
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33201.1 510 aa
(510 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
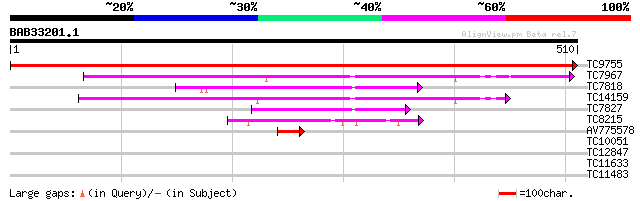
TC9755 UP|ATPA_LOTJA (Q9BBS3) ATP synthase alpha chain , complete 966 0.0
TC7967 UP|ATPB_LOTJA (Q9BBU0) ATP synthase beta chain , complete 100 7e-22
TC7818 UP|Q94AY7 (Q94AY7) AT4g38510/F20M13_70, partial (87%) 94 5e-20
TC14159 homologue to UP|ATP2_HEVBR (P29685) ATP synthase beta ch... 89 2e-18
TC7827 PIR|A31886|A31886 H+-exporting ATPase 57K chain - Arabid... 77 7e-15
TC8215 homologue to UP|VATA_CITUN (Q9SM09) Vacuolar ATP synthase... 62 3e-10
AV775578 49 3e-06
TC10051 homologue to UP|Q93W07 (Q93W07) Vacuolar ATPase B subuni... 39 0.002
TC12847 similar to GB|AAM66074.1|21595125|AY088542 endosperm-spe... 32 0.19
TC11633 28 3.6
TC11483 homologue to UP|Q9M3H9 (Q9M3H9) Tubby-like protein, part... 28 4.6
>TC9755 UP|ATPA_LOTJA (Q9BBS3) ATP synthase alpha chain , complete
Length = 2005
Score = 966 bits (2496), Expect = 0.0
Identities = 510/510 (100%), Positives = 510/510 (100%)
Frame = +1
Query: 1 MVTIRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGT 60
MVTIRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGT
Sbjct: 1 MVTIRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGT 180
Query: 61 IGIALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRG 120
IGIALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRG
Sbjct: 181 IGIALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRG 360
Query: 121 EISSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVA 180
EISSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVA
Sbjct: 361 EISSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVA 540
Query: 181 TDTILNQQGQNVICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAP 240
TDTILNQQGQNVICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAP
Sbjct: 541 TDTILNQQGQNVICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAP 720
Query: 241 YTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLE 300
YTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLE
Sbjct: 721 YTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLE 900
Query: 301 RAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINV 360
RAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINV
Sbjct: 901 RAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINV 1080
Query: 361 GISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLARGQRLREL 420
GISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLARGQRLREL
Sbjct: 1081GISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLARGQRLREL 1260
Query: 421 LKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNEIISSTK 480
LKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNEIISSTK
Sbjct: 1261LKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNEIISSTK 1440
Query: 481 TFTGEAEALLKEAIQEQMELFLLQEQVEKN 510
TFTGEAEALLKEAIQEQMELFLLQEQVEKN
