
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33187.1 734 aa
(734 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
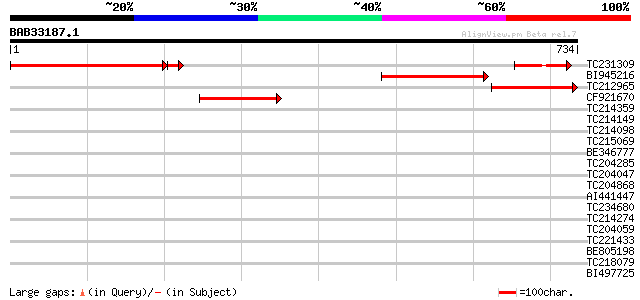
Score E
Sequences producing significant alignments: (bits) Value
TC231309 homologue to UP|PSAB_LOTJA (P58385) Photosystem I P700 ... 416 e-116
BI945216 SP|P06407|PSAB Photosystem I P700 chlorophyll A apoprot... 299 3e-81
TC212965 UP|PSAB_ANTMA (Q33332) Photosystem I P700 chlorophyll A... 236 3e-62
CF921670 210 2e-54
TC214359 homologue to UP|PRP1_SOYBN (P08012) Repetitive proline-... 39 0.006
TC214149 homologue to UP|PRP1_SOYBN (P08012) Repetitive proline-... 37 0.024
TC214098 UP|PRP1_SOYBN (P08012) Repetitive proline-rich cell wal... 37 0.031
TC215069 similar to UP|O24300 (O24300) PtxA protein precursor, p... 33 0.45
BE346777 33 0.59
TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 33 0.59
TC204047 hydroxyproline-rich glycoprotein 32 0.77
TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, par... 32 1.3
AI441447 SP|P08012|PRP1 Repetitive proline-rich cell wall protei... 31 2.3
TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor... 30 3.8
TC214274 homologue to UP|Q00508 (Q00508) Ribosomal protein L41, ... 30 5.0
TC204059 similar to UP|Q94ES3 (Q94ES3) Root nodule extensin (Fra... 30 5.0
TC221433 weakly similar to UP|Q7EYQ4 (Q7EYQ4) DnaJ protein famil... 30 5.0
BE805198 similar to PIR|T06455|T064 Myb26 protein - garden pea, ... 30 5.0
TC218079 similar to UP|Q945B9 (Q945B9) Growth-on protein GRO11, ... 29 6.5
BI497725 homologue to SP|P13993|PRP2 Repetitive proline-rich cel... 29 6.5
>TC231309 homologue to UP|PSAB_LOTJA (P58385) Photosystem I P700 chlorophyll
A apoprotein A2 (PsaB) (PSI-B), partial (27%)
Length = 910
Score = 416 bits (1069), Expect = e-116
Identities = 194/204 (95%), Positives = 196/204 (95%)
Frame = +3
Query: 1 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLAIIFLW 60
MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQL IIFLW
Sbjct: 264 MALRFPRFSQGLAQDPTTRRIWFGIATAHDFESHDDITEERLYQNIFASHFGQLVIIFLW 443
Query: 61 TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 120
TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV
Sbjct: 444 TSGNLFHVAWQGNFEAWVQDPLHVRPIAHAIWDPHFGQPAVEAFTRGGALGPVNIAYSGV 623
Query: 121 YQWWYTIGLRTNGDLYSGALFLLFLSAISLLAGWLHLQPKWKPSVSWFKNAESRLNHHLS 180
YQWWYTIGLRTNGDLY+GALFLLFLS ISL+AGWLHLQPKWKPSVSWFKNAESRLNHHLS
Sbjct: 624 YQWWYTIGLRTNGDLYTGALFLLFLSTISLIAGWLHLQPKWKPSVSWFKNAESRLNHHLS 803
Query: 181 GLFGVSSLAWTGHLVHVAIPGSRG 204
GLFGVSSLAWTGHLVH P G
Sbjct: 804 GLFGVSSLAWTGHLVHGPNPSG*G 875
Score = 74.7 bits (182), Expect = 1e-13
Identities = 38/74 (51%), Positives = 47/74 (63%)
Frame = +2
Query: 654 HLVWATGFMFLISWRGYWQELIETLAWAHERTPLANLIRWRDKPVALSIVQARLVGLAHF 713
H VWA MFL S RGYWQELIE++ WAH + +A +P ALSIVQ R VG+ H+
Sbjct: 8 HFVWAFSLMFLFSGRGYWQELIESIVWAHNKLKVAP----ATQPRALSIVQGRAVGVTHY 175
Query: 714 SVGYIFTYAAFLIA 727
+G I T AF +A
Sbjct: 176 LLGGIATTWAFFLA 217
