

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0627.15
(557 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
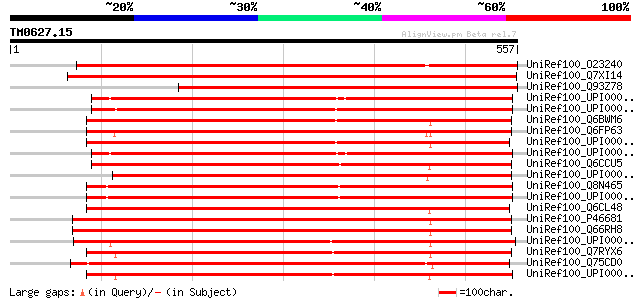
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O23240 Actin interacting protein [Arabidopsis thaliana] 750 0.0
UniRef100_Q7XI14 Putative actin interacting protein [Oryza sativa] 711 0.0
UniRef100_Q93Z78 At4g36400/C7A10_960 [Arabidopsis thaliana] 598 e-169
UniRef100_UPI0000360E67 UPI0000360E67 UniRef100 entry 530 e-149
UniRef100_UPI00003AED50 UPI00003AED50 UniRef100 entry 530 e-149
UniRef100_Q6BWM6 Debaryomyces hansenii chromosome B of strain CB... 526 e-148
UniRef100_Q6FP63 Candida glabrata strain CBS138 chromosome J com... 526 e-148
UniRef100_UPI000004DF7D UPI000004DF7D UniRef100 entry 525 e-147
UniRef100_UPI00003380C5 UPI00003380C5 UniRef100 entry 523 e-147
UniRef100_Q6CCU5 Similar to sp|P46681 Saccharomyces cerevisiae A... 519 e-145
UniRef100_UPI00003C14D1 UPI00003C14D1 UniRef100 entry 518 e-145
UniRef100_Q8N465 Hypothetical protein MGC25181 [Homo sapiens] 513 e-144
UniRef100_UPI0000456F51 UPI0000456F51 UniRef100 entry 512 e-143
UniRef100_Q6CL48 Kluyveromyces lactis strain NRRL Y-1140 chromos... 511 e-143
UniRef100_P46681 D-lactate dehydrogenase [cytochrome] 2, mitocho... 511 e-143
UniRef100_Q66RH8 YDL178W [Saccharomyces cerevisiae] 511 e-143
UniRef100_UPI000023F053 UPI000023F053 UniRef100 entry 509 e-142
UniRef100_Q7RYX6 Hypothetical protein [Neurospora crassa] 506 e-142
UniRef100_Q75CD0 ACL021Cp [Ashbya gossypii] 506 e-142
UniRef100_UPI0000236145 UPI0000236145 UniRef100 entry 505 e-141
>UniRef100_O23240 Actin interacting protein [Arabidopsis thaliana]
Length = 524
Score = 750 bits (1936), Expect = 0.0
Identities = 367/485 (75%), Positives = 428/485 (87%), Gaps = 4/485 (0%)
Query: 74 QHKCY-ASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGS 132
Q+KC+ +S L+QRNP FS L+ DV YF+EILG+KNVV+D+E+L T NTDWMHKY+GS
Sbjct: 42 QYKCFGSSAASLIQRNPLFSSLDSKDVSYFKEILGEKNVVEDKERLETANTDWMHKYKGS 101
Query: 133 SKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISF 192
SKL+L P NT +VSQIL+YC+SR LAVVPQGGNTGLVGGSVPVFDEVIV++ MNK++SF
Sbjct: 102 SKLMLLPKNTQEVSQILEYCDSRRLAVVPQGGNTGLVGGSVPVFDEVIVNVGLMNKILSF 161
Query: 193 DKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSL 252
D+VSG+LVCEAGCILEN+ +FLD +GFIMPLDLGAKGSC IGGNVSTNAGGLRL+RYGSL
Sbjct: 162 DEVSGVLVCEAGCILENLATFLDTKGFIMPLDLGAKGSCHIGGNVSTNAGGLRLIRYGSL 221
Query: 253 HGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSV 312
