
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33187.1 734 aa
(734 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
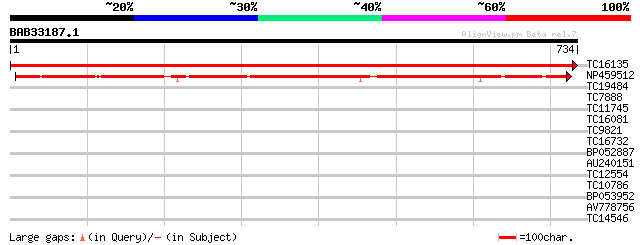
Score E
Sequences producing significant alignments: (bits) Value
TC16135 UP|PSAB_LOTJA (P58385) Photosystem I P700 chlorophyll A ... 1545 0.0
NP459512 PSI P700 apoprotein A1 [Lotus japonicus] 628 e-180
TC19484 similar to UP|BAC81651 (BAC81651) Polygalacturonase inhi... 30 1.1
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 30 1.4
TC11745 30 1.8
TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), par... 29 3.1
TC9821 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like prot... 29 3.1
TC16732 28 5.2
BP052887 28 5.2
AU240151 28 5.2
TC12554 similar to AAR99909 (AAR99909) Aminotransferase AGD2, pa... 28 6.8
TC10786 weakly similar to UP|Q9FMM3 (Q9FMM3) Gb|AAC80623.1, part... 27 8.9
BP053952 27 8.9
AV778756 27 8.9
TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxida... 27 8.9
>TC16135 UP|PSAB_LOTJA (P58385) Photosystem I P700 chlorophyll A apoprotein
A2 (PsaB) (PSI-B), complete
Length = 2647
Score = 1545 bits (3999), Expect = 0.0
Identities = 734/734 (100%), Positives = 734/734 (100%)
Frame = +1
Query: 1 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW 60
MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW
Sbjct: 1 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW 180
Query: 61 TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 120
TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV
Sbjct: 181 TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 360
Query: 121 YQWWYTIGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS 180
YQWWYTIGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS
Sbjct: 361 YQWWYTIGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS 540
Query: 181 GLFGVSSLAWTGHLVHVAIPGSRGEYVRWNNFLSILPHPQGLGPLFTGQWNLYAQNPDSS 240
GLFGVSSLAWTGHLVHVAIPGSRGEYVRWNNFLSILPHPQGLGPLFTGQWNLYAQNPDSS
Sbjct: 541 GLFGVSSLAWTGHLVHVAIPGSRGEYVRWNNFLSILPHPQGLGPLFTGQWNLYAQNPDSS 720
Query: 241 SHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLLAGHMYRTNFGIGHS 300
SHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLLAGHMYRTNFGIGHS
Sbjct: 721 SHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLLAGHMYRTNFGIGHS 900
Query: 301 IKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAYAF 360
IKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAYAF
Sbjct: 901 IKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAYAF 1080
Query: 361 IAQDYTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAIIS 420
IAQDYTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAIIS
Sbjct: 1081IAQDYTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAIIS 1260
Query: 421 HLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSS 480
HLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSS
Sbjct: 1261HLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQILIEPIFAQWIQSAHGKTSYGFDVLLSS 1440
Query: 481 TNSPAFNAGRSIWLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALD 540
TNSPAFNAGRSIWLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALD
