
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33190.1 473 aa
(473 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
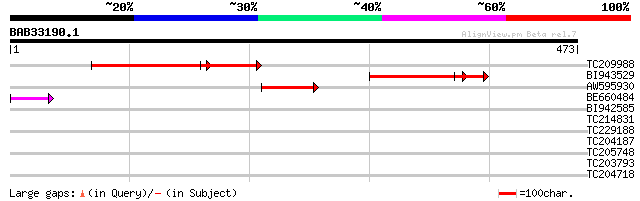
Score E
Sequences producing significant alignments: (bits) Value
TC209988 UP|Q6SSE1 (Q6SSE1) Photosystem II CP43 protein, partial... 196 4e-75
BI943529 GP|11022865|gb photosystem II CP43 protein {Acorus cala... 154 6e-38
AW595930 GP|11022865|gb photosystem II CP43 protein {Acorus cala... 106 3e-23
BE660484 homologue to GP|11022864|gb photosystem II D2 protein {... 44 2e-04
BI942585 homologue to GP|27446507|gb photosystem II CP47 protein... 35 0.095
TC214831 weakly similar to UP|Q8LPQ0 (Q8LPQ0) At1g17450/F1L3.20,... 31 1.4
TC229188 29 4.0
TC204187 homologue to UP|TBB6_ARATH (P29514) Tubulin beta-6 chai... 29 5.2
TC205748 similar to GB|AAR04550.1|37813554|AY337455 dual-specifi... 28 6.8
TC203793 similar to PIR|T02602|T02602 vacuolar sorting receptor ... 28 8.9
TC204718 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17... 28 8.9
>TC209988 UP|Q6SSE1 (Q6SSE1) Photosystem II CP43 protein, partial (30%)
Length = 427
Score = 196 bits (499), Expect(2) = 4e-75
Identities = 94/100 (94%), Positives = 95/100 (95%)
Frame = +1
Query: 69 LFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLISSAVLGFG 128
LFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLISSAVLGFG
Sbjct: 1 LFEVAHFVPEKPMYEQGLILLPHLATLGWGVGPGGEVIDTFPYFVSGVLHLISSAVLGFG 180
Query: 129 GIYHALLGPETLEESFPFFGYVWKDRNKMTTILGIHLILL 168
GIYHALLGPETLEESFPFFGYVWKDRNKMTT G L L+
Sbjct: 181 GIYHALLGPETLEESFPFFGYVWKDRNKMTTNFGYSLNLV 300
Score = 103 bits (257), Expect(2) = 4e-75
Identities = 50/51 (98%), Positives = 50/51 (98%)
Frame = +2
Query: 160 ILGIHLILLGIGAFLLVFKASYFGGIYDTWAPGGGDVRKITNLTLSPSIIF 210
ILGIHLILLGIGAFLLVFKA YFGGIYDTWAPGGGDVRKITNLTLSPSIIF
Sbjct: 275 ILGIHLILLGIGAFLLVFKALYFGGIYDTWAPGGGDVRKITNLTLSPSIIF 427
>BI943529 GP|11022865|gb photosystem II CP43 protein {Acorus calamus},
partial (23%)
Length = 299
Score = 154 bits (390), Expect = 6e-38
Identities = 74/82 (90%), Positives = 75/82 (91%)
Frame = +1
Query: 301 FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD 360
FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD
Sbjct: 1 FYGPTGPEASQAQAFTFLVRDQRLGANVGSAQGPTGLGKYLMRSPTGEVIFGGETMRFWD 180
Query: 361 LRAPWLEPLRGPNGLDLSRLKK 382
LRAPWLEPLRGP L +K
Sbjct: 181 LRAPWLEPLRGPKWFRLE*TEK 246
Score = 62.8 bits (151), Expect = 3e-10
Identities = 28/28 (100%), Positives = 28/28 (100%)
Frame = +2
Query: 372 PNGLDLSRLKKDIQPWQERRSAEYMTHA 399
PNGLDLSRLKKDIQPWQERRSAEYMTHA
Sbjct: 215 PNGLDLSRLKKDIQPWQERRSAEYMTHA 298
>AW595930 GP|11022865|gb photosystem II CP43 protein {Acorus calamus},
partial (11%)
Length = 141
Score = 106 bits (264), Expect = 3e-23
Identities = 47/47 (100%), Positives = 47/47 (100%)
Frame = +1
Query: 211 GYLLKSPFGGEGWIVSVDDLEDIIGGHVWLGSICILGGIWHILTKPF 257
GYLLKSPFGGEGWIVSVDDLEDIIGGHVWLGSICILGGIWHILTKPF
Sbjct: 1 GYLLKSPFGGEGWIVSVDDLEDIIGGHVWLGSICILGGIWHILTKPF 141
>BE660484 homologue to GP|11022864|gb photosystem II D2 protein {Acorus
calamus}, partial (66%)
Length = 723
Score = 43.9 bits (102), Expect = 2e-04
Identities = 21/37 (56%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Frame = +2
Query: 1 MKTLYSLRRFYHVETLFNGTLALTGR-DQETTGFAWW 36
MKTLYSLR YHVETLFNG + GF WW
Sbjct: 599 MKTLYSLRXVYHVETLFNGNFSFNWS*PGNLPGFLWW 709
