
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33190.1 473 aa
(473 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
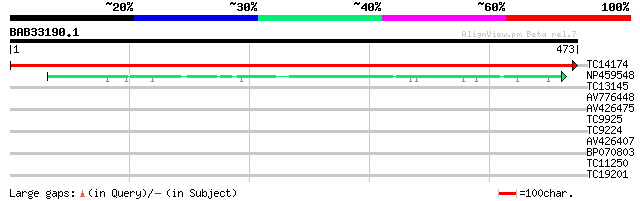
TC14174 UP|PSBC_LOTJA (Q9BBT1) Photosystem II 44 kDa reaction ce... 982 0.0
NP459548 PSII 47KDa protein [Lotus japonicus] 94 4e-20
TC13145 30 1.1
AV776448 29 1.5
AV426475 29 1.9
TC9925 29 1.9
TC9224 similar to GB|AAP21230.1|30102624|BT006422 At3g49750 {Ara... 28 3.2
AV426407 27 5.5
BP070803 27 7.2
TC11250 homologue to UP|AAQ72359 (AAQ72359) B-type cell cycle sw... 27 9.4
TC19201 similar to UP|Q8W1N7 (Q8W1N7) CSLC7 (Fragment), partial ... 27 9.4
>TC14174 UP|PSBC_LOTJA (Q9BBT1) Photosystem II 44 kDa reaction center protein
(P6 protein) (CP43), complete
Length = 3406
Score = 982 bits (2539), Expect = 0.0
Identities = 473/473 (100%), Positives = 473/473 (100%)
Frame = +1
Query: 1 MKTLYSLRRFYHVETLFNGTLALTGRDQETTGFAWWAGNARLINLSGKLLGAHVAHAGLI 60
MKTLYSLRRFYHVETLFNGTLALTGRDQETTGFAWWAGNARLINLSGKLLGAHVAHAGLI
Sbjct: 1513 MKTLYSLRRFYHVETLFNGTLALTGRDQETTGFAWWAGNARLINLSGKLLGAHVAHAGLI 1692
Query: 61 VFWAGAMNLFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLI 120
VFWAGAMNLFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLI
Sbjct: 1693 VFWAGAMNLFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLI 1872
Query: 121 SSAVLGFGGIYHALLGPETLEESFPFFGYVWKDRNKMTTILGIHLILLGIGAFLLVFKAS 180
SSAVLGFGGIYHALLGPETLEESFPFFGYVWKDRNKMTTILGIHLILLGIGAFLLVFKAS
Sbjct: 1873 SSAVLGFGGIYHALLGPETLEESFPFFGYVWKDRNKMTTILGIHLILLGIGAFLLVFKAS 2052
Query: 181 YFGGIYDTWAPGGGDVRKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWL 240
YFGGIYDTWAPGGGDVRKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWL
Sbjct: 2053 YFGGIYDTWAPGGGDVRKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWL 2232
Query: 241 GSICILGGIWHILTKPFAWARRALVWSGEAYLSYSLAALSVFGFIACCFVWFNNTAYPSE 300
GSICILGGIWHILTKPFAWARRALVWSGEAYLSYSLAALSVFGFIACCFVWFNNTAYPSE
Sbjct: 2233 GSICILGGIWHILTKPFAWARRALVWSGEAYLSYSLAALSVFGFIACCFVWFNNTAYPSE 2412
Query: 301 FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD 360
FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD
Sbjct: 2413 FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD 2592
Query: 361 LRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINAVNYV 420
LRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINAVNYV
Sbjct: 2593 LRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAPLGSLNSVGGVATEINAVNYV 2772
Query: 421 SPRSWLATSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDFEPVLSMTPLN 473
SPRSWLATSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDFEPVLSMTPLN
Sbjct: 2773 SPRSWLATSHFVLGFFLFVGHLWHAGRARAAAAGFEKGIDRDFEPVLSMTPLN 2931
