to PIR|T06538|T06538 probable H+-transporting ATP
synthase (EC 3.6.1.34) beta chain mitochondrial -
garden pea, partial (96%)
Length = 2129
Score = 90.1 bits (222), Expect = 2e-18
Identities = 100/421 (23%), Positives = 174/421 (40%), Gaps = 30/421 (7%)
Frame = +2
Query: 54 VEFEEGTIGI--ALNLESKNVGVVL-----MGDGLM----------IQEGSSVKATGRIA 96
V FEEG I AL + + +VL +G+G++ + G V TG
Sbjct: 353 VRFEEGLPPILTALEVLDHSSRLVLEVAQHLGEGVVRTIAMDATEGVVRGWRVLNTGSPI 532
Query: 97 QIPVSEGYLGRVINALAKPIDGRGEISSSESRLIESPAPGIISRRSVYEPLQTGLIAIDS 156
+PV LGR++N + +PID +GE + I AP + + + + L TG+ +D
Sbjct: 533 SVPVGRATLGRIMNVIGEPIDHKGEFITEHYLPIHREAPAFVEQATEQQILVTGIKVVDL 712
Query: 157 MIPIGRGQRELIIGDRQTGKTAVATDTILN-QQGQNVICVYVAVGQKASSVAQVVNTLQE 215
+ P RG + + G GKT + + I N + V+ VG++ + + E
Sbjct: 713 LAPYQRGGKIGLFGGAGVGKTVLIMELINNVAKAHGGFSVFAGVGERTREGNDLYREMIE 892
Query: 216 RGAMEY--------TIVVAETADSPATLQYLAPYTGAALAEFFMYRE-RHTLIIYDDLSK 266
G ++ +V + P + TG +AE F E + L+ D++ +
Sbjct: 893 SGVIKLGEKQSESKCALVYGQMNEPPGARARVGLTGLTVAEHFRDAEGQDVLLFVDNIFR 1072
Query: 267 QAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSSQLGEGSMTALPIVETQSGD 326
QA ++S LL R P Y + L ER +GS+T++ + + D
Sbjct: 1073FTQANSEVSALLGRIPSAVGYQPTLSTDLGGLQERITTTK----KGSITSVQAIYVPADD 1240
Query: 327 VSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAAQI-KAMKQVAGKLKL 385
++ P + D LS + GI PA++ S SR+ S + Q A ++
Sbjct: 1241LTDPAPATTFAHLDATTVLSRQISELGIYPAVDPLDSTSRMLSPLILGDEHYQTARGVQQ 1420
Query: 386 ELAQFAELEAFAQF--ASDLDKATQNQLARGQRLRELLKQSQSAPLTVEEQVITIYTGTN 443
L + L+ +L + + +AR ++++ L Q P V E ++TG
Sbjct: 1421VLQNYKNLQDIIAILGMDELSEDDKLTVARARKIQRFLSQ----PFHVAE----VFTGAP 1576
Query: 444 G 444
G
Sbjct: 1577G 1579
>TC76340 homologue to SP|Q9SM09|VATA_CITUN Vacuolar ATP synthase catalytic
subunit A (EC 3.6.3.14) (V-ATPase A subunit), complete
Length = 2417
Score = 79.7 bits (195), Expect = 2e-15
Identities = 73/261 (27%), Positives = 121/261 (45%), Gaps = 18/261 (6%)
Frame = +1
Query: 130 IESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATDTILNQQG 189
+ +P P + S+ + PL TG +D++ P G I G GKT ++ L++
Sbjct: 742 VRTPRP-VASKLAADTPLLTGQRVLDALFPSVLGGTCAIPGAFGCGKTVISQ--ALSKYS 912
Query: 190 QNVICVYVAVGQKASSVAQVVNTL----------QERGAMEYTIVVAETADSPATLQYLA 239
+ +YV G++ + +A+V+ +E M+ T +VA T++ P + +
Sbjct: 913 NSDAVIYVGCGERGNEMAEVLMDFPQLTMTLPDGREESVMKRTTLVANTSNMPVAAREAS 1092
Query: 240 PYTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRL- 298
YTG LAE+F + ++ D S+ A+A R++S L P YP YL +RL
Sbjct: 1093IYTGITLAEYFRDMGYNVSMMADSTSRWAEALREISGRLAEMPADSGYPA---YLAARLA 1263
Query: 299 --LERAAKLSSQLG---EGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSAD--LFN 351
ERA K+ G GS+T + V GD S + + +SI Q+F D L
Sbjct: 1264SFYERAGKVKCLGGPERTGSVTIVGAVSPPGGDFSDPVTSATLSIV--QVFWGLDKKLAQ 1437
Query: 352 AGIRPAINVGISVSRVGSAAQ 372
P++N IS S+ +A +
Sbjct: 1438RKHFPSVNWLISYSKYSTALE 1500
>TC90412 homologue to SP|P23704|ATPB_NEUCR ATP synthase beta chain
mitochondrial precursor (EC 3.6.3.14). {Neurospora
crassa}, partial (67%)
Length = 1181
Score = 62.8 bits (151), Expect = 3e-10
Identities = 58/238 (24%), Positives = 101/238 (42%), Gaps = 8/238 (3%)
Frame = +1
Query: 138 ISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATDTILN-QQGQNVICVY 196
I + + E L TG+ +D + P RG + + G GKT + I N + V+
Sbjct: 1 IDQSTTAEILVTGIKVVDLLAPYARGGKIGLFGGAGVGKTVFIQELINNIAKAHGGYSVF 180
Query: 197 VAVGQKASSVAQVVNTLQER------GAMEYTIVVAETADSPATLQYLAPYTGAALAEFF 250
VG++ + + +QE G + +V + + P +A TG +AE+F
Sbjct: 181 TGVGERTREGNDLYHEMQETSVIQLDGESKVALVFGQMNEPPGARARVA-LTGLTIAEYF 357