>BI942585 homologue to GP|27446507|gb photosystem II CP47 protein {Pisum
sativum}, partial (22%)
Length = 408
Score = 34.7 bits (78), Expect = 0.095
Identities = 19/73 (26%), Positives = 27/73 (36%), Gaps = 2/73 (2%)
Frame = -1
Query: 394 EYMTHAPLGSLNSVGGVATEINAVNYVSPRSWLATSHFVLGFFLFVGHLWHAGRA--RAA 451
+Y A G + + + + V S R W H F GH+WH R R
Sbjct: 333 KYARRAQ*GEIFELDRATLKSDGVFRSSQRGWFTLGHASFALLFFFGHIWHGERTLYRDV 154
Query: 452 AAGFEKGIDRDFE 464
AG + +D E
Sbjct: 153 FAGIDPDLDAQVE 115
>TC214831 weakly similar to UP|Q8LPQ0 (Q8LPQ0) At1g17450/F1L3.20, partial (28%)
Length = 1599
Score = 30.8 bits (68), Expect = 1.4
Identities = 16/43 (37%), Positives = 24/43 (55%)
Frame = +3
Query: 402 GSLNSVGGVATEINAVNYVSPRSWLATSHFVLGFFLFVGHLWH 444
G+L + G +I + PR++L SH +FL VG+LWH
Sbjct: 1209 GTLLTSGTYRKQIQSTEVDMPRAFLRQSH---EYFLVVGNLWH 1328
>TC229188
Length = 882
Score = 29.3 bits (64), Expect = 4.0
Identities = 18/42 (42%), Positives = 22/42 (51%)
Frame = +3
Query: 396 MTHAPLGSLNSVGGVATEINAVNYVSPRSWLATSHFVLGFFL 437
MT P GSL SVG + +NA+N + L TS LG L
Sbjct: 54 MTRVPWGSLESVGDQSEYVNAINLI-----LTTSIPALGSLL 164
>TC204187 homologue to UP|TBB6_ARATH (P29514) Tubulin beta-6 chain, partial
(98%)
Length = 1882
Score = 28.9 bits (63), Expect = 5.2
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = -1
Query: 410 VATEINAVNYVSPRSWLATSHFVLGFFLFVGHLWH 444
+ATE V+P W+ ++H VLG + LWH
Sbjct: 1012 LATEHG*CREVTPVPWICSTHHVLGIPHLLCQLWH 908
>TC205748 similar to GB|AAR04550.1|37813554|AY337455 dual-specificity
phosphatase-like protein {Arabidopsis thaliana;} ,
partial (77%)
Length = 1419
Score = 28.5 bits (62), Expect = 6.8
Identities = 22/67 (32%), Positives = 30/67 (43%)
Frame = -3
Query: 187 DTWAPGGGDVRKITNLTLSPSIIFGYLLKSPFGGEGWIVSVDDLEDIIGGHVWLGSICIL 246
+ W PGGG ++ + LS S+I + GEG D+ I W+ ICI
Sbjct: 1141 EIWQPGGGTLQTLPQDKLSSSLI------TQVAGEGSRTIHLDISAI-----WICWICIK 995
Query: 247 GGIWHIL 253
G HIL
Sbjct: 994 GISSHIL 974
>TC203793 similar to PIR|T02602|T02602 vacuolar sorting receptor protein
homolog At2g14740 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (31%)
Length = 711
Score = 28.1 bits (61), Expect = 8.9
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = -1
Query: 424 SWLATSHFVLGFFLFVGHL 442
S A++HF+L FF F+GHL
Sbjct: 207 SMTASAHFLLYFFSFIGHL 151
>TC204718 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17H15.1
{Arabidopsis thaliana;} , partial (5%)
Length = 1417
Score = 28.1 bits (61), Expect = 8.9
Identities = 18/55 (32%), Positives = 28/55 (50%), Gaps = 4/55 (7%)
Frame = -1
Query: 236 GH--VWLGSICILGGIW-HILTKPFAWARRALVWS-GEAYLSYSLAALSVFGFIA 286
GH WL CI+G IW +L P R WS G L + +++++G++A
Sbjct: 589 GHPITWLSLWCIVGSIWARLLASPITLTWRLGRWSLG*IPLGSVIGSVALWGWVA 425
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.143 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,813,093
Number of Sequences: 63676
Number of extensions: 377107
Number of successful extensions: 1832
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1817
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1832
length of query: 473
length of database: 12,639,632
effective HSP length: 101
effective length of query: 372
effective length of database: 6,208,356
effective search space: 2309508432
effective search space used: 2309508432
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Description of BAB33190.1

