
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33201.1 510 aa
(510 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
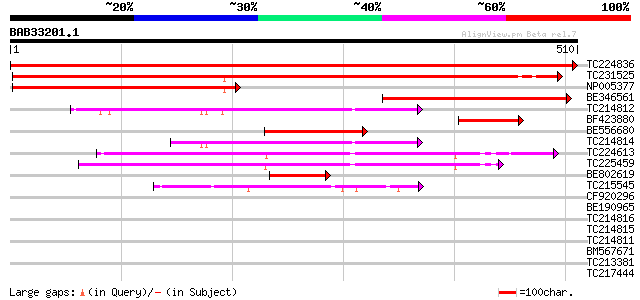
TC224836 homologue to UP|ATPA_LOTJA (Q9BBS3) ATP synthase alpha ... 951 0.0
TC231525 atpA 573 e-164
NP005377 pseudo-atpA 238 4e-63
BE346561 similar to SP|Q9BBS3|ATPA ATP synthase alpha chain (EC ... 234 5e-62
TC214812 homologue to UP|Q9ZQS1 (Q9ZQS1) Vacuolar H+-ATPase B su... 124 1e-28
BF423880 homologue to GP|20068316|emb ATPase alpha subunit {Atro... 112 5e-25
BE556680 weakly similar to GP|14715604|gb| ATP synthase alpha su... 102 5e-22
TC214814 homologue to UP|Q8GUB5 (Q8GUB5) Vacuolar ATPase subunit... 100 1e-21
TC224613 homologue to UP|Q95FJ6 (Q95FJ6) ATP synthase beta subun... 100 2e-21
TC225459 homologue to UP|O04275 (O04275) F1 ATPase , partial (96%) 96 5e-20
BE802619 similar to GP|20146763|gb| ATPase alpha subunit {Oryza ... 89 4e-18
TC215545 homologue to GB|AAC49174.1|849136|VRU26709 vacuolar H+-... 80 2e-15
CF920296 38 0.009
BE190965 similar to GP|3676296|gb|A mitochondrial ATPase beta su... 34 0.18
TC214816 similar to UP|Q9M5Z8 (Q9M5Z8) Vacuolar H+-ATPase B subu... 33 0.30
TC214815 similar to UP|VAB1_GOSHI (Q43432) Vacuolar ATP synthase... 33 0.30
TC214811 homologue to UP|VAB2_HORVU (Q40079) Vacuolar ATP syntha... 33 0.40
BM567671 31 1.2
TC213381 30 3.4
TC217444 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like prote... 30 3.4
>TC224836 homologue to UP|ATPA_LOTJA (Q9BBS3) ATP synthase alpha chain ,
complete
Length = 2664
Score = 951 bits (2459), Expect = 0.0
Identities = 500/510 (98%), Positives = 505/510 (98%)
Frame = +1
Query: 1 MVTIRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGT 60
MVTIRADEISKIIRERIEQYNTE+KIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGT
Sbjct: 1009 MVTIRADEISKIIRERIEQYNTEVKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGT 1188
Query: 61 IGIALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRG 120
IGIALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSE YLGRVINALAKPIDGRG
Sbjct: 1189 IGIALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEAYLGRVINALAKPIDGRG 1368
Query: 121 EISSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVA 180
EIS+SESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVA
Sbjct: 1369 EISASESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVA 1548
Query: 181 TDTILNQQGQNVICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAP 240
TDTILNQQGQNVICVYVA+GQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAP
Sbjct: 1549 TDTILNQQGQNVICVYVAIGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAP 1728
Query: 241 YTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLE 300
YTGAALAE+FMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLE
Sbjct: 1729 YTGAALAEYFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLE 1908
Query: 301 RAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINV 360
RAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINV
Sbjct: 1909 RAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINV 2088
Query: 361 GISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLARGQRLREL 420
GISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLARGQRLREL
Sbjct: 2089 GISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLARGQRLREL 2268