HG+VLG+EAV A+G VLDML TLRKDNTGYDLKHLFIGSEGSLGIVTKV+ILT PKLSSV
Sbjct: 222 HGTVLGLEAVTANGNVLDMLGTLRKDNTGYDLKHLFIGSEGSLGIVTKVSILTQPKLSSV 281
Query: 313 NVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFY 372
N+A +ACKDY CQKLL EAKR LGEILSAFEFLD+ SM+LV+NHL+G RNP+ + +FY
Sbjct: 282 NLAFIACKDYLSCQKLLVEAKRNLGEILSAFEFLDNNSMDLVLNHLDGVRNPVSSSENFY 341
Query: 373 VLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVG 432
+LIETTGSDE++D++KLEAFLL S+E L+SDGV+AQDINQASSFWR+REGITEAL + G
Sbjct: 342 ILIETTGSDETNDREKLEAFLLKSLEKGLVSDGVIAQDINQASSFWRIREGITEALQKAG 401
Query: 433 AVYKYDLSIPLENLYNLVEEMRSRLGNAANVVGYGHLGDGNLHLNISVPQYDDKILSQIE 492
AVYKYDLS+P+E +YN+V ++R RL ANV+GYGHLGDGNLHLNIS +Y+DK+L IE
Sbjct: 402 AVYKYDLSLPVEEIYNIVNDLRGRL---ANVMGYGHLGDGNLHLNISAAEYNDKLLGLIE 458
Query: 493 PFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
P+VYEWTSKH GSISAEHGLG+MKANEI YSKS ETV LMASIK LLDP ILNPYKVLP
Sbjct: 459 PYVYEWTSKHRGSISAEHGLGVMKANEIFYSKSPETVALMASIKKLLDPKGILNPYKVLP 518
Query: 553 HSLIA 557
HSL +
Sbjct: 519 HSLFS 523
>UniRef100_Q7XI14 Putative actin interacting protein [Oryza sativa]
Length = 559
Score = 711 bits (1836), Expect = 0.0
Identities = 352/494 (71%), Positives = 414/494 (83%), Gaps = 1/494 (0%)
Query: 64 QMGVGISTGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLSTVNT 123
Q ++ +Q + ++S VQRNP +S LN DDV YF+ ILG VVQDE+++S N
Sbjct: 66 QQSARMNFEVQKRSFSSAAAHVQRNPAYSVLNSDDVSYFKSILGDSGVVQDEDRVSVANM 125
Query: 124 DWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSL 183
DWM KY+GSS+LLL P +T +VS+IL YCNSR LAVVPQGGNTGLVGGSVPV+DEVI+SL
Sbjct: 126 DWMGKYKGSSQLLLLPKSTAEVSKILSYCNSRRLAVVPQGGNTGLVGGSVPVYDEVIISL 185
Query: 184 SSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGG 243
M+K+I+FD V+GIL CEAGC+LEN+ S+++N+GFIMPLDLGAKGSC IGGN+STNAGG
Sbjct: 186 GGMDKIITFDNVNGILTCEAGCVLENLSSYVENKGFIMPLDLGAKGSCHIGGNISTNAGG 245
Query: 244 LRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAI 303
LR +RYGSLHGSVLG+E VLADGTVLDML TLRKDNTGYDLKHLFIGSEGSLGIVTK+AI
Sbjct: 246 LRFIRYGSLHGSVLGLEVVLADGTVLDMLTTLRKDNTGYDLKHLFIGSEGSLGIVTKIAI 305
Query: 304 LTPPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARN 363
LTP KL S NVA L+C DY CQKLL A+R LGEILSAFEF+D +NL + +LEG N
Sbjct: 306 LTPAKLPSTNVAFLSCNDYISCQKLLLAARRSLGEILSAFEFMDRHCINLAMKYLEGVHN 365
Query: 364 PLP-TLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLRE 422
PLP + +FYVLIETTGSDES DK KLEAFLL SME+ L++DGV+AQDI+QAS+FWR+RE