Score = 45.8 bits (107), Expect = 7e-05
Identities = 18/20 (90%), Positives = 19/20 (95%)
Frame = -1
Query: 205 EYVRWNNFLSILPHPQGLGP 224
EYVRWNN LSILPHP+GLGP
Sbjct: 910 EYVRWNNLLSILPHPEGLGP 851
>BI945216 SP|P06407|PSAB Photosystem I P700 chlorophyll A apoprotein A2
(PsaB) (PSI-B). [Common tobacco] {Nicotiana tabacum},
partial (18%)
Length = 419
Score = 299 bits (765), Expect = 3e-81
Identities = 135/139 (97%), Positives = 137/139 (98%)
Frame = +2
Query: 482 NSPAFNAGRSIWLPGWLNAINENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALDA 541
+ PAFNAGRSIWLPGWLNA+NENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALDA
Sbjct: 2 HEPAFNAGRSIWLPGWLNAVNENSNSLFLTIGPGDFLVHHAIALGLHTTTLILVKGALDA 181
Query: 542 RGSKLMPDKKDFGYSFPCDGPGRGGTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHITL 601
RGSKLMPDKKDFGYSFPCDGPGRG TCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHITL
Sbjct: 182 RGSKLMPDKKDFGYSFPCDGPGRGRTCDISAWDAFYLAVFWMLNTIGWVTFYWHWKHITL 361
Query: 602 WQGNVSQFNESSTYLMGWL 620
WQGNVSQFNESSTYLMGWL
Sbjct: 362 WQGNVSQFNESSTYLMGWL 418
>TC212965 UP|PSAB_ANTMA (Q33332) Photosystem I P700 chlorophyll A apoprotein
A2 (PsaB) (PSI-B), partial (15%)
Length = 466
Score = 236 bits (602), Expect = 3e-62
Identities = 111/111 (100%), Positives = 111/111 (100%)
Frame = +3
Query: 624 LWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWAHE 683
LWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWAHE
Sbjct: 3 LWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWAHE 182
Query: 684 RTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFTYAAFLIASTSGKFG 734
RTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFTYAAFLIASTSGKFG
Sbjct: 183 RTPLANLIRWRDKPVALSIVQARLVGLAHFSVGYIFTYAAFLIASTSGKFG 335
>CF921670
Length = 727
Score = 210 bits (534), Expect = 2e-54
Identities = 101/106 (95%), Positives = 103/106 (96%)
Frame = -3
Query: 246 TSQGAGTAILTLLGGFHPQTQSLWLTDMAHHHLAIAILFLLAGHMYRTNFGIGHSIKDLL 305
T QGAGTAILTLLGGFHPQTQSLWLTD+AHHHLAIA LFL+AGHMYRTNFGIGHSIKDLL
Sbjct: 320 TPQGAGTAILTLLGGFHPQTQSLWLTDIAHHHLAIAFLFLVAGHMYRTNFGIGHSIKDLL 141
Query: 306 EAHIPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQH 351
EAH PPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQH
Sbjct: 140 EAHTPPGGRLGRGHKGLYDTINNSIHFQLGLALASLGVITSLVAQH 3
>TC214359 homologue to UP|PRP1_SOYBN (P08012) Repetitive proline-rich cell
wall protein 1 precursor, partial (52%)
Length = 414
Score = 39.3 bits (90), Expect = 0.006
Identities = 27/101 (26%), Positives = 41/101 (39%), Gaps = 3/101 (2%)
Frame = -3
Query: 570 ISAWDAFYLAVFWMLNTIGWVTFYW---HWKHITLWQGNVSQFNESSTYLMGWLRDYLWL 626
I+ W ++ V W L I W YW HW+ + W L+ W + WL
Sbjct: 337 INWWFVYWWLVHWRL--INWWFVYWWLVHWRLVHWW-------------LLNWWLVHWWL 203
Query: 627 NSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISW 667
+ LING+ V W F +G + W +L++W
Sbjct: 202 VNWWLINGW--------FVHWWFFNWGLINWRCVNRWLVNW 104
>TC214149 homologue to UP|PRP1_SOYBN (P08012) Repetitive proline-rich cell