Query: 421 LKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNEIISSTK 480
LKQSQSAPLTVEEQ+ITIYTGTNGYLDSLEI QVRKFLVELRAYL TNKPQF EIISSTK
Sbjct: 2269 LKQSQSAPLTVEEQIITIYTGTNGYLDSLEIGQVRKFLVELRAYLNTNKPQFKEIISSTK 2448
Query: 481 TFTGEAEALLKEAIQEQMELFLLQEQVEKN 510
TFTGEAE LLKEAIQEQMELFLLQEQVEKN
Sbjct: 2449 TFTGEAEVLLKEAIQEQMELFLLQEQVEKN 2538
>TC231525 atpA
Length = 1527
Score = 573 bits (1478), Expect = e-164
Identities = 301/505 (59%), Positives = 375/505 (73%), Gaps = 10/505 (1%)
Frame = +1
Query: 3 TIRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGTIG 62
++RA E++ ++ RI + T ++ G V+ VGDGIAR+YGL+E+ AGE+VEF G G
Sbjct: 10 SVRAAELTTLLESRITNFYTNFQVDEIGRVVSVGDGIARVYGLNEIQAGEMVEFASGVKG 189
Query: 63 IALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEI 122
IALNLE++NVG+V+ G I+EG VK TG I +P + LGRV++AL PIDGRG +
Sbjct: 190 IALNLENENVGIVVFGSDTAIKEGDLVKRTGSIVDVPAGKAMLGRVVDALGVPIDGRGAL 369
Query: 123 SSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATD 182
S E R +E APGII R+SV+EP+QTGL A+DS++PIGRGQRELIIGDRQTGKTA+A D
Sbjct: 370 SDHERRRVEVKAPGIIERKSVHEPMQTGLKAVDSLVPIGRGQRELIIGDRQTGKTAIAID 549
Query: 183 TILNQQGQN---------VICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPA 233
TILNQ+ N + CVYVA+GQK S+VAQ+V L E A+EY+I+VA TA PA
Sbjct: 550 TILNQKQMNSRATSESETLYCVYVAIGQKRSTVAQLVQILSEANALEYSILVAATASDPA 729
Query: 234 TLQYLAPYTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFY 293
LQ+LAPY+G A+ E+F H LIIYDDLSKQA AYRQMSLLLRRPPGREA+PGDVFY
Sbjct: 730 PLQFLAPYSGCAMGEYFRDNGMHALIIYDDLSKQAVAYRQMSLLLRRPPGREAFPGDVFY 909
Query: 294 LHSRLLERAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAG 353
LHSRLLERAAK S Q G GS+TALP++ETQ+GDVSAYIPTNVISITDGQI L +LF G
Sbjct: 910 LHSRLLERAAKRSDQTGAGSLTALPVIETQAGDVSAYIPTNVISITDGQICLETELFYRG 1089
Query: 354 IRPAINVGISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEAFAQFASDLDKATQNQLAR 413
IRPAINVG+SVSRVGSAAQ+KAMKQV G LKLELAQ+ E+ AFAQF SDLD ATQ L R
Sbjct: 1090IRPAINVGLSVSRVGSAAQLKAMKQVCGSLKLELAQYREVAAFAQFGSDLDAATQALLNR 1269
Query: 414 GQRLRELLKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFN 473
G RL E+LKQ Q APL +E+Q++ IY NG+ D + + ++ ++ R L T KP
Sbjct: 1270GARLTEVLKQPQYAPLPIEKQILVIYAAVNGFCDRMPLDKIPQY---ERDILTTIKP--- 1431
Query: 474 EIISSTK-TFTGEAEALLKEAIQEQ 497
E++ S K T E + L++ ++E+
Sbjct: 1432ELLQSLKGGLTSERKIELEKFLKEK 1506
>NP005377 pseudo-atpA
Length = 981
Score = 238 bits (608), Expect = 4e-63
Identities = 119/214 (55%), Positives = 155/214 (71%), Gaps = 9/214 (4%)
Frame = +1
Query: 3 TIRADEISKIIRERIEQYNTEIKIVNTGTVLQVGDGIARIYGLDEVMAGELVEFEEGTIG 62
++RA E++ ++ RI + T ++ G V+ VGDGIAR+YGL+E+ AGE+VEF G G
Sbjct: 10 SVRAAELTTLLESRITNFYTNFQVDEIGRVVSVGDGIARVYGLNEIQAGEMVEFASGVKG 189
Query: 63 IALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEI 122
IALNLE++NVG+V+ G I+EG VK TG I +P + LGRV++AL PIDGRG +
Sbjct: 190 IALNLENENVGIVVFGSDTAIKEGDLVKRTGSIVDVPAGKAMLGRVVDALGVPIDGRGAL 369
Query: 123 SSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATD 182
S E R +E APGII R+SV+EP+QTGL A+DS++PIGRGQRELIIGDRQTGKTA+A D
Sbjct: 370 SDHERRRVEVKAPGIIERKSVHEPMQTGLKAVDSLVPIGRGQRELIIGDRQTGKTAIAID 549
Query: 183 TILNQQGQN---------VICVYVAVGQKASSVA 207
TILNQ+ N + CVYVA+GQK S+VA
Sbjct: 550 TILNQKQMNSRATSESETLYCVYVAIGQKRSTVA 651
>BE346561 similar to SP|Q9BBS3|ATPA ATP synthase alpha chain (EC 3.6.3.14).
