
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33190.1 473 aa
(473 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
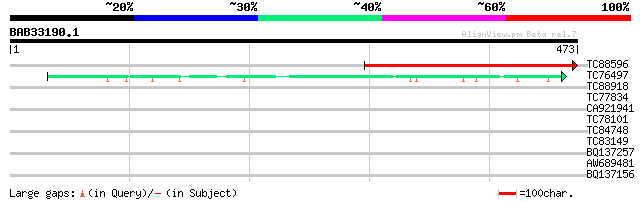
Sequences producing significant alignments: (bits) Value
TC88596 homologue to GP|4581045|gb|AAD24583.1| chlorophyll a-bin... 359 1e-99
TC76497 homologue to SP|P06411|PSBB_TOBAC Photosystem II P680 ch... 94 9e-20
TC88918 similar to GP|18447763|gb|AAL67992.1 putative serine car... 30 2.0
TC77834 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 29 3.4
CA921941 weakly similar to GP|22136974|gb| putative villin 2 pro... 29 3.4
TC78101 similar to GP|17381084|gb|AAL36354.1 unknown protein {Ar... 29 4.5
TC84748 similar to GP|9758300|dbj|BAB08843.1 contains similarity... 28 5.8
TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk p... 28 7.6
BQ137257 similar to GP|21536912|gb| unknown {Arabidopsis thalian... 28 7.6
AW689481 weakly similar to GP|11994595|db receptor-like serine/t... 28 7.6
BQ137156 28 10.0
>TC88596 homologue to GP|4581045|gb|AAD24583.1| chlorophyll a-binding
protein PsbC {Populus deltoides}, partial (37%)
Length = 1650
Score = 359 bits (922), Expect = 1e-99
Identities = 174/177 (98%), Positives = 174/177 (98%)
Frame = +3
Query: 297 YPSEFYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETM 356
YPSEFYGPTG EASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETM
Sbjct: 3 YPSEFYGPTGXEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETM 182
Query: 357 RFWDLRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINA 416
RFWDLRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINA
Sbjct: 183 RFWDLRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINA 362
Query: 417 VNYVSPRSWLATSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDFEPVLSMTPLN 473
VNYVSPRSWLATSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRD EPVL MTPLN
Sbjct: 363 VNYVSPRSWLATSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDSEPVLFMTPLN 533
>TC76497 homologue to SP|P06411|PSBB_TOBAC Photosystem II P680 chlorophyll A
apoprotein (CP-47 protein). [Common tobacco] {Nicotiana
tabacum}, complete
Length = 4045
Score = 94.4 bits (233), Expect = 9e-20
Identities = 121/519 (23%), Positives = 191/519 (36%), Gaps = 86/519 (16%)
Frame = +1
Query: 32 GFAWWAGNARLINLSGKLLGAHVAHAGLIVFWAGAMNLFEVAHFVPEKP----MYEQGLI 87
G W+ + ++N G+LL H+ H L+ WAG+M L+E+A F P P M+ QG+
Sbjct: 148 GLPWYRVHTVVLNDPGRLLSVHIMHTALVAGWAGSMALYELAVFDPSDPVLDPMWRQGMF 327
Query: 88 LLPHLATLG-------WGVGPGGEVIDTFPYFVSGVL--HLISSAVLGFGGIYHALLGPE 138
++P + LG W + GG + + + GV H++ S + I+H +
Sbjct: 328 VIPFMTRLGITNSWGGWNI-TGGTITNPGIWSYEGVAAAHIVFSGLCFLAAIWHWVYWDL 504
Query: 139 TL--EESFPFFGYVWKDRNKMTTILGIHLILLGIGAFLLVFKASYFGGIYDTWAPGGG-- 194