Sbjct: 1441TFTGEAEALLKEAIQEQMELFLLQEQVEKN 1530
>TC7967 UP|ATPB_LOTJA (Q9BBU0) ATP synthase beta chain , complete
Length = 2457
Score = 100 bits (248), Expect = 7e-22
Identities = 109/455 (23%), Positives = 192/455 (41%), Gaps = 13/455 (2%)
Frame = +3
Query: 67 LESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEISSSE 126
L + V V M + G V TG +PV LGR+ N L +PID G + +
Sbjct: 744 LGNNRVRAVAMSATDGLTRGMEVIDTGAALSVPVGGVTLGRIFNVLGEPIDNLGPVDTRT 923
Query: 127 SRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATDTILN 186
+ I AP I + +TG+ +D + P RG + + G GKT + + I N
Sbjct: 924 TSPIHRSAPAFIQLDTKLSIFETGIKVVDLLAPYRRGGKIGLFGGAGVGKTVLIMELINN 1103
Query: 187 -QQGQNVICVYVAVGQKASSVAQVVNTLQERGAM-EYTIVVAETA------DSPATLQYL 238
+ + V+ VG++ + ++E G + E I ++ A + P +
Sbjct: 1104 IAKAHGGVSVFGGVGERTREGNDLYMEMKESGVINEKNIAESKVALVYGQMNEPPGARMR 1283
Query: 239 APYTGAALAEFFM-YRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSR 297
T +AE+F E+ L+ D++ + QA ++S LL R P Y +
Sbjct: 1284 VGLTALTMAEYFRDVNEQDVLLFIDNIFRFVQAGSEVSALLGRMPSAVGYQPTLSTEMGS 1463
Query: 298 LLERAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPA 357
L ER EGS+T++ V + D++ P + D LS L GI PA
Sbjct: 1464 LQERITSTK----EGSITSIQAVYVPADDLTDPAPATTFAHLDATTVLSRGLAAKGIYPA 1631
Query: 358 INVGISVSRVGSAAQI-KAMKQVAGKLKLELAQFAELEAFAQF--ASDLDKATQNQLARG 414
++ S S + + + + A ++K L ++ EL+ +L + + +AR
Sbjct: 1632 VDPLDSTSTMLQPRIVGEEHYETAQRVKQTLQRYKELQDIIAILGLDELSEEDRLTVARA 1811
Query: 415 QRLRELLKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNE 474
+++ L Q P V E ++TG+ G L +R F + L L + Q
Sbjct: 1812 RKIERFLSQ----PFFVAE----VFTGSPGKYVGL-AETIRGFKLILSGELDSLPEQAFY 1964
Query: 475 IISSTKTFTGEAEALLKEA-IQEQMELFLLQEQVE 508
++ + T +A L KE+ +++ + +F+ Q+E
Sbjct: 1965 LVGNIDEATVKATNLEKESKLKK*L*IFVY*PQIE 2069
>TC7818 UP|Q94AY7 (Q94AY7) AT4g38510/F20M13_70, partial (87%)
Length = 1481
Score = 94.0 bits (232), Expect = 5e-20
Identities = 69/239 (28%), Positives = 110/239 (45%), Gaps = 17/239 (7%)
Frame = +3
Query: 150 GLIAIDSMIPIGRGQRELIIGD-------------RQTG--KTAVATDTILNQ-QGQNVI 193
G+ ID M I RGQ+ + RQ G K +D +L + N
Sbjct: 3 GISTIDVMNSIARGQKIPLFSAAGLPHNEIAAQICRQAGLVKRLEKSDNLLEGGEDDNFA 182
Query: 194 CVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAPYTGAALAEFFMYR 253
V+ A+G + +E G+ME + A+ P + + P AE+ Y
Sbjct: 183 IVFAAMGVNMETAQFFKRDFEENGSMERVTLFLNLANDPTIERIITPRIALTTAEYLAYE 362
Query: 254 -ERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSSQLGEG 312
+H L+I D+S A A R++S PGR YPG ++ + + ERA ++ + +G
Sbjct: 363 CGKHVLVILTDMSSYADALREVSAAREEVPGRRGYPGYMYTDLATIYERAGRIEGR--KG 536
Query: 313 SMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAA 371
S+T +PI+ + D++ P IT+GQI++ L N I P INV S+SR+ +A
Sbjct: 537 SITQIPILTMPNDDITHPTPDLTGYITEGQIYIDRQLHNRQIYPPINVLPSLSRLMKSA 713
>TC14159 homologue to UP|ATP2_HEVBR (P29685) ATP synthase beta chain,
mitochondrial precursor , partial (93%)
Length = 2056