{Lotus japonicus}, partial (33%)
Length = 511
Score = 234 bits (598), Expect = 5e-62
Identities = 133/170 (78%), Positives = 136/170 (79%)
Frame = -1
Query: 336 ISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAAQIKAMKQVAGKLKLELAQFAELEA 395
ISITDGQIFLS DLF+ GIR INVGISVSRV S A I AMK VAGKLKLEL FA+LE
Sbjct: 511 ISITDGQIFLSVDLFHVGIRLVINVGISVSRVESLAYITAMKSVAGKLKLELV*FAQLEV 332
Query: 396 FAQFASDLDKATQNQLARGQRLRELLKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVR 455
F QFASD D AT N L RG L EL KQ A LTVEEQ+I IYTG NGY DSLEI QVR
Sbjct: 331 FVQFASDFDTAT*N*LVRG**LCELXKQF*LALLTVEEQIIIIYTGMNGYFDSLEIGQVR 152
Query: 456 KFLVELRAYLKTNKPQFNEIISSTKTFTGEAEALLKEAIQEQMELFLLQE 505
KFLVELR YL TNKPQF EIISSTKTFTGEAE LLKEAIQEQMELFLLQE
Sbjct: 151 KFLVELRDYLNTNKPQFKEIISSTKTFTGEAEVLLKEAIQEQMELFLLQE 2
>TC214812 homologue to UP|Q9ZQS1 (Q9ZQS1) Vacuolar H+-ATPase B subunit ,
complete
Length = 1777
Score = 124 bits (310), Expect = 1e-28
Identities = 102/347 (29%), Positives = 157/347 (44%), Gaps = 30/347 (8%)
Frame = +3
Query: 55 EFEEGTIGIALNLESKNVGVVLMGDG----LMIQEGSS--------VKATGRIAQIPVSE 102
+F+E + I L + G VL DG + + EG+S V+ TG + + PVS
Sbjct: 252 KFQE-IVNIRLGDGTTRRGQVLEVDGEKAVVQVFEGTSGIDNKFTTVQFTGEVLKTPVSL 428
Query: 103 GYLGRVINALAKPIDGRGEISSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGR 162
LGR+ N KPID I I + R E +QTG+ ID M I R
Sbjct: 429 DMLGRIFNGSGKPIDNGPPILPEAYLDISGSSINPSERTYPEEMIQTGISTIDVMNSIAR 608
Query: 163 GQRELIIGD-------------RQTG--KTAVATDTILNQQGQ--NVICVYVAVGQKASS 205
GQ+ + RQ G K +D +L G+ N V+ A+G +
Sbjct: 609 GQKIPLFSAAGLPHNEIAAQICRQAGLVKRLEKSDNLLEGGGEEDNFAIVFAAMGVNMET 788
Query: 206 VAQVVNTLQERGAMEYTIVVAETADSPATLQYLAPYTGAALAEFFMYR-ERHTLIIYDDL 264
+E G+ME + A+ P + + P AE+ Y +H L+I D+
Sbjct: 789 AQFFKRDFEENGSMERVTLFLNLANDPTIERIITPRIALTTAEYLAYECGKHVLVILTDM 968
Query: 265 SKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSSQLGEGSMTALPIVETQS 324
S A A R++S PGR YPG ++ + + ERA ++ + +GS+T +PI+ +
Sbjct: 969 SSYADALREVSAAREEVPGRRGYPGYMYTDLATIYERAGRIEGR--KGSITQIPILTMPN 1142
Query: 325 GDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAA 371
D++ P IT+GQI++ L+N I P INV S+SR+ +A
Sbjct: 1143DDITHPTPDLTGYITEGQIYIDRQLYNRQIYPPINVLPSLSRLMKSA 1283
>BF423880 homologue to GP|20068316|emb ATPase alpha subunit {Atropa
belladonna}, partial (11%)
Length = 401
Score = 112 bits (279), Expect = 5e-25
Identities = 57/59 (96%), Positives = 58/59 (97%)
Frame = +1
Query: 404 DKATQNQLARGQRLRELLKQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELR 462
DKATQNQLARGQRLRELLKQSQSAPLTVEEQ+ITIYTGTNGYLDSLEI QVRKFLVELR