wall protein 1 precursor, partial (96%)
Length = 942
Score = 37.4 bits (85), Expect = 0.024
Identities = 29/104 (27%), Positives = 44/104 (41%), Gaps = 6/104 (5%)
Frame = -1
Query: 570 ISAWDAFYLAVFWMLNTIGWVTFYW---HWKHITLWQGNVSQFNESSTYLMGWLRDYLWL 626
I+ W ++ V W L I W YW HW+ + W L+ W + WL
Sbjct: 609 INWWFVYWWLVHWRL--INWWFVYWWLVHWRLVHWW-------------LLNWWLVHWWL 475
Query: 627 NSSQLINGYNPFGMNSLSVWA--WMFLFGHLV-WATGFMFLISW 667
+ LING+ + L+ W W + G LV W +L++W
Sbjct: 474 VNWWLINGWFVYWW-LLNWWLVHWWLVNGWLVNWRLVHWWLVNW 346
Score = 36.2 bits (82), Expect = 0.054
Identities = 23/95 (24%), Positives = 40/95 (41%), Gaps = 2/95 (2%)
Frame = -1
Query: 576 FYLAVFWMLNTIGWVTFYW--HWKHITLWQGNVSQFNESSTYLMGWLRDYLWLNSSQLIN 633
++L +W++N GW ++W +W + W N +L+ W + WL + LIN
Sbjct: 483 WWLVNWWLIN--GWFVYWWLLNWWLVHWWLVN--------GWLVNWRLVHWWLVNWWLIN 334
Query: 634 GYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWR 668
G W F +G + W + +WR
Sbjct: 333 G-------------WFFNWGFIHWRLVNRWFFNWR 268
>TC214098 UP|PRP1_SOYBN (P08012) Repetitive proline-rich cell wall protein 1
precursor, complete
Length = 1144
Score = 37.0 bits (84), Expect = 0.031
Identities = 24/95 (25%), Positives = 41/95 (42%), Gaps = 2/95 (2%)
Frame = -2
Query: 576 FYLAVFWMLNTIGWVTFYW--HWKHITLWQGNVSQFNESSTYLMGWLRDYLWLNSSQLIN 633
++L +W++N GW ++W +W + W N +L+ W + WL + LIN
Sbjct: 507 WWLVNWWLIN--GWFVYWWLLNWWLVHWWLVN--------WWLIHWRLVHWWLVNWWLIN 358
Query: 634 GYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWR 668
G W F +G + W + I+WR
Sbjct: 357 G-------------WFFNWGFIHWRLVNRWFINWR 292
Score = 32.0 bits (71), Expect = 1.0
Identities = 24/102 (23%), Positives = 41/102 (39%), Gaps = 12/102 (11%)
Frame = -2
Query: 578 LAVFWMLNT-------IGWVTFYW---HWKHITLWQGNVSQFNESSTYLMGWLRDYLWLN 627
L +W++N I W YW HW+ + W L+ W + WL
Sbjct: 636 LVNWWLVNRRFLNWRLINWWFVYWWLVHWRLVHWW-------------LLNWWLVHWWLV 496
Query: 628 SSQLINGYNPFG--MNSLSVWAWMFLFGHLVWATGFMFLISW 667
+ LING+ + +N V W+ + + W +L++W
Sbjct: 495 NWWLINGWFVYWWLLNWWLVHWWLVNWWLIHWRLVHWWLVNW 370
>TC215069 similar to UP|O24300 (O24300) PtxA protein precursor, partial (83%)
Length = 1405
Score = 33.1 bits (74), Expect = 0.45
Identities = 27/107 (25%), Positives = 39/107 (36%), Gaps = 11/107 (10%)
Frame = -1
Query: 582 WMLNTIGWVTFYWHWKHITLWQG---NVSQFNESSTYLMGWLR------DYLWLNSSQLI 632
W L IGW +W W +W V ++ ++ W R D W+
Sbjct: 388 WWLRDIGWFNNWWFWDIRWMWMQWRLRVRFWDMGWMWMNWWFRVVRRFWDIGWV------ 227
Query: 633 NGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWR--GYWQELIET 677
Y + L W W +LF + + G +F WR G W I T
Sbjct: 226 --YMGWFWR*LGWWTWFWLFVNWWFHNGRLF---WRRLGRWASAIRT 101
>BE346777
Length = 461
Score = 32.7 bits (73), Expect = 0.59
Identities = 18/62 (29%), Positives = 26/62 (41%), Gaps = 3/62 (4%)
Frame = +3
Query: 227 TGQWNLYAQNPDSSSHLFGTSQGAGTAILTLLGGFHPQTQSLWLTDM---AHHHLAIAIL 283
T W L + QGA T L ++GG + + LW M HHHL + ++
Sbjct: 111 TSSWTFIPNGEIDGQELLASYQGALTMRLRIIGGLT*ERKRLWRKSMHQHQHHHLFLLVV 290
Query: 284 FL 285
L
