
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33187.1 734 aa
(734 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
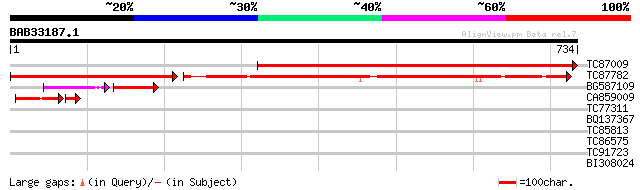
Score E
Sequences producing significant alignments: (bits) Value
TC87009 homologue to SP|P58385|PSAB_LOTJA Photosystem I P700 chl... 855 0.0
TC87782 homologue to SP|P05310|PSAA_PEA Photosystem I P700 chlor... 467 e-132
BG587109 homologue to GP|22091671|em photosystem1 subunit A {Pin... 70 9e-25
CA859009 homologue to PIR|S00703|S00 photosystem I protein A1 - ... 61 5e-11
TC77311 homologue to GP|6164957|gb|AAF04597.1| vacuolar H+-ATP s... 32 0.88
BQ137367 similar to SP|P08927|RUBB_ RuBisCO subunit binding-prot... 31 2.0
TC85813 homologue to GP|6503253|gb|AAC09468.2| putative NADPH-cy... 30 2.5
TC86575 similar to GP|10140758|gb|AAG13589.1 putative ubiquitin ... 29 5.7
TC91723 similar to PIR|T10585|T10585 serine proteinase homolog F... 29 7.4
BI308024 28 9.7
>TC87009 homologue to SP|P58385|PSAB_LOTJA Photosystem I P700 chlorophyll A
apoprotein A2 (PsaB) (PSI-B). {Lotus japonicus}, partial
(56%)
Length = 1445
Score = 855 bits (2210), Expect = 0.0
Identities = 407/413 (98%), Positives = 410/413 (98%)
Frame = +1
Query: 322 LYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAYAFIAQDYTTQAALYTHHQYIAGF 381
LYDTINNSIHFQLGLAL SLGVITSLVAQHMYSLPAYAFIAQD+TTQAALYTHHQYIAGF
Sbjct: 1 LYDTINNSIHFQLGLALXSLGVITSLVAQHMYSLPAYAFIAQDFTTQAALYTHHQYIAGF 180
Query: 382 IMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAIISHLSWASLFLGFHTLGLYVHND 441
IMTGAFAHGAIFFIRDYNPEQNEDNVLARML+HKEAIISHLSWASLFLGFHTLGLYVHND
Sbjct: 181 IMTGAFAHGAIFFIRDYNPEQNEDNVLARMLEHKEAIISHLSWASLFLGFHTLGLYVHND 360
Query: 442 VMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSSTNSPAFNAGRSIWLPGWLNAI 501
VMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSSTNSPA +AGRSIWLPGWLNAI
Sbjct: 361 VMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSSTNSPALSAGRSIWLPGWLNAI 540
Query: 502 NENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALDARGSKLMPDKKDFGYSFPCDG 561
NENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALDARGSKLMPDKKDFGYSFPCDG
Sbjct: 541 NENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALDARGSKLMPDKKDFGYSFPCDG 720
Query: 562 PGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHITLWQGNVSQFNESSTYLMGWLR 621
PGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHITLWQGNVSQFNESSTYLMGWLR
Sbjct: 721 PGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHITLWQGNVSQFNESSTYLMGWLR 900
Query: 622 DYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWA 681
DYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWA
Sbjct: 901 DYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWA 1080
Query: 682 HERTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFTYAAFLIASTSGKFG 734
HERTPLANLIRWRDKPVALSIVQARLVGL HFSVGYIFTYAAFLIASTSGKFG
Sbjct: 1081HERTPLANLIRWRDKPVALSIVQARLVGLVHFSVGYIFTYAAFLIASTSGKFG 1239
>TC87782 homologue to SP|P05310|PSAA_PEA Photosystem I P700 chlorophyll A
apoprotein A1 (PsaA) (PSI-A). [Garden pea] {Pisum
sativum}, partial (64%)
Length = 2152
Score = 467 bits (1202), Expect = e-132
Identities = 246/520 (47%), Positives = 320/520 (61%), Gaps = 17/520 (3%)
Frame = +2
Query: 225 LFTGQWNLYAQNPDSSSHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILF 284
+FT W+ YA LT G P T LWLTD+AHHHLAIAILF
Sbjct: 2 IFTLNWSKYAD------------------FLTFRXGLDPLTGGLWLTDIAHHHLAIAILF 127
Query: 285 LLAGHMYRTNFGIGHSIKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVI 344