+ +E K + I GIHL L G+ F FG + T G G
Sbjct: 505 EIFCDER------TGKPSLDLPKIFGIHLFLAGVACF-------GFGAFHVTGLFGPGIW 645
Query: 195 --DVRKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWLGSICILGGIWHI 252
D +T S + +G PF G I H+ G++ IL G++H+
Sbjct: 646 VSDPYGLTGRVQSVNPAWGVDGFDPFVPGG----------IASHHIAAGTLGILAGLFHL 795
Query: 253 LTKPFAWARRALVWSG-EAYLSYSLAALSVFGFIACCFVWFNNTAYPSEFYGPTGPEASQ 311
+P + L E LS S+AA+ F+ +W+ + P E +GPT + Q
Sbjct: 796 SVRPPQRLYKGLRMGNIETVLSSSIAAVFFAAFVVAGTMWYGSATTPIELFGPTRYQWDQ 975
Query: 312 AQAFTFLVRDQRLGANVGSAQG--------PTGLG--KYLMRSPTGEVIFGGETMRFWD- 360
+ R R+GA + Q P L Y+ +P +F +M D
Sbjct: 976 GYFQQEIYR--RVGAGLAENQSLSEAWSKIPEKLAFYDYIGNNPAKGGLFRAGSMDNGDG 1149
Query: 361 LRAPWL-EPL-RGPNGLDL--SRLKKDIQPWQ----------------ERRSAEYMTHAP 400
+ WL P+ R G +L R+ + + R ++Y
Sbjct: 1150IAVGWLGHPIFRDKEGRELFVRRMPTFFETFPVVLVDGDGIVRADVPFRRAESKYSVEQV 1329
Query: 401 LGSLNSVGGVATEINAVNYVSP---------------------------------RSWLA 427
++ GG E+N V+Y P R W
Sbjct: 1330GVTVEFFGG---ELNGVSYSDPATVKKYARRAQLGEIFELDRATLKSDGVFRSSPRGWFT 1500
Query: 428 TSHFVLGFFLFVGHLWHAGRA--RAAAAGFEKGIDRDFE 464
H F GH+WH R R AG + +D E
Sbjct: 1501FGHASFALLFFFGHIWHGARTLFRDVFAGIDPDLDAQVE 1617
>TC88918 similar to GP|18447763|gb|AAL67992.1 putative serine
carboxypeptidase precursor {Gossypium hirsutum}, partial
(58%)
Length = 1145
Score = 30.0 bits (66), Expect = 2.0
Identities = 24/68 (35%), Positives = 32/68 (46%), Gaps = 1/68 (1%)
Frame = -3
Query: 105 VIDTFPYFVSGVLHLISSAVLGFGGIYHALLGPETLEESFPFF-GYVWKDRNKMTTILGI 163
++ FP F V HL S VL GI + + SF F G +WK + + L +
Sbjct: 357 ILLVFPGFKEKVEHLSSMRVLN--GIISSTMSQVINANSFVVFAGEIWKLKRLLNDDLVV 184
Query: 164 HLILLGIG 171
LILL IG
Sbjct: 183 VLILLHIG 160
>TC77834 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, complete
Length = 1604
Score = 29.3 bits (64), Expect = 3.4
Identities = 20/49 (40%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Frame = -1
Query: 142 ESFPFFGYVWKDRNKMTTILGIHL----ILLGIGAFLLVFKASYFGGIY 186
ESFP Y R K+ ILG L LL + AFL F AS+ G +
Sbjct: 953 ESFPVIQYHLAWRKKIKWILGFGLGCLFFLLLLSAFLAPFLASFLGSSF 807
>CA921941 weakly similar to GP|22136974|gb| putative villin 2 protein
{Arabidopsis thaliana}, partial (6%)
Length = 834
Score = 29.3 bits (64), Expect = 3.4
Identities = 16/36 (44%), Positives = 21/36 (57%), Gaps = 8/36 (22%)
Frame = -1
Query: 164 HLILLGIGAFL--LVFKASY------FGGIYDTWAP 191
H++L+GI +L +V K SY F GIY TW P
Sbjct: 384 HMLLMGIRIYLS*VVLKCSYLLNGVVFFGIYHTWIP 277
>TC78101 similar to GP|17381084|gb|AAL36354.1 unknown protein {Arabidopsis
thaliana}, partial (58%)
Length = 1267
Score = 28.9 bits (63), Expect = 4.5
Identities = 13/22 (59%), Positives = 14/22 (63%), Gaps = 1/22 (4%)
Frame = -2
Query: 221 EGWIVSVDDLED-IIGGHVWLG 241
EGW S D ED +GGH WLG