Sbjct: 1441TNSPAFNAGRSIWLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALD 1620
Query: 541 ARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHIT 600
ARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHIT
Sbjct: 1621ARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHIT 1800
Query: 601 LWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATG 660
LWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATG
Sbjct: 1801LWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATG 1980
Query: 661 FMFLISWRGYWQELIETLAWAHERTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFT 720
FMFLISWRGYWQELIETLAWAHERTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFT
Sbjct: 1981FMFLISWRGYWQELIETLAWAHERTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFT 2160
Query: 721 YAAFLIASTSGKFG 734
YAAFLIASTSGKFG
Sbjct: 2161YAAFLIASTSGKFG 2202
>NP459512 PSI P700 apoprotein A1 [Lotus japonicus]
Length = 2253
Score = 628 bits (1619), Expect = e-180
Identities = 344/746 (46%), Positives = 449/746 (60%), Gaps = 26/746 (3%)
Frame = +1
Query: 8 FSQGLAQDP-TTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLWTSGNLF 66
FS+ +A+ P TT IW A AHDF+SH E+ + + IF++HFGQL+IIFLW SG F
Sbjct: 94 FSRTIAKGPDTTTWIWNLHADAHDFDSHTSDLED-ISRKIFSAHFGQLSIIFLWLSGMYF 270
Query: 67 HVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGVYQWWYT 126
H A N+EAW+ DP H+RP A +W P GQ + GG G + I SG +Q W
Sbjct: 271 HGARFSNYEAWLSDPTHIRPSAQVVW-PIVGQEILNGDVGGGFRG-IQIT-SGFFQIWRA 441
Query: 127 IGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLSGLFGVS 186
G+ + LY A+ L +A+ L AGW H K P ++WF++ ES LNHHL+GL G+
Sbjct: 442 SGITSELQLYCTAIGGLVFAALMLFAGWFHYH-KAAPKLAWFQDVESMLNHHLAGLLGLG 618
Query: 187 SLAWTGHLVHVAIPGSRGEYVRWNNFLSI--------LPHPQGLGPLFTGQWNLYAQNPD 238
SL+W GH VHV++P N FL+ LPH L Q LY +
Sbjct: 619 SLSWAGHQVHVSLP--------INQFLNAGVDPKEIPLPHEFILNRDLLAQ--LYPSFAE 768
Query: 239 SSSHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLLAGHMYRTNFGIG 298
++ F + LT GG P T LWLTD+AHHHLAIAILFL+AGHMYRTN+GIG
Sbjct: 769 GATPFFTLNWAKYADFLTFRGGLDPVTGGLWLTDIAHHHLAIAILFLIAGHMYRTNWGIG 948
Query: 299 HSIKDLLEAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQHMYSLPAY 358
H IKD+LEAH G G+GHKGLY+ + S H QL + LA LG +T +VA HMYS+P Y
Sbjct: 949 HGIKDILEAH--KGPFTGQGHKGLYEILTTSWHAQLSINLAMLGSLTIIVAHHMYSMPPY 1122
Query: 359 AFIAQDYTTQAALYTHHQYIAGFIMTGAFAHGAIFFIRDYNPEQNEDNVLARMLDHKEAI 418
++A DY TQ +L+THH +I GF++ GA AH AIF +RDY+P +++L R+L H++AI
Sbjct: 1123PYLATDYGTQLSLFTHHMWIGGFLIVGAAAHAAIFMVRDYDPTTRYNDLLDRVLRHRDAI 1302
Query: 419 ISHLSWASLFLGFHTLGLYVHNDVMLAFGTPEKQ-----ILIEPIFAQWIQSAHGKTSYG 473
ISHL+W +FLGFH+ GLY+HND M A G P+ I ++PIFAQWIQ+ H
Sbjct: 1303ISHLNWVCIFLGFHSFGLYIHNDTMSALGRPQDMFSDTAIQLQPIFAQWIQNTH------ 1464
Query: 474 FDVLLSSTNSPAFNAGRSI-WLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTL 532
L +P G S+ W G L A+ L + +G DFLVHH A +H T L
Sbjct: 1465--ALAPGITAPGATTGTSLTWGGGDLVAVGNKVALLPIPLGTADFLVHHIHAFTIHVTVL 1638
Query: 533 ILVKGALDARGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTF 592