Sbjct: 366 PLPVSPFNFYVLIETTGSDESYDKAKLEAFLLRSMEDGLVADGVIAQDISQASNFWRIRE 425
Query: 423 GITEALMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNAANVVGYGHLGDGNLHLNISVPQ 482
GI+EA ++VGAVYKYDLSIP+E LY++VEEMRSR+G+ V+GYGHLGDGNLHLNI +
Sbjct: 426 GISEASVKVGAVYKYDLSIPVEKLYDIVEEMRSRVGDMGQVLGYGHLGDGNLHLNILSTK 485
Query: 483 YDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPN 542
Y DK+L+QIEPFVYEWTSK GSISAEHGLGLMKA +I YSKS E VQLMASIK LLDPN
Sbjct: 486 YSDKMLAQIEPFVYEWTSKQRGSISAEHGLGLMKAEKIHYSKSSEAVQLMASIKKLLDPN 545
Query: 543 HILNPYKVLPHSLI 556
ILNPYKVLP S++
Sbjct: 546 SILNPYKVLPQSVL 559
>UniRef100_Q93Z78 At4g36400/C7A10_960 [Arabidopsis thaliana]
Length = 373
Score = 598 bits (1541), Expect = e-169
Identities = 289/372 (77%), Positives = 334/372 (89%)
Query: 186 MNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLR 245
MNK++SFD+VSG+LVCEAGCILEN+ +FLD +GFIMPLDLGAKGSC IGGNVSTNAGGLR
Sbjct: 1 MNKILSFDEVSGVLVCEAGCILENLATFLDTKGFIMPLDLGAKGSCHIGGNVSTNAGGLR 60
Query: 246 LVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILT 305
L+RYGSLHG+VLG+EAV A+G VLDML TLRKDNTGYDLKHLFIGSEGSLGIVTKV+ILT
Sbjct: 61 LIRYGSLHGTVLGLEAVTANGNVLDMLGTLRKDNTGYDLKHLFIGSEGSLGIVTKVSILT 120
Query: 306 PPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPL 365
PKLSSVN+A +ACKDY CQKLL EAKR LGEILSAFEFLD+ SM+LV+NHL+G RNP+
Sbjct: 121 QPKLSSVNLAFIACKDYLSCQKLLVEAKRNLGEILSAFEFLDNNSMDLVLNHLDGVRNPV 180
Query: 366 PTLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGIT 425
+ +FY+LIETTGSDE++D++KLEAFLL S+E L+SDGV+AQDINQASSFWR+REGIT
Sbjct: 181 SSSENFYILIETTGSDETNDREKLEAFLLKSLEKGLVSDGVIAQDINQASSFWRIREGIT 240
Query: 426 EALMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNAANVVGYGHLGDGNLHLNISVPQYDD 485
EAL + GAVYKYDLS+P+E +YN+V ++R RLG+ ANV+GYGHLGDGNLHLNIS +Y+D
Sbjct: 241 EALQKAGAVYKYDLSLPVEEIYNIVNDLRGRLGDLANVMGYGHLGDGNLHLNISAAEYND 300
Query: 486 KILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHIL 545
K+L IEP+VYEWTSKH GSISAEHGLG+MKANEI YSKS ETV LMASIK LLDP IL
Sbjct: 301 KLLGLIEPYVYEWTSKHRGSISAEHGLGVMKANEIFYSKSPETVALMASIKKLLDPKGIL 360
Query: 546 NPYKVLPHSLIA 557
NPYKVLPHSL +
Sbjct: 361 NPYKVLPHSLFS 372
>UniRef100_UPI0000360E67 UPI0000360E67 UniRef100 entry
Length = 480
Score = 530 bits (1366), Expect = e-149
Identities = 269/463 (58%), Positives = 343/463 (73%), Gaps = 4/463 (0%)
Query: 91 FSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILK 150
FS + ++D+ +F+ IL + VV D + L + N DW+ GSS++LL+P + +VSQIL+
Sbjct: 19 FSRITEEDLAFFRNILPGR-VVTDPDLLESNNEDWLKSVRGSSEVLLRPQTSQEVSQILR 77