Sbjct: 132 EGWSRSSDYCED*FVGGHCWLG 67
>TC84748 similar to GP|9758300|dbj|BAB08843.1 contains similarity to unknown
protein~dbj|BAA89593.1~gene_id:MJH22.6 {Arabidopsis
thaliana}, partial (58%)
Length = 848
Score = 28.5 bits (62), Expect = 5.8
Identities = 11/33 (33%), Positives = 20/33 (60%)
Frame = +3
Query: 33 FAWWAGNARLINLSGKLLGAHVAHAGLIVFWAG 65
F++W NAR++ L+ K G ++ I+F +G
Sbjct: 543 FSFWFNNARIVPLASKEYGTRISTGSSILFSSG 641
>TC83149 similar to GP|13561982|gb|AAK30594.1 flagelliform silk protein
{Argiope trifasciata}, partial (3%)
Length = 1177
Score = 28.1 bits (61), Expect = 7.6
Identities = 25/88 (28%), Positives = 41/88 (46%)
Frame = -1
Query: 162 GIHLILLGIGAFLLVFKASYFGGIYDTWAPGGGDVRKITNLTLSPSIIFGYLLKSPFGGE 221
G+ L G+G LL+ + GG + W GGG S ++ G L E
Sbjct: 271 GVAAGLEGLGLLLLILGGDF*GG-FGCWGRGGGGGG-------SGVLLLGLL-------E 137
Query: 222 GWIVSVDDLEDIIGGHVWLGSICILGGI 249
G + V DLE+++G +G + ++GG+
Sbjct: 136 GLVDGV-DLEEVLGAD-GVGELGLVGGV 59
>BQ137257 similar to GP|21536912|gb| unknown {Arabidopsis thaliana}, partial
(15%)
Length = 1064
Score = 28.1 bits (61), Expect = 7.6
Identities = 14/31 (45%), Positives = 16/31 (51%), Gaps = 3/31 (9%)
Frame = -3
Query: 419 YVSPRSWLATSH---FVLGFFLFVGHLWHAG 446
+V P W +SH F L FFLF W AG
Sbjct: 177 FVPPGFWSFSSHSPSFFLFFFLFSSRFWSAG 85
>AW689481 weakly similar to GP|11994595|db receptor-like serine/threonine
kinase {Arabidopsis thaliana}, partial (4%)
Length = 528
Score = 28.1 bits (61), Expect = 7.6
Identities = 18/62 (29%), Positives = 30/62 (48%)
Frame = +3
Query: 284 FIACCFVWFNNTAYPSEFYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMR 343
F+ C V+ +N+ + S +Y P E+S + LVR ++ +A +GL Y
Sbjct: 39 FVQCHLVYISNSFW-SNYYSP**KESS*RYSLNHLVRKIGTSTSILAATNLSGLQMYQTM 215
Query: 344 SP 345
SP
Sbjct: 216 SP 221
>BQ137156
Length = 1010
Score = 27.7 bits (60), Expect = 10.0
Identities = 18/51 (35%), Positives = 24/51 (46%), Gaps = 1/51 (1%)
Frame = -3
Query: 400 PLGSLNSVGGVATEINAVNYVSPRSW-LATSHFVLGFFLFVGHLWHAGRAR 449
PL +N + A E + + +V R L+T F L FFL V WH R
Sbjct: 312 PLTGVNRLLFSAQEFHEIEWVMSRFIVLSTDSFPLSFFLRVSRGWHLSPMR 160
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.143 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,369,719
Number of Sequences: 36976
Number of extensions: 256410
Number of successful extensions: 1244
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1232
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1242
length of query: 473
length of database: 9,014,727
effective HSP length: 100
effective length of query: 373
effective length of database: 5,317,127
effective search space: 1983288371
effective search space used: 1983288371
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Description of BAB33190.1