IL+KG L AR S+L+PDK + G+ FPCDGPGRGGTC +SAWD +L +FWM N I V F
Sbjct: 1639ILLKGVLFARSSRLIPDKANLGFRFPCDGPGRGGTCQVSAWDHVFLGLFWMYNAISVVIF 1818
Query: 593 YWHWKHITLWQGNVS-----------QFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMN 641
++ WK + G++S F +SS + GWLRD+LW +SQ+I Y +
Sbjct: 1819HFSWKMQSDVWGSISDQGVVTHITGGNFAQSSITINGWLRDFLWAQASQVIQSYG----S 1986
Query: 642 SLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWAHERTPLANLIRWRDKPVALS 701
SLS + FL H VWA MFL S RGYWQELIE++ WAH + +A +P ALS
Sbjct: 1987SLSAYGLFFLGAHFVWAFSLMFLFSGRGYWQELIESIVWAHNKLKVAP----ATQPRALS 2154
Query: 702 IVQARLVGLAHFSVGYIFTYAAFLIA 727
IVQ R VG+ H+ +G I T AF +A
Sbjct: 2155IVQGRAVGVTHYLLGGIATTWAFFLA 2232
>TC19484 similar to UP|BAC81651 (BAC81651) Polygalacturonase inhibiting
protein (Fragment), partial (89%)
Length = 595
Score = 30.4 bits (67), Expect = 1.1
Identities = 29/108 (26%), Positives = 44/108 (39%), Gaps = 6/108 (5%)
Frame = +3
Query: 222 LGPLFTGQWNLYAQNPDSSSHLFGTSQGAGTAILTL------LGGFHPQTQSLWLTDMAH 275
LGP+ GQ L+ + +S LFG ++ L+ T+SL D+ H
Sbjct: 147 LGPVRPGQDRLFKEQEGDASFLFGPNKKIRILDLSKNLLSFDFSNVEIPTKSLIWLDLNH 326
Query: 276 HHLAIAILFLLAGHMYRTNFGIGHSIKDLLEAHIPPGGRLGRGHKGLY 323
+ + +I L F + + + L IP GG LGR K Y
Sbjct: 327 NMIHGSIPIGLTAVENLQQFNVSY---NQLSGKIPKGGELGRFDKYSY 461
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 30.0 bits (66), Expect = 1.4
Identities = 33/137 (24%), Positives = 48/137 (34%), Gaps = 24/137 (17%)
Frame = -3
Query: 582 WMLNTIGWVTFYWHWKHI----TLWQGN------VSQFNESSTYLMGWLRDY-----LWL 626
W L I W W ++HI W GN + F+ + MGW ++ W
Sbjct: 551 WWLRNIRWWFHNWWFRHIRRFHNWWFGNMGWFWHIRWFHNWWFWNMGWFHNWWFWHMRWF 372
Query: 627 NSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGY----WQELIETL---- 678
N +L G + L W W F + W + F+ W Y W+ + L
Sbjct: 371 NHWRL--GIVRWFW*WLGWWTW---FRAMRWRVLWWFMNWWLHYRRMLWRFMNRGLHYWR 207
Query: 679 -AWAHERTPLANLIRWR 694
W L N + WR
Sbjct: 206 VLWRFVNRRLHNRVLWR 156
>TC11745
Length = 440
Score = 29.6 bits (65), Expect = 1.8
Identities = 18/56 (32%), Positives = 22/56 (39%)
Frame = -3
Query: 602 WQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVW 657
+ G VS N G R + S +NG FG VW W F G L+W
Sbjct: 243 FSGGVSDGNVRFVLRSGLRRFHPAARESAYLNGVGSFGF----VWNWNFERGELIW 88
>TC16081 similar to UP|Q9XGQ5 (Q9XGQ5) ESTs AU064813(E40579), partial (37%)
Length = 1189
Score = 28.9 bits (63), Expect = 3.1
Identities = 18/56 (32%), Positives = 25/56 (44%)
Frame = +1
Query: 546 LMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHITL 601
L P +F SFP G G C +S W FY W+ + + F W + HI +
Sbjct: 334 LEPSSLNFSLSFPVSGLGGSIMCLVSCWWFFYC---WLSSVL---KFQW-FSHICI 480
>TC9821 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like protein,
partial (61%)
Length = 1080
Score = 28.9 bits (63), Expect = 3.1
Identities = 20/63 (31%), Positives = 28/63 (43%), Gaps = 7/63 (11%)
Frame = -2
Query: 580 VFW------MLNTIGWVTFYWHWKHITLWQGNVSQFNESSTYLMGWLRDYL-WLNSSQLI 632
VFW +L + W F W H T GN+S L+G + Y+ + NSS+
Sbjct: 884 VFWEGTCSKLLFCLPWQEFTWLRSHSTGNPGNISASPRHFFSLLGQFKGYMGYANSSKRG 705
Query: 633 NGY 635
GY
Sbjct: 704 TGY 696
>TC16732
Length = 676
Score = 28.1 bits (61), Expect = 5.2