>NP459548 PSII 47KDa protein [Lotus japonicus]
Length = 1527
Score = 94.4 bits (233), Expect = 4e-20
Identities = 119/515 (23%), Positives = 190/515 (36%), Gaps = 82/515 (15%)
Frame = +1
Query: 32 GFAWWAGNARLINLSGKLLGAHVAHAGLIVFWAGAMNLFEVAHFVPEKP----MYEQGLI 87
G W+ + ++N G+LL H+ H L+ WAG+M L+E+A F P P M+ QG+
Sbjct: 4 GLPWYRVHTVVLNDPGRLLAVHIMHTALVAGWAGSMALYELAVFDPSDPVLDPMWRQGMF 183
Query: 88 LLPHLATLG-------WGVGPGGEVIDTFPYFVSGVL--HLISSAVLGFGGIYHALLGPE 138
++P + LG W + GG + + + GV H++ S + I+H +
Sbjct: 184 VIPFMTRLGITNSWGGWNI-TGGTITNPGIWSYEGVAGAHIVFSGLCFLAAIWHWVYWDL 360
Query: 139 TLEESFPFFGYVWKDRNKMTTILGIHLILLGIGAFLLVFKASYFGGIYDTWAPG--GGDV 196
+ K + I GIHL L G+ F F A + G+Y PG D
Sbjct: 361 EIFSD----ERTGKPSLDLPKIFGIHLFLAGLACF--GFGAFHVTGLY---GPGIWVSDP 513
Query: 197 RKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWLGSICILGGIWHILTKP 256
+T + +G PF G + H+ G++ IL G++H+ +P
Sbjct: 514 YGLTGRVQPVNPAWGVEGFDPFVPGG----------VASHHIAAGTLGILAGLFHLSVRP 663
Query: 257 FAWARRALVWSG-EAYLSYSLAALSVFGFIACCFVWFNNTAYPSEFYGPTGPEASQAQAF 315
+ L E LS S+AA+ F+ +W+ + P E +GPT + Q
Sbjct: 664 PQRLYKGLRMGNIETVLSSSIAAVFFAAFVVAGTMWYGSATTPIELFGPTRYQWDQGYFQ 843
Query: 316 TFLVRDQRLGANVGSAQG--------PTGLG--KYLMRSPTGEVIFGGETMRFWD-LRAP 364
+ R R+GA + Q P L Y+ +P +F +M D +
Sbjct: 844 QEIYR--RVGAGLAENQSFSEAWSKIPEKLAFYDYIGNNPAKGGLFRAGSMDNGDGIAVG 1017
Query: 365 WL-EPL-RGPNGLDL--SRLKKDIQPWQ----------------ERRSAEYMTHAPLGSL 404
WL P+ R G +L R+ + + R ++Y ++
Sbjct: 1018WLGHPIFRDKEGRELFVRRMPTFFETFPVVLVDGDGIVRADVPFRRAESKYSVEQVGVTV 1197
Query: 405 NSVGGVATEINAVNYVSP---------------------------------RSWLATSHF 431
GG E+N V+Y P R W H
Sbjct: 1198EFYGG---ELNGVSYSDPATVKKYARRAQLGEIFELDRATLKSDGVFRSSPRGWFTFGHA 1368
Query: 432 VLGFFLFVGHLWHAGRA--RAAAAGFEKGIDRDFE 464
F GH+WH R R AG + +D E
Sbjct: 1369SFALLFFFGHIWHGSRTLFRDVFAGIDPDLDSQVE 1473
>TC13145
Length = 661
Score = 29.6 bits (65), Expect = 1.1
Identities = 16/55 (29%), Positives = 26/55 (47%)
Frame = +1
Query: 346 TGEVIFGGETMRFWDLRAPWLEPLRGPNGLDLSRLKKDIQPWQERRSAEYMTHAP 400
T + I+ G T R W L +L+P NG L+ +K + W ++ Y+ P
Sbjct: 412 TSKHIWAGPTPRKWGLPWNYLQPTTDINGTLLASHRKMDRRWWRAQAVRYLMRFP 576
>AV776448
Length = 569
Score = 29.3 bits (64), Expect = 1.5
Identities = 17/63 (26%), Positives = 28/63 (43%)
Frame = -1
Query: 4 LYSLRRFYHVETLFNGTLALTGRDQETTGFAWWAGNARLINLSGKLLGAHVAHAGLIVFW 63
L+ + F +T+++ L T G G N +G + G H+ + GL FW
Sbjct: 353 LHDMSSFPKTQTIYSSILLYTQLPAILVGSRS*YGKD---NATGYMEGRHIVYQGLCTFW 183
Query: 64 AGA 66
+GA
Sbjct: 182 SGA 174
>AV426475
Length = 300
Score = 28.9 bits (63), Expect = 1.9
Identities = 15/42 (35%), Positives = 22/42 (51%)
Frame = +2
Query: 343 RSPTGEVIFGGETMRFWDLRAPWLEPLRGPNGLDLSRLKKDI 384