Score = 88.6 bits (218), Expect = 2e-18
Identities = 93/402 (23%), Positives = 167/402 (41%), Gaps = 14/402 (3%)
Frame = +3
Query: 63 IALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEI 122
+A +L V + M + G V TG +PV LGR++N + +PID +G+
Sbjct: 423 VAQHLGEGVVRTIAMDGTEGVVRGWRVLNTGSPITVPVGRVTLGRIMNVIGEPIDEKGDF 602
Query: 123 SSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATD 182
+ I AP + + + + L TG+ +D + P RG + + G GKT + +
Sbjct: 603 KTEHYLPIHREAPSFVEQATEQQILVTGIKVVDMLAPYQRGGKIGLFGGAGVGKTVLIME 782
Query: 183 TILN-QQGQNVICVYVAVGQKASSVAQVVNTLQERGAMEY--------TIVVAETADSPA 233
I N + V+ VG++ + + E G ++ +V + P
Sbjct: 783 LINNVAKAHGGFSVFAGVGERTREGNDLYREMIESGVIKLGDKQSESKCALVYGQMNEPP 962
Query: 234 TLQYLAPYTGAALAEFFMYRE-RHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVF 292
+ TG +AE F E + L+ D++ + QA ++S LL R P Y +
Sbjct: 963 GARARVGLTGLTVAEHFRDAEGQDVLLFVDNIFRFTQANSEVSALLGRIPSAVGYQPTLS 1142
Query: 293 YLHSRLLERAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNA 352
L ER +GS+T++ + + D++ P + D LS +
Sbjct: 1143TDLGGLQERITTTK----KGSITSVQAIYVPADDLTDPAPATTFAHLDATTVLSRQISEL 1310
Query: 353 GIRPAINVGISVSRVGSAAQI-KAMKQVAGKLKLELAQFAELEAFAQF--ASDLDKATQN 409
GI PA++ S SR+ S + + + A ++ L + L+ +L + +
Sbjct: 1311GIYPAVDPLDSTSRMLSPLILGEDHYETARGVQKVLQNYKNLQDIIAILGMDELSEDDKL 1490
Query: 410 QLARGQRLRELLKQSQSAPLTVEEQVITIYTGTNG-YLDSLE 450
+AR ++++ L Q P V E ++TG G Y+D E
Sbjct: 1491TVARARKIQRFLSQ----PFHVAE----VFTGAPGKYVDLKE 1592
>TC7827 PIR|A31886|A31886 H+-exporting ATPase 57K chain - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial
(29%)
Length = 519
Score = 77.0 bits (188), Expect = 7e-15
Identities = 45/144 (31%), Positives = 73/144 (50%), Gaps = 1/144 (0%)
Frame = +2
Query: 218 AMEYTIVVAETADSPATLQYLAPYTGAALAEFFMYR-ERHTLIIYDDLSKQAQAYRQMSL 276
+ME + A+ P + + P AE+ Y +H L+I D+S A A R++S
Sbjct: 92 SMERVTLFLNLANDPTIERIITPRIALTTAEYLAYECGKHVLVILTDMSSYADALREVSA 271
Query: 277 LLRRPPGREAYPGDVFYLHSRLLERAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVI 336
PGR YPG ++ + + ERA ++ + +GS+T +PI+ + D++ P
Sbjct: 272 AREEVPGRRGYPGYMYTDLATIYERAGRIEGR--KGSITQIPILTMPNDDITHPTPDLTG 445
Query: 337 SITDGQIFLSADLFNAGIRPAINV 360
IT+GQI++ L N I P INV
Sbjct: 446 YITEGQIYIDRQLHNRQIYPPINV 517
>TC8215 homologue to UP|VATA_CITUN (Q9SM09) Vacuolar ATP synthase catalytic
subunit A (V-ATPase A subunit) (Vacuolar proton pump
alpha subunit) (V-ATPase 69 kDa subunit) , partial (56%)
Length = 1563
Score = 61.6 bits (148), Expect = 3e-10
Identities = 56/194 (28%), Positives = 90/194 (45%), Gaps = 18/194 (9%)
Frame = +3
Query: 197 VAVGQKASSVAQVVNTL----------QERGAMEYTIVVAETADSPATLQYLAPYTGAAL 246
V G++ + +A+V+ +E M+ T +VA T++ P + + YTG L
Sbjct: 15 VGCGERGNEMAEVLMDFPQLTMTLPDGREESVMKRTTLVANTSNMPVAAREASIYTGITL 194
Query: 247 AEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRL---LERAA 303
AE+F + ++ D S+ A+A R++S L P YP YL +RL ERA