Sbjct: 1 DKATQNQLARGQRLRELLKQSQSAPLTVEEQIITIYTGTNGYLDSLEIGQVRKFLVELR 177
>BE556680 weakly similar to GP|14715604|gb| ATP synthase alpha subunit
{Clostridium pasteurianum}, partial (8%)
Length = 283
Score = 102 bits (253), Expect = 5e-22
Identities = 56/93 (60%), Positives = 67/93 (71%)
Frame = +2
Query: 230 DSPATLQYLAPYTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPG 289
DS AT QYLA YT +ALAE++M RER+TLI+ D+LSK A AY QMS+LLRRPP YP
Sbjct: 5 DSSATSQYLASYT*SALAEYYMCRERNTLIMNDNLSKLAHAYLQMSVLLRRPPAL*DYPR 184
Query: 290 DVFYLHSRLLERAAKLSSQLGEGSMTALPIVET 322
D FYLH+RLL A L + E MT LP+V+T
Sbjct: 185 DRFYLHTRLL*IAGNLILHVDERRMTDLPLVDT 283
>TC214814 homologue to UP|Q8GUB5 (Q8GUB5) Vacuolar ATPase subunit B ,
partial (76%)
Length = 1639
Score = 100 bits (250), Expect = 1e-21
Identities = 72/244 (29%), Positives = 115/244 (46%), Gaps = 17/244 (6%)
Frame = +2
Query: 145 EPLQTGLIAIDSMIPIGRGQRELIIGD-------------RQTG--KTAVATDTILNQ-Q 188
E +QTG+ ID M I RGQ+ + RQ G K +D +L +
Sbjct: 86 EMIQTGISTIDVMNSIARGQKIPLFSAAGLPHNEIAAQICRQAGLVKRLEKSDNLLEGGE 265
Query: 189 GQNVICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAETADSPATLQYLAPYTGAALAE 248
N V+ A+G + +E G+ME + A+ P + + P AE
Sbjct: 266 EDNFAIVFAAMGVNMETAQFFKRDFEENGSMERVTLFLNLANDPTIERIITPRIALTTAE 445
Query: 249 FFMYR-ERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSS 307
+ Y +H L+I D+S A A R++S PGR YPG ++ + + ERA ++
Sbjct: 446 YLAYECGKHVLVILTDMSSYADALREVSAAREEVPGRRGYPGYMYTDLATIYERAGRIEG 625
Query: 308 QLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRV 367
+ +GS+T +PI+ + D++ P IT+GQI++ L+N I P INV S+SR+
Sbjct: 626 R--KGSITQIPILTMPNDDITHPTPDLTGYITEGQIYIDRQLYNRQIYPPINVLPSLSRL 799
Query: 368 GSAA 371
+A
Sbjct: 800 MKSA 811
>TC224613 homologue to UP|Q95FJ6 (Q95FJ6) ATP synthase beta subunit, complete
Length = 2810
Score = 100 bits (248), Expect = 2e-21
Identities = 105/427 (24%), Positives = 179/427 (41%), Gaps = 12/427 (2%)
Frame = +3
Query: 79 DGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEISSSESRLIESPAPGII 138
+GLM G V TG +PV LGR+ N L +PID G + + + I AP I
Sbjct: 924 EGLM--RGMEVIDTGAALSVPVGGATLGRIFNVLGEPIDNLGPVDTRTTSPIHRSAPAFI 1097
Query: 139 SRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATDTILN-QQGQNVICVYV 197
+ +TG+ +D + P RG + + G GKT + + I N + + V+
Sbjct: 1098 QLDTKLAIFETGIKVVDLLAPYRRGGKIGLFGGAGVGKTVLIMELINNIAKAHGGVSVFG 1277
Query: 198 AVGQKASSVAQVVNTLQERGAM-EYTIVVAETA------DSPATLQYLAPYTGAALAEFF 250
VG++ + ++E G + E I ++ A + P + T +AE+F
Sbjct: 1278 GVGERTREGNDLYMEMKESGVINEQNIAESKVALVYGQMNEPPGARMRVGLTALTMAEYF 1457
Query: 251 M-YRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRLLERAAKLSSQL 309