L+AGHMYRTN+GIGH I+D+LEAH G G+GHKGLY+ + S H QL + LA LG +
Sbjct: 128 LIAGHMYRTNWGIGHGIRDILEAH--KGPFTGQGHKGLYEILTTSWHAQLSINLAMLGSL 301
Query: 345 TSLVAQHMYSLPAYAFIAQDYTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNE 404
T +VA HMY++P Y ++A DY TQ +L+THH +I GF++ GA AH AIF +RDY+P
Sbjct: 302 TIIVAHHMYAMPPYPYLATDYGTQLSLFTHHMWIGGFLIVGAAAHAAIFMVRDYDPTTRY 481
Query: 405 DNVLARMLDHKEAIISHLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQ-----ILIEPIF 459
+++L R+L H++AIISHL+W +FLGFH+ GLY+HND M A G P+ I ++P+F
Sbjct: 482 NDLLDRILRHRDAIISHLNWVCIFLGFHSFGLYIHNDTMSALGRPQDMFSDTAIQLQPVF 661
Query: 460 AQWIQSAHGKTSYGFDVLLSSTNSPAFNAGRSI-WLPGWLNAINENSNSLFLTIGPGDFL 518
AQWIQ+ H L T +P A S+ W G L A+ L + +G DFL
Sbjct: 662 AQWIQNTH--------ALAPGTTAPGATASTSLTWGGGDLVAVGGKVALLPIPLGTADFL 817
Query: 519 VHHAIALGLHTTTLILVKGALDARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYL 578
VHH A +H T LIL+KG L AR S+L+PDK + G+ FPCDGPGRGGTC +SAWD +L
Sbjct: 818 VHHIHAFTIHVTVLILLKGVLFARSSRLIPDKANLGFRFPCDGPGRGGTCQVSAWDHVFL 997
Query: 579 AVFWMLNTIGWVTFYWHWK-HITLW-----QGNVS-----QFNESSTYLMGWLRDYLWLN 627
+FWM N+I V F++ WK +W QG V+ F +SS + GWLRD+LW
Sbjct: 998 GLFWMYNSISVVIFHFSWKMQSDVWGSINDQGVVTHITGGNFAQSSITINGWLRDFLWAQ 1177
Query: 628 SSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWAHERTPL 687
+ +I Y +SLS + FL H VWA MFL S RGYWQELIE++ WAH + +
Sbjct: 1178AXXVIQSYG----SSLSAYGLFFLGAHFVWAFSLMFLFSGRGYWQELIESIVWAHNKLKV 1345
Query: 688 ANLIRWRDKPVALSIVQARLVGLAHFSVGYIFTYAAFLIA 727
A +P ALSIVQ R VG+ H+ +G I T AF +A
Sbjct: 1346AP----ATQPRALSIVQGRAVGVTHYLLGGIATTWAFFLA 1453
Score = 450 bits (1158), Expect = e-127
Identities = 211/217 (97%), Positives = 212/217 (97%)
Frame = +3
Query: 1 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW 60
MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW
Sbjct: 1500 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW 1679
Query: 61 TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 120
TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV
Sbjct: 1680 TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 1859
Query: 121 YQWWYTIGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS 180
YQWWYTIGLRTN DLY+GALFLLFLS ISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS
Sbjct: 1860 YQWWYTIGLRTNEDLYTGALFLLFLSVISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS 2039
Query: 181 GLFGVSSLAWTGHLVHVAIPGSRGEYVRWNNFLSILP 217
GLFGVSSLAW GHLVHVAIPGSR EYVRWNNFLS LP
Sbjct: 2040 GLFGVSSLAWAGHLVHVAIPGSRXEYVRWNNFLSGLP 2150
>BG587109 homologue to GP|22091671|em photosystem1 subunit A {Pinus
parviflora}, partial (29%)
Length = 791
Score = 70.1 bits (170), Expect(2) = 9e-25
Identities = 37/86 (43%), Positives = 50/86 (58%)
Frame = +2
Query: 44 QNIFASHFGQLAIIFLWTSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEA 103
+ +F++HFGQL+IIFLW SG FH A N+EA + DP H+ P A +W P GQ +
Sbjct: 170 EKVFSAHFGQLSIIFLWLSGMYFHGARFSNYEACLSDPTHIGPSAQVVW-PIVGQEILNG 346
Query: 104 FTRGGALGPVNIAYSGVYQWWYTIGL 129
GG G + I SG +Q W G+
Sbjct: 347 DVGGGFRG-IQIT-SGFFQIWRASGI 418
Score = 62.4 bits (150), Expect(2) = 9e-25
Identities = 28/58 (48%), Positives = 38/58 (65%)
Frame = +1
Query: 135 LYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLSGLFGVSSLAWTG 192
LY A+ L +A+ L AGW H K P ++WF++ ES LNHHL+GL G+ SL+W G
Sbjct: 430 LYCTAIGALVFAALMLFAGWFHYH-KAAPKLAWFQDVESMLNHHLAGLLGLGSLSWAG 600
>CA859009 homologue to PIR|S00703|S00 photosystem I protein A1 - garden pea
chloroplast, partial (16%)
Length = 823
Score = 61.2 bits (147), Expect(2) = 5e-11