Sbjct: 291 NL 296
>TC204285 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (32%)
Length = 1539
Score = 32.7 bits (73), Expect = 0.59
Identities = 26/109 (23%), Positives = 43/109 (38%)
Frame = -1
Query: 573 WDAFYLAVFWMLNTIGWVTFYWHWKHITLWQGNVSQFNESSTYLMGWLRDYLWLNSSQLI 632
W ++L FW + + F+W W+ + LW + +LR + W+ L
Sbjct: 576 WAWWFLRWFWWMRRLNLWWFWWMWR-LNLW------------WAWWFLRWFWWMRRLNLW 436
Query: 633 NGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETLAWA 681
+ + +N WAW FL+ W L W +W + L WA
Sbjct: 435 WFWWMWRLNLW--WAWWFLW----WMWWMRRLNLWWFWWMRRL-NLWWA 310
>TC204047 hydroxyproline-rich glycoprotein
Length = 887
Score = 32.3 bits (72), Expect = 0.77
Identities = 30/147 (20%), Positives = 55/147 (37%), Gaps = 10/147 (6%)
Frame = -3
Query: 558 PCDGPGRGGTCDISAWDAFYLAVFWMLNTIGW---------VTFYWHWKHITLWQGNVSQ 608
PC G R + W ++ V W + W + +W W+ + W GN+ +
Sbjct: 528 PCGGVKRKCYLVVVRWRG*FVVVVWWRWRVDWGRRRLVLVRLLVWWWWRRVVNWGGNMGR 349
Query: 609 FNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLIS-W 667
+ W R ++++ G +G + W + + G L W L+
Sbjct: 348 VG-----VNWWWR*WIFV-------GLLWWGWRRVVHWVRVRV-GFLHWRRRRWVLVGLL 208
Query: 668 RGYWQELIETLAWAHERTPLANLIRWR 694
RG+W +++ W R L+ WR
Sbjct: 207 RGWWWRVVD---WVWVR---VGLLHWR 145
>TC204868 similar to UP|Q9LKY8 (Q9LKY8) Proline-rich protein, partial (91%)
Length = 865
Score = 31.6 bits (70), Expect = 1.3
Identities = 23/99 (23%), Positives = 40/99 (40%), Gaps = 4/99 (4%)
Frame = -3
Query: 589 WVTFYWHWKHITLWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGMNSLSVWAW 648
WV + W W + L L+ W+ +W S + + P+ VW W
Sbjct: 395 WVVWVWVWVWVVL-----------VLELVVWV---VWA-SEEKVEDNQPWVW---VVWVW 270
Query: 649 MFLFGHLVWATGFMFLISWRGYWQ----ELIETLAWAHE 683
+++ L W + L++W W+ EL+ + WA E
Sbjct: 269 VWVVWALEWVVLVLELVAWALEWEVLVLELVVWVVWASE 153
>AI441447 SP|P08012|PRP1 Repetitive proline-rich cell wall protein 1
precursor. [Soybean] {Glycine max}, partial (30%)
Length = 411
Score = 30.8 bits (68), Expect = 2.3
Identities = 24/90 (26%), Positives = 35/90 (38%), Gaps = 3/90 (3%)
Frame = -1
Query: 582 WMLNTIGWVTFYW---HWKHITLWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPF 638
W L I W YW HW+ + W L+ W + WL + LING+
Sbjct: 213 WRL--INWWFVYWWLVHWRLVHWW-------------LLNWWLVHWWLVNWWLINGW--- 88
Query: 639 GMNSLSVWAWMFLFGHLVWATGFMFLISWR 668
V+ W+ + + W +LI WR
Sbjct: 87 -----FVYWWLLNWWLVHWWLVNWWLIHWR 13
>TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor, partial
(49%)
Length = 571
Score = 30.0 bits (66), Expect = 3.8
Identities = 15/47 (31%), Positives = 22/47 (45%)
Frame = -3
Query: 632 INGYNPFGMNSLSVWAWMFLFGHLVWATGFMFLISWRGYWQELIETL 678
+ G N G+ L +AW FL L W G+W+++IE L
Sbjct: 116 LKGVNN*GLRLLPCFAWFFL------------LFFWGGFWKKIIEAL 12
>TC214274 homologue to UP|Q00508 (Q00508) Ribosomal protein L41, complete
Length = 737
Score = 29.6 bits (65), Expect = 5.0
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = -2
Query: 582 WMLNTIGWVTFYWHWKHITLW 602
W LNTI WV F + WK +W
Sbjct: 292 WSLNTIFWVVFAFWWKTGLVW 230