E+ L+ D++ + QA ++S LL R P Y + L ER
Sbjct: 1458 RDVNEQDVLLFIDNIFRFVQAGSEVSALLGRMPSAVGYQPTLSTEMGSLQERITSTK--- 1628
Query: 310 GEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRVGS 369
EGS+T++ V + D++ P + D LS L GI PA++ S S +
Sbjct: 1629 -EGSITSIQAVYVPADDLTDPAPATTFAHLDATTVLSRGLAAKGIYPAVDPLDSTSTMLQ 1805
Query: 370 AAQI-KAMKQVAGKLKLELAQFAELEAFAQF--ASDLDKATQNQLARGQRLRELLKQSQS 426
+ + + A ++K L ++ EL+ +L + + +AR +++ L Q
Sbjct: 1806 PRIVGEEHYETAQRVKQTLQRYKELQDIIAILGLDELSEEDRLTVARARKIERFLSQ--- 1976
Query: 427 APLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNEIISSTKTFTGEA 486
P V E ++TG+ G SL +R F + L L Q ++ + T +A
Sbjct: 1977 -PFFVAE----VFTGSPGKYVSL-AETIRGFKLILSGELDGLPEQAFYLVGNIDEATAKA 2138
Query: 487 EALLKEA 493
L E+
Sbjct: 2139 TNLEMES 2159
>TC225459 homologue to UP|O04275 (O04275) F1 ATPase , partial (96%)
Length = 2079
Score = 95.5 bits (236), Expect = 5e-20
Identities = 94/395 (23%), Positives = 166/395 (41%), Gaps = 13/395 (3%)
Frame = +2
Query: 63 IALNLESKNVGVVLMGDGLMIQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEI 122
+A +L V + M + G V TG +PV LGR+IN + +PID +GEI
Sbjct: 419 VAQHLGEGVVRTIAMDATEGVVRGWRVLNTGSPITVPVGRATLGRIINVIGEPIDDKGEI 598
Query: 123 SSSESRLIESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATD 182
++ I AP + + + + L TG+ +D + P RG + + G GKT + +
Sbjct: 599 NTEHYLPIHREAPAFVEQETAQQILVTGIKVVDLLAPYQRGGKIGLFGGAGVGKTVLIME 778
Query: 183 TILN-QQGQNVICVYVAVGQKASSVAQVVNTLQERGAMEYTIVVAET--------ADSPA 233
I N + V+ VG++ + + E G ++ +E+ + P
Sbjct: 779 LINNVAKAHGGFSVFAGVGERTREGNDLYREMIESGVIKLDDKQSESKCALVYGQMNEPP 958
Query: 234 TLQYLAPYTGAALAEFFMYRE-RHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVF 292
+ TG +AE F E + L+ D++ + QA ++S LL R P Y +
Sbjct: 959 GARARVGLTGLTVAEHFRDAEGQDVLLFVDNIFRFTQANSEVSALLGRIPSAVGYQPTLS 1138
Query: 293 YLHSRLLERAAKLSSQLGEGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSADLFNA 352
L ER +GS+T++ + + D++ P + D LS +
Sbjct: 1139TDLGALQERITTTK----KGSITSVQAIYVPADDLTDPAPATTFAHLDATTVLSRQISEL 1306
Query: 353 GIRPAINVGISVSRVGSAAQIKA-MKQVAGKLKLELAQFAELEAFAQF--ASDLDKATQN 409
GI PA++ S SR+ S + A + A ++ L + L+ +L + +
Sbjct: 1307GIYPAVDPLDSTSRMLSPLILGADHYETARGVQKVLQNYKNLQDIIAILGMDELSEDDKL 1486
Query: 410 QLARGQRLRELLKQSQSAPLTVEEQVITIYTGTNG 444
+AR ++++ L Q P V E ++TG G
Sbjct: 1487TVARARKIQRFLSQ----PFHVAE----VFTGAPG 1567
>BE802619 similar to GP|20146763|gb| ATPase alpha subunit {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 169
Score = 89.4 bits (220), Expect = 4e-18
Identities = 43/55 (78%), Positives = 49/55 (88%)
Frame = +2