Query: 151 YCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENI 210
YCNSR+LAV PQGGNTGLVGGSVPV+DE+I+S + MN + SFD +SGIL C+AGC+LE++
Sbjct: 78 YCNSRNLAVNPQGGNTGLVGGSVPVYDEIILSTALMNHIRSFDSISGILTCQAGCVLEDL 137
Query: 211 ISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLD 270
+L+ IMPLDLGAKGSC IGGNV+TNAGGLRL+RYGSLHG+VLG+E VLADG VLD
Sbjct: 138 SLYLEQRDHIMPLDLGAKGSCHIGGNVATNAGGLRLLRYGSLHGTVLGLEVVLADGQVLD 197
Query: 271 MLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQ 330
L TLRKDNTGYDLK LFIGSEG+LG++T +IL P K SVNVA L C+ + K Q
Sbjct: 198 CLATLRKDNTGYDLKQLFIGSEGTLGVITAASILCPRKPKSVNVAFLGCETFEQLLKTFQ 257
Query: 331 EAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDESSDKQKLE 390
+ LGEILSA+EFLDS+ M L+ HL+ NP+ FY++IET+GS+ + D++KL
Sbjct: 258 LCRGMLGEILSAYEFLDSECMRLLNTHLK-LTNPVRDC-QFYIVIETSGSEPTHDEEKLH 315
Query: 391 AFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIPLENLYNLV 450
FL +M + L+S+G +A + ++ + W LRE +TEAL G YKYD+S+P+E LY LV
Sbjct: 316 NFLEEAMTSSLVSEGTVATEDSKIKALWSLRERVTEALAHDGYTYKYDVSLPVEELYQLV 375
Query: 451 EEMRSRL-GNAANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAE 509
+MR R+ G A +VVGYGH+GDGNLHLNI+ D +L+ IEPFVYEWT+KH GSISAE
Sbjct: 376 TDMRERVGGRAKSVVGYGHVGDGNLHLNITSQAKDPALLADIEPFVYEWTAKHRGSISAE 435
Query: 510 HGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
HGLGL K N I YSK + V LM IK LLDP ILNPYK LP
Sbjct: 436 HGLGLKKRNYIYYSKPSQAVALMGQIKALLDPKGILNPYKTLP 478
>UniRef100_UPI00003AED50 UPI00003AED50 UniRef100 entry
Length = 486
Score = 530 bits (1365), Expect = e-149
Identities = 273/464 (58%), Positives = 344/464 (73%), Gaps = 5/464 (1%)
Query: 91 FSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILK 150
FS L+ DV +F+ +L + EE L N DW+ G S+LLL+P +V+Q+L+
Sbjct: 26 FSRLSRGDVAFFEGLLPGRACTNPEE-LKACNVDWLKSVRGCSELLLKPKTAAEVAQVLR 84
Query: 151 YCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENI 210
YC+ R+LAV PQGGNTGLVGGSVPVFDE+I+S + MN++ISFD VSGILVC+AGC+LE +
Sbjct: 85 YCHERNLAVNPQGGNTGLVGGSVPVFDEIILSTALMNQIISFDPVSGILVCQAGCVLEQL 144
Query: 211 ISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLD 270
+L+ +GFIMPLDLGAKGSC IGGNV+TNAGGLRL+RYGSL G+VLG+E VLADGTVLD
Sbjct: 145 NEYLEEQGFIMPLDLGAKGSCHIGGNVATNAGGLRLLRYGSLRGTVLGLEVVLADGTVLD 204
Query: 271 MLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQ 330
L +LRKDNTGYDLK LFIGSEG+LG++T V+IL P K +VN+A L C+D++ Q+
Sbjct: 205 CLASLRKDNTGYDLKQLFIGSEGTLGVITAVSILCPQKPKAVNLAFLGCQDFSRVQETFT 264