>TC204059 similar to UP|Q94ES3 (Q94ES3) Root nodule extensin (Fragment),
partial (96%)
Length = 853
Score = 29.6 bits (65), Expect = 5.0
Identities = 27/144 (18%), Positives = 50/144 (33%), Gaps = 30/144 (20%)
Frame = -1
Query: 558 PCDGPG-RGGTCDISAWDAFYLAVFWMLNTIGW---------VTFYWHWKHITLWQGNVS 607
PC G +G + W ++ V W + W + +W W+ + W GN+
Sbjct: 553 PCGGVI*KGYILVVVGWRG*FVVVVWWRWRVDWGRRRLVLIRLLVWWWWRRVVDWGGNMG 374
Query: 608 QFNESS----TYLMGWLRDYL---------------WLNSSQLINGYNPFGMNSLSVWAW 648
+ + +G LR + W ++ G +G + W W
Sbjct: 373 RVGVNWWRR*WVFVGLLRCWWWGVVHWVWVRVGFLHWRRRRWVLVGLLRWGWWRVVDWVW 194
Query: 649 MFL-FGHLVWATGFMFLISWRGYW 671
+++ F H W G + W +W
Sbjct: 193 VWVGFLHWRWG*GVFVGLLWWWWW 122
>TC221433 weakly similar to UP|Q7EYQ4 (Q7EYQ4) DnaJ protein family-like
protein, partial (34%)
Length = 588
Score = 29.6 bits (65), Expect = 5.0
Identities = 10/26 (38%), Positives = 19/26 (72%)
Frame = +2
Query: 396 RDYNPEQNEDNVLARMLDHKEAIISH 421
++Y+P+QN+DN++A KE + S+
Sbjct: 206 KEYHPDQNQDNIVAAEAKFKEVLTSY 283
>BE805198 similar to PIR|T06455|T064 Myb26 protein - garden pea, partial
(24%)
Length = 256
Score = 29.6 bits (65), Expect = 5.0
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +2
Query: 437 YVHNDVMLAFGTPEKQILIEPIFAQW 462
Y+H+DV TPE+Q+LI ++A W
Sbjct: 80 YLHSDVRKGNITPEEQLLIMDLYAVW 157
>TC218079 similar to UP|Q945B9 (Q945B9) Growth-on protein GRO11, partial
(33%)
Length = 1909
Score = 29.3 bits (64), Expect = 6.5
Identities = 20/75 (26%), Positives = 31/75 (40%), Gaps = 8/75 (10%)
Frame = +1
Query: 586 TIGWVTFYWHWKHITLWQGNVSQF--------NESSTYLMGWLRDYLWLNSSQLINGYNP 637
++GW+ +W W H L + + S F E S +++ D LW S I G+
Sbjct: 208 SLGWLQHHWFWGH-HLTRASSSIFTKV*LVGKEEPSRWVLLVGNDRLWFRSPYHIRGF-- 378
Query: 638 FGMNSLSVWAWMFLF 652
L+ W W F
Sbjct: 379 ----ELNGWTWSTSF 411
>BI497725 homologue to SP|P13993|PRP2 Repetitive proline-rich cell wall
protein 2 precursor. [Soybean] {Glycine max}, partial
(47%)
Length = 413
Score = 29.3 bits (64), Expect = 6.5
Identities = 19/77 (24%), Positives = 31/77 (39%)
Frame = -3
Query: 581 FWMLNTIGWVTFYWHWKHITLWQGNVSQFNESSTYLMGWLRDYLWLNSSQLINGYNPFGM 640
+W++N W YW H L + + + +L+ W + WL + LING
Sbjct: 189 WWLVN---WRLLYWWLVHWRLLNWGLVYWWLVNGWLVNWRLVHWWLVNWWLING------ 37
Query: 641 NSLSVWAWMFLFGHLVW 657
W F +G + W
Sbjct: 36 -------WFFNWGFIHW 7
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.140 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,113,058
Number of Sequences: 63676
Number of extensions: 744533
Number of successful extensions: 4877
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 4780
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4871
length of query: 734
length of database: 12,639,632
effective HSP length: 104
effective length of query: 630
effective length of database: 6,017,328
effective search space: 3790916640
effective search space used: 3790916640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Description of BAB33187.1