Query: 234 TLQYLAPYTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYP 288
TLQYLAPYT AA+AE++M+RERHTL+I DDLS Q QAYRQMSLLLRRPP R+A P
Sbjct: 2 TLQYLAPYT*AAMAEYYMFRERHTLLINDDLS*QTQAYRQMSLLLRRPPDRKAMP 166
>TC215545 homologue to GB|AAC49174.1|849136|VRU26709 vacuolar H+-ATPase
subunit A {Vigna radiata;} , complete
Length = 2358
Score = 80.1 bits (196), Expect = 2e-15
Identities = 74/261 (28%), Positives = 121/261 (46%), Gaps = 18/261 (6%)
Frame = +1
Query: 130 IESPAPGIISRRSVYEPLQTGLIAIDSMIPIGRGQRELIIGDRQTGKTAVATDTILNQQG 189
+ +P P + S+ + PL TG +D++ P G I G GKT ++ L++
Sbjct: 691 VRTPRP-VASKLAADTPLLTGQRVLDALFPSVLGGTCAIPGAFGCGKTVISQ--ALSKYS 861
Query: 190 QNVICVYVAVGQKASSVAQVVNTL----------QERGAMEYTIVVAETADSPATLQYLA 239
+ VYV G++ + +A+V+ +E M+ T +VA T++ P + +
Sbjct: 862 NSDAVVYVGCGERGNEMAEVLMDFPQLTMTLPDGREESVMKRTTLVANTSNMPVAAREAS 1041
Query: 240 PYTGAALAEFFMYRERHTLIIYDDLSKQAQAYRQMSLLLRRPPGREAYPGDVFYLHSRL- 298
YTG LAE+F + ++ D S+ A+A R++S L P YP YL +RL
Sbjct: 1042IYTGITLAEYFRDMGYNVSMMADSTSRWAEALREISGRLAEMPADSGYPA---YLAARLA 1212
Query: 299 --LERAAKLSSQLG---EGSMTALPIVETQSGDVSAYIPTNVISITDGQIFLSAD--LFN 351
ERA K+ G GS+T + V GD S + + +SI Q+F D L
Sbjct: 1213SFYERAGKVKCLGGPERTGSVTIVGAVSPPGGDFSDPVTSATLSIV--QVFWGLDKKLAQ 1386
Query: 352 AGIRPAINVGISVSRVGSAAQ 372
P++N IS S+ +A +
Sbjct: 1387RKHFPSVNWLISYSKYSTALE 1449
>CF920296
Length = 518
Score = 38.1 bits (87), Expect = 0.009
Identities = 21/59 (35%), Positives = 34/59 (57%)
Frame = +3
Query: 422 KQSQSAPLTVEEQVITIYTGTNGYLDSLEIRQVRKFLVELRAYLKTNKPQFNEIISSTK 480
KQ Q APL +E+Q++ IY NG+ D + + ++ ++ R L T KP E++ S K
Sbjct: 3 KQPQYAPLPIEKQILVIYAAVNGFCDRMPLDKIPQY---ERDILTTIKP---ELLQSLK 161
>BE190965 similar to GP|3676296|gb|A mitochondrial ATPase beta subunit
{Nicotiana sylvestris}, partial (10%)
Length = 181
Score = 33.9 bits (76), Expect = 0.18
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = +2
Query: 83 IQEGSSVKATGRIAQIPVSEGYLGRVINALAKPIDGRGEI 122
+ G V TG +P+ LGR+IN + +PID GEI
Sbjct: 62 VVRGCRVINTGSPITVPLVMATLGRIINVIGEPIDSTGEI 181
>TC214816 similar to UP|Q9M5Z8 (Q9M5Z8) Vacuolar H+-ATPase B subunit, partial
(45%)
Length = 799
Score = 33.1 bits (74), Expect = 0.30
Identities = 26/75 (34%), Positives = 37/75 (48%), Gaps = 12/75 (16%)
Frame = +1
Query: 55 EFEEGTIGIALNLESKNVGVVLMGDG----LMIQEGSS--------VKATGRIAQIPVSE 102
+F+E + I L + G VL DG + + EG+S V+ TG + + PVS
Sbjct: 253 KFQE-IVNIRLGDGTTRRGQVLEVDGEKAVVQVFEGTSGIDNKFTTVQFTGEVLKTPVSL 429
Query: 103 GYLGRVINALAKPID 117
LGR+ N KPID
Sbjct: 430 DMLGRIFNGSGKPID 474
>TC214815 similar to UP|VAB1_GOSHI (Q43432) Vacuolar ATP synthase subunit B