Sbjct: 195 AEYFRDMGYNVSMMADSTSRWAEALREISGRLAEMPADSGYPA---YLAARLASFYERAG 365
Query: 304 KLSSQLG---EGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSAD--LFNAGIRPAI 358
K+ G GS+T + V GD S + + +SI Q+F D L P++
Sbjct: 366 KVKCLGGPERTGSVTIVGAVSPPGGDFSDPVTSATLSIV--QVFWGLDKKLAQRKHFPSV 539
Query: 359 NVGISVSRVGSAAQ 372
N IS S+ +A +
Sbjct: 540 NWLISYSKYSTALE 581
>AV775578
Length = 485
Score = 48.5 bits (114), Expect = 3e-06
Identities = 22/24 (91%), Positives = 22/24 (91%)
Frame = -2
Query: 242 TGAALAEFFMYRERHTLIIYDDLS 265
TGA LAEFFMYRERH LIIYDDLS
Sbjct: 187 TGAGLAEFFMYRERHALIIYDDLS 116
>TC10051 homologue to UP|Q93W07 (Q93W07) Vacuolar ATPase B subunit, partial
(30%)
Length = 545
Score = 39.3 bits (90), Expect = 0.002
Identities = 43/150 (28%), Positives = 62/150 (40%), Gaps = 13/150 (8%)
Frame = +1
Query: 21 NTEIKIVNTGTVLQVGDGIAR-IYGLDEVMAGELVEFEEGTIGIALNLESKNVGVVLMGD 79
+TE + G + G+A + LD+V + E + I L + G VL D
Sbjct: 100 DTEDGTLEIGMEYRTVSGVAGPLVILDKVKGPKFQEI----VNIRLGDGTTRRGQVLEVD 267
Query: 80 G----LMIQEGSS--------VKATGRIAQIPVSEGYLGRVINALAKPIDGRGEISSSES 127
G + + EG+S V+ TG + + PVS LGR+ N KPID I
Sbjct: 268 GEKAVVQVFEGTSGIDNKFTTVQFTGEVLKTPVSLDMLGRIFNGSGKPIDNGPPILPEAY 447
Query: 128 RLIESPAPGIISRRSVYEPLQTGLIAIDSM 157
I + R E +QTG+ ID M
Sbjct: 448 LDISGSSINPSERTYPEEMIQTGISTIDVM 537
>TC12847 similar to GB|AAM66074.1|21595125|AY088542 endosperm-specific
protein-like protein {Arabidopsis thaliana;} , partial
(11%)
Length = 245
Score = 32.3 bits (72), Expect = 0.19
Identities = 20/53 (37%), Positives = 29/53 (53%)
Frame = -1
Query: 36 GDGIARIYGLDEVMAGELVEFEEGTIGIALNLESKNVGVVLMGDGLMIQEGSS 88
GDG+A + + GEL EEG +G + + +N+G V+ GDG EG S
Sbjct: 227 GDGLAAVD-----LVGELGLGEEGVVGAVVGVGGENLGDVVGGDGGGGGEGES 84
>TC11633
Length = 436
Score = 28.1 bits (61), Expect = 3.6
Identities = 13/24 (54%), Positives = 17/24 (70%)
Frame = +3
Query: 59 GTIGIALNLESKNVGVVLMGDGLM 82
GT+ A LES++VG L+GDG M
Sbjct: 30 GTVDSATGLESESVGESLLGDGEM 101
>TC11483 homologue to UP|Q9M3H9 (Q9M3H9) Tubby-like protein, partial (15%)
Length = 541
Score = 27.7 bits (60), Expect = 4.6
Identities = 13/37 (35%), Positives = 24/37 (64%), Gaps = 1/37 (2%)
Frame = +3
Query: 228 TADSPATLQYLAPYTGAALA-EFFMYRERHTLIIYDD 263
TA SP+TL++L PY+ A A + F+ R +++ + +
Sbjct: 99 TASSPSTLEFLDPYSKVAAAFDLFVLRFNFSVLCWKE 209
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.134 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,146,732
Number of Sequences: 28460
Number of extensions: 52036
Number of successful extensions: 251
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 248
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 249
length of query: 510
length of database: 4,897,600
effective HSP length: 94
effective length of query: 416
effective length of database: 2,222,360
effective search space: 924501760
effective search space used: 924501760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Description of BAB33201.1