Query: 251 MYRE-RHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSSQL 309
E + L+ D++ + QA ++S LL R P Y + + ER
Sbjct: 358 RDEEGQDVLLFIDNIFRFTQAGSEVSALLGRIPSAVGYQPTLAIDMGGMQERITTTK--- 528
Query: 310 GEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRV 367
+GS+T++ V + D++ P + D LS + GI PA++ S SR+
Sbjct: 529 -KGSITSVQAVYVPADDLTDPAPATTFAHLDATTVLSRGISELGIYPAVDPLDSKSRM 699
>TC85576 homologue to GP|6715512|gb|AAF26445.1| vacuolar H+-ATPase B subunit
{Nicotiana tabacum}, partial (40%)
Length = 668
Score = 46.2 bits (108), Expect = 3e-05
Identities = 47/157 (29%), Positives = 65/157 (40%), Gaps = 19/157 (12%)
Frame = +2
Query: 28 NTGTVLQVGDGIARIYGLDEVMAGELVEFE-------EGTIGIALNLESKNVGVVLMGDG 80
N L++G + G+ AG LV E + + I L + G VL DG
Sbjct: 98 NDDGTLEIGMEYRTVSGV----AGPLVILEKVKGPKYQEIVNIRLGDGTTRRGQVLEVDG 265
Query: 81 ----LMIQEGSS--------VKATGRIAQIPVSEGYLGRVINALAKPIDGRGEISSSESR 128
+ + EG+S V+ TG + + PVS LGR+ N KPID I
Sbjct: 266 EKAVVQVFEGTSGIDNKFTTVQFTGEVLKTPVSLDMLGRIFNGSGKPIDNGPPILPEAYL 445
Query: 129 LIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQR 165
I + R E +QTG+ ID M I RGQ+
Sbjct: 446 DISGSSINPSERTYPEEMIQTGISTIDVMNSIARGQK 556
>TC93774 homologue to SP|Q9TKI7|ATPB_MEDSA ATP synthase beta chain (EC
3.6.3.14). [Alfalfa] {Medicago sativa}, partial (29%)
Length = 497
Score = 40.4 bits (93), Expect = 0.002
Identities = 25/85 (29%), Positives = 37/85 (43%)
Frame = +1
Query: 67 LESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEISSSE 126
L + V V M + G V TG +PV LGR+ N L +PID G +++
Sbjct: 235 LGNNRVRAVAMSATDGLXXGMVVINTGAPLSVPVGGATLGRIFNVLGEPIDNLGPVNTGT 414
Query: 127 SRLIESPAPGIISRRSVYEPLQTGL 151
+ I AP I + +TG+
Sbjct: 415 TSPIHRSAPAFIQLDTKLSIFETGI 489
>BQ751302 similar to SP|P23704|ATPB ATP synthase beta chain mitochondrial
precursor (EC 3.6.3.14). {Neurospora crassa}, partial
(23%)
Length = 373
Score = 36.2 bits (82), Expect = 0.031
Identities = 22/65 (33%), Positives = 30/65 (45%)
Frame = -3
Query: 63 IALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEI 122
+A +L V + M + G TG IPV LGR+IN PID RG I
Sbjct: 218 VAQHLGENVVRCIAMDGTEGLVRGRKATDTGAPITIPVGPETLGRIINVTGDPIDERGPI 39
Query: 123 SSSES 127
+S++
Sbjct: 38 KTSKT 24
>BI269702
Length = 405
Score = 29.3 bits (64), Expect = 3.8
Identities = 18/53 (33%), Positives = 29/53 (53%)
Frame = +3
Query: 458 LVELRAYLKTNKPQFNEIISSTKTFTGEAEALLKEAIQEQMELFLLQEQVEKN 510
LV L ++L NK Q + I+SST T + ++ L +Q E + ++EKN
Sbjct: 39 LVLLLSFLVRNKNQEHSIVSSTSFCTIKDDSCLNARWTQQREWSNYESELEKN 197
>TC81414
Length = 826
Score = 28.9 bits (63), Expect = 4.9
Identities = 18/76 (23%), Positives = 38/76 (49%)
Frame = +1
Query: 4 IRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGTIGI 63
I+ DEI + + + + ++ ++ VGD I + ++EV+ ++ E T+ I
Sbjct: 508 IKDDEIMLAVDFQCLHNSGSQSVADSPIIVTVGDEIHGKFNINEVLPA--LKAEXNTVSI 681
Query: 64 ALNLESKNVGVVLMGD 79
A+++E +N V D
Sbjct: 682 AMDIERENXSVSXEAD 729
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.134 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,374,361
Number of Sequences: 36976
Number of extensions: 96410
Number of successful extensions: 469
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 460
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 461
length of query: 510
length of database: 9,014,727
effective HSP length: 100
effective length of query: 410
effective length of database: 5,317,127
effective search space: 2180022070
effective search space used: 2180022070
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Description of BAB33201.1