Identities = 33/63 (52%), Positives = 43/63 (67%), Gaps = 1/63 (1%)
Frame = +2
Query: 8 FSQGLAQDP-TTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLWTSGNLF 66
FS+ +A+ P TT IW A AHDF+SH EE + + +F++HFGQL+IIFLW SG F
Sbjct: 572 FSRTIAKGPDTTTWIWNLHADAHDFDSHTSDLEE-ISRKVFSAHFGQLSIIFLWLSGMYF 748
Query: 67 HVA 69
H A
Sbjct: 749 HGA 757
Score = 24.6 bits (52), Expect(2) = 5e-11
Identities = 8/19 (42%), Positives = 12/19 (63%)
Frame = +1
Query: 73 NFEAWVQDPLHVRPIAHAI 91
N+EAW+ H+RP A +
Sbjct: 766 NYEAWLN*STHIRPSAQVV 822
>TC77311 homologue to GP|6164957|gb|AAF04597.1| vacuolar H+-ATP synthase
16kDa proteolipid subunit {Dendrobium crumenatum},
complete
Length = 932
Score = 32.0 bits (71), Expect = 0.88
Identities = 27/109 (24%), Positives = 40/109 (35%), Gaps = 2/109 (1%)
Frame = +3
Query: 589 WVTFYWHWKHITLWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYN-PFGMNSLSV-W 646
W + W W I + + + + GW YLW + ++ P G LS W
Sbjct: 177 WNSEEWCWSSINGCDET*TCYEINRSCCYGWCFGYLWFDYCCDY*YWD*P*G*ILLSF*W 356
Query: 647 AWMFLFGHLVWATGFMFLISWRGYWQELIETLAWAHERTPLANLIRWRD 695
F +W+ +L+ W GYW W + P A RW D
Sbjct: 357 ICSSFFWSCLWSC---WLVCWYGYW---CCW*CWC*SKCPTAETFRWND 485
>BQ137367 similar to SP|P08927|RUBB_ RuBisCO subunit binding-protein beta
subunit chloroplast precursor (60 kDa chaperonin beta
subunit), partial (8%)
Length = 723
Score = 30.8 bits (68), Expect = 2.0
Identities = 16/40 (40%), Positives = 18/40 (45%), Gaps = 1/40 (2%)
Frame = +2
Query: 667 WRGYWQELIETLAWAHERTPLANLIRWRD-KPVALSIVQA 705
WR W E I L W ++R N W D K SIV A
Sbjct: 437 WRDTWLEKIALLTWLYKRVATTNNSAWADVKRTTFSIVLA 556
>TC85813 homologue to GP|6503253|gb|AAC09468.2| putative NADPH-cytochrome
P450 reductase {Pisum sativum}, complete
Length = 2581
Score = 30.4 bits (67), Expect = 2.5
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +1
Query: 648 WMFLFGHLVWATGFMFLISWRGYW 671
W+ L L W+TG +F+I+W W
Sbjct: 877 WLHLILLLFWSTGLLFVINWMQLW 948
>TC86575 similar to GP|10140758|gb|AAG13589.1 putative ubiquitin protein
{Oryza sativa}, partial (58%)
Length = 2169
Score = 29.3 bits (64), Expect = 5.7
Identities = 10/29 (34%), Positives = 14/29 (47%)
Frame = +1
Query: 576 FYLAVFWMLNTIGWVTFYWHWKHITLWQG 604
F A FW + W WHW+ +W+G
Sbjct: 469 FRRARFWSF-IVSWTRCQWHWEKYLIWRG 552
>TC91723 similar to PIR|T10585|T10585 serine proteinase homolog F9F13.80 -
Arabidopsis thaliana, partial (41%)
Length = 1149
Score = 28.9 bits (63), Expect = 7.4
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = +3
Query: 108 GALGPVNIAYSGVYQWWYTIGLRTNGDLYSGALFL 142
G GP ++ S W +TIG ++ +YS +LFL
Sbjct: 63 GNTGPSPMSMSSFSPWIFTIGATSHDRVYSNSLFL 167
>BI308024
Length = 683
Score = 28.5 bits (62), Expect = 9.7
Identities = 11/17 (64%), Positives = 12/17 (69%)
Frame = -2
Query: 122 QWWYTIGLRTNGDLYSG 138
QWW T LR +GD YSG
Sbjct: 88 QWWRTTTLRNDGD*YSG 38
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.140 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,103,792
Number of Sequences: 36976
Number of extensions: 511896
Number of successful extensions: 3080
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 2208
Number of HSP's successfully gapped in prelim test: 128
Number of HSP's that attempted gapping in prelim test: 823
Number of HSP's gapped (non-prelim): 2397
length of query: 734
length of database: 9,014,727
effective HSP length: 103
effective length of query: 631
effective length of database: 5,206,199
effective search space: 3285111569
effective search space used: 3285111569
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Description of BAB33187.1