R+PT +++FGG ++ RGPNGL LK D+
Sbjct: 23 RNPTAKLVFGGTHLQVKPSPVVAAFSSRGPNGLTPKILKPDL 148
>TC9925
Length = 546
Score = 28.9 bits (63), Expect = 1.9
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +2
Query: 261 RRALVWSGEAYLSYSLAALSVFGFIACCFVWF 292
+R L+W + Y +Y L L VF FI+ WF
Sbjct: 251 KRCLIW**DVYRNYFLLCLLVFHFISFI*FWF 346
>TC9224 similar to GB|AAP21230.1|30102624|BT006422 At3g49750 {Arabidopsis
thaliana;}, partial (77%)
Length = 1318
Score = 28.1 bits (61), Expect = 3.2
Identities = 19/56 (33%), Positives = 23/56 (40%), Gaps = 4/56 (7%)
Frame = -2
Query: 304 PTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIF----GGET 355
P PE Q A L+ D+R G P G + R TGE + GGET
Sbjct: 1002 PFTPERRQRVAVEILIPDERASVESGQVAAPVGERR---RDRTGEAVVGDVEGGET 844
>AV426407
Length = 415
Score = 27.3 bits (59), Expect = 5.5
Identities = 19/64 (29%), Positives = 28/64 (43%), Gaps = 8/64 (12%)
Frame = +1
Query: 375 LDLSRLKKDIQPW------QERRSAEYMTHAPLGSLNSVGGVATEINAVNYVSPRSW--L 426
L LS+++ QP + +RS H P L V +A + +Y+ PR W L
Sbjct: 118 LSLSQIRSRFQPAPALSPRRRQRSRREFWHPPWSRLQQVPALAPPLCRFSYLFPRLWCSL 297
Query: 427 ATSH 430
SH
Sbjct: 298 PRSH 309
>BP070803
Length = 407
Score = 26.9 bits (58), Expect = 7.2
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = -3
Query: 136 GPETLEESFPFFGYVWKDRNKMTTIL 161
G E + P FGY +RN++TT+L
Sbjct: 171 GAEWNKNFIPLFGYFMTERNQVTTLL 94
>TC11250 homologue to UP|AAQ72359 (AAQ72359) B-type cell cycle switch
protein ccs52B, partial (28%)
Length = 564
Score = 26.6 bits (57), Expect = 9.4
Identities = 12/25 (48%), Positives = 16/25 (64%), Gaps = 3/25 (12%)
Frame = +2
Query: 340 YLMRSPTGEVIF---GGETMRFWDL 361
YL SP G+ I G ET+RFW++
Sbjct: 290 YLAMSPDGQTIVTGAGDETLRFWNV 364
>TC19201 similar to UP|Q8W1N7 (Q8W1N7) CSLC7 (Fragment), partial (19%)
Length = 544
Score = 26.6 bits (57), Expect = 9.4
Identities = 10/30 (33%), Positives = 19/30 (63%)
Frame = -3
Query: 377 LSRLKKDIQPWQERRSAEYMTHAPLGSLNS 406
L R D+Q W++R ++ + H+ LG ++S
Sbjct: 380 LLRRGSDLQAWRDRSMSKKINHSALGPIHS 291
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.143 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,313,440
Number of Sequences: 28460
Number of extensions: 146209
Number of successful extensions: 673
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 662
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 669
length of query: 473
length of database: 4,897,600
effective HSP length: 94
effective length of query: 379
effective length of database: 2,222,360
effective search space: 842274440
effective search space used: 842274440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Description of BAB33190.1