Identities = 11/24 (45%), Positives = 14/24 (57%)
Frame = -1
Query: 411 MLDHKEAIISHLSWASLFLGFHTL 434
++D K + HLSW L L FH L
Sbjct: 97 LVDEKMVVQQHLSWVYLHLAFHML 26
>BP052887
Length = 420
Score = 28.1 bits (61), Expect = 5.2
Identities = 18/78 (23%), Positives = 31/78 (39%), Gaps = 9/78 (11%)
Frame = +3
Query: 592 FYWHWKHITLWQ--GNVSQFNESST-------YLMGWLRDYLWLNSSQLINGYNPFGMNS 642
F+ W H+ +W S F++S +L+ W+R W+ I+ +
Sbjct: 177 FFPFWAHLAIWPYLKPGSCFSDSGVSDGRRERFLVSWMRLNGWVPPLDGISIFFSLASTG 356
Query: 643 LSVWAWMFLFGHLVWATG 660
L W W + + W TG
Sbjct: 357 LG*WPWWYTW*WPRWYTG 410
>AU240151
Length = 286
Score = 28.1 bits (61), Expect = 5.2
Identities = 10/29 (34%), Positives = 13/29 (44%)
Frame = +1
Query: 643 LSVWAWMFLFGHLVWATGFMFLISWRGYW 671
L W W G W TGF++ W +W
Sbjct: 184 LGTWTWRSGLGSWPWRTGFLWTWWWFFWW 270
>TC12554 similar to AAR99909 (AAR99909) Aminotransferase AGD2, partial (13%)
Length = 491
Score = 27.7 bits (60), Expect = 6.8
Identities = 16/39 (41%), Positives = 22/39 (56%)
Frame = +3
Query: 108 GALGPVNIAYSGVYQWWYTIGLRTNGDLYSGALFLLFLS 146
GA+ +N A S + GL N D+YSG F++FLS
Sbjct: 258 GAMSSLNNAISFLGI*TMM*GLINNDDVYSGICFIMFLS 374
>TC10786 weakly similar to UP|Q9FMM3 (Q9FMM3) Gb|AAC80623.1, partial (9%)
Length = 756
Score = 27.3 bits (59), Expect = 8.9
Identities = 15/42 (35%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Frame = -1
Query: 420 SHLSWASLFLGFHTLGLY---VHNDVMLAFGTPEKQILIEPI 458
SH++ S FLGF +Y +H+DV+L E + I+P+
Sbjct: 612 SHVADQS*FLGFLNHSVYDYQMHHDVLLLSSLKEDYLAIDPL 487
>BP053952
Length = 462
Score = 27.3 bits (59), Expect = 8.9
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = +3
Query: 666 SWRGYWQELIETLAWAHERTPLANL 690
SW+ +W +L L W+ +T L NL
Sbjct: 18 SWKIWWNQLFHVLDWSPIKTHLINL 92
>AV778756
Length = 513
Score = 27.3 bits (59), Expect = 8.9
Identities = 16/51 (31%), Positives = 23/51 (44%), Gaps = 1/51 (1%)
Frame = +2
Query: 177 HHLSGLF-GVSSLAWTGHLVHVAIPGSRGEYVRWNNFLSILPHPQGLGPLF 226
H LS + G+ S +W+ G Y +W+ + PHP GL LF
Sbjct: 359 HQLSKME*GLGSRSWSSIGSSSRKVGWASTYTQWSTGPEVAPHPTGLVSLF 511
>TC14546 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxidase
precursor , partial (66%)
Length = 738
Score = 27.3 bits (59), Expect = 8.9
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = -3
Query: 46 IFASHFGQLAIIFLWTSGNLFHVAWQGNFEAWVQDPLHVR 85
+F FGQ+ + L +LFH+ WQ V + LH+R
Sbjct: 484 LFLHLFGQVVLAMLVLQADLFHLLWQ------VLEYLHMR 383
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.140 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,546,322
Number of Sequences: 28460
Number of extensions: 288341
Number of successful extensions: 1775
Number of sequences better than 10.0: 31
Number of HSP's better than 10.0 without gapping: 1744
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1764
length of query: 734
length of database: 4,897,600
effective HSP length: 97
effective length of query: 637
effective length of database: 2,136,980
effective search space: 1361256260
effective search space used: 1361256260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Description of BAB33187.1