isoform 1 (V-ATPase B subunit 1) (Vacuolar proton pump
B subunit 1) , partial (35%)
Length = 619
Score = 33.1 bits (74), Expect = 0.30
Identities = 26/75 (34%), Positives = 37/75 (48%), Gaps = 12/75 (16%)
Frame = +2
Query: 55 EFEEGTIGIALNLESKNVGVVLMGDG----LMIQEGSS--------VKATGRIAQIPVSE 102
+F+E + I L + G VL DG + + EG+S V+ TG + + PVS
Sbjct: 224 KFQE-IVNIRLGDGTTRRGQVLEVDGEKAVVQVFEGTSGIDNKFTTVQFTGEVLKTPVSL 400
Query: 103 GYLGRVINALAKPID 117
LGR+ N KPID
Sbjct: 401 DMLGRIFNGSGKPID 445
>TC214811 homologue to UP|VAB2_HORVU (Q40079) Vacuolar ATP synthase subunit B
isoform 2 (V-ATPase B subunit 2) (Vacuolar proton pump
B subunit 2) , partial (31%)
Length = 789
Score = 32.7 bits (73), Expect = 0.40
Identities = 18/46 (39%), Positives = 27/46 (58%)
Frame = +2
Query: 326 DVSAYIPTNVISITDGQIFLSADLFNAGIRPAINVGISVSRVGSAA 371
D++ P IT+GQI++ L+N I P INV S+SR+ +A
Sbjct: 5 DITHPTPDLTGYITEGQIYIDRQLYNRQIYPPINVLPSLSRLMKSA 142
>BM567671
Length = 428
Score = 31.2 bits (69), Expect = 1.2
Identities = 19/54 (35%), Positives = 27/54 (49%), Gaps = 3/54 (5%)
Frame = +3
Query: 147 LQTGLIAIDSMIPIG---RGQRELIIGDRQTGKTAVATDTILNQQGQNVICVYV 197
+ TG A+D + IG +G+ I G +GKT +A I Q Q CV+V
Sbjct: 267 VSTGSFALDVALGIGGLPKGRVVEIYGPEASGKTTLALHVIAEAQKQGGYCVFV 428
>TC213381
Length = 421
Score = 29.6 bits (65), Expect = 3.4
Identities = 16/60 (26%), Positives = 32/60 (52%), Gaps = 1/60 (1%)
Frame = -1
Query: 231 SPATLQYLAPYTGAALAEFFMYRERHTLIIY-DDLSKQAQAYRQMSLLLRRPPGREAYPG 289
+P ++++ ++ A+ + + +E TL ++ DDL + YR S LLR+ R+ G
Sbjct: 406 TPLAMRFICAFSNASSLKSLIGKEPTTLFLFHDDLCPRITEYRTNSALLRKLTDRKGSDG 227
>TC217444 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like protein
(AT5g59010/k19m22_210), partial (91%)
Length = 1655
Score = 29.6 bits (65), Expect = 3.4
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = -2
Query: 104 YLGRVINALAKPIDGRGEISSSESRL 129
Y G I+ L ++GRGE+SS ESR+
Sbjct: 1336 YHGSTIDELCIAVNGRGEVSSMESRI 1259
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.134 0.355
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,273,412
Number of Sequences: 63676
Number of extensions: 135269
Number of successful extensions: 602
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 593
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 595
length of query: 510
length of database: 12,639,632
effective HSP length: 101
effective length of query: 409
effective length of database: 6,208,356
effective search space: 2539217604
effective search space used: 2539217604
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Description of BAB33201.1