Query: 331 EAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDESSDKQKLE 390
+ LGEILSA+EF+D + M LV HL G NP+ FYVLIET+GS+ + D++KL
Sbjct: 265 TCRTMLGEILSAYEFMDEKCMELVEKHL-GLSNPVRG-SPFYVLIETSGSNSTHDEEKLN 322
Query: 391 AFLLHSMENELISDGVLAQDINQ-ASSFWRLREGITEALMRVGAVYKYDLSIPLENLYNL 449
+FL +M + L++DG +A D + ++ W LRE ITEAL R G VYKYD+S+P+ LY+L
Sbjct: 323 SFLEQAMTSGLVTDGTVAVDDKKIKANLWSLRERITEALTRDGCVYKYDVSLPVGKLYDL 382
Query: 450 VEEMRSRLGNAA-NVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISA 508
V +MR+RLG +A NVVGYGHLGDGNLHLNI+ Y +L IEPFVYEWT++ NGSISA
Sbjct: 383 VTDMRARLGQSAKNVVGYGHLGDGNLHLNITAESYSHSLLDAIEPFVYEWTARCNGSISA 442
Query: 509 EHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
EHGLG K I YSK E V LM K +LDP ILNPYK LP
Sbjct: 443 EHGLGFKKKQFIQYSKPNEAVFLMQRFKAMLDPKGILNPYKTLP 486
>UniRef100_Q6BWM6 Debaryomyces hansenii chromosome B of strain CBS767 of Debaryomyces
hansenii [Debaryomyces hansenii]
Length = 536
Score = 526 bits (1355), Expect = e-148
Identities = 266/481 (55%), Positives = 342/481 (70%), Gaps = 15/481 (3%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKN-VVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTD 143
V+R+ RF +L D D+ +F +L + V+ EE + N DWM KY+G SKL+L+P T+
Sbjct: 57 VKRDDRFKQLEDSDIEFFHSVLQDPHGVITGEEDIEFYNEDWMRKYKGQSKLVLRPKTTE 116
Query: 144 QVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEA 203
Q+SQI+KYCN + LAVVPQGGNTGLVGGS+PVFDE+I+SL+ MNK+ SFD VSGIL C+A
Sbjct: 117 QISQIVKYCNEQKLAVVPQGGNTGLVGGSIPVFDEIIISLAGMNKIRSFDSVSGILKCDA 176
Query: 204 GCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVL 263
G ILEN + L EG+I PLDLGAKGSC +GGNV+TNAGGLRL++YGSLHGSVLG+EAVL
Sbjct: 177 GLILENADNALSEEGYIFPLDLGAKGSCHVGGNVATNAGGLRLLKYGSLHGSVLGLEAVL 236
Query: 264 ADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYN 323
DGT+ + + +LRKDNTGYDLK LFIGSEG+LG++T V+IL P + VN+A LA Y
Sbjct: 237 PDGTIYNSMNSLRKDNTGYDLKQLFIGSEGTLGLITGVSILCPSRPQVVNIAFLAVSSYE 296
Query: 324 CCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPT-LHDFYVLIETTGSDE 382
QK+ A+R+L EILSAFEF+D +S L HL A +P+ + + FYVLIET+GS++
Sbjct: 297 AVQKVFVGARRELSEILSAFEFMDDKSQLLTARHL-NADHPIESGEYPFYVLIETSGSNK 355
Query: 383 SSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIP 442
D +KLE FL +ME+ L+ DG++AQD Q S W RE I EA G VYKYD+S+P
Sbjct: 356 EHDDEKLEGFLERAMEDGLVDDGIIAQDETQLQSLWSWRESIPEASTIGGGVYKYDVSLP 415
Query: 443 LENLYNLVEEMRSRLGNA------------ANVVGYGHLGDGNLHLNISVPQYDDKILSQ 490
L +LY LV+ ++L A +GYGH+GDGNLHLN++V +Y +I S
Sbjct: 416 LSDLYGLVDAASAKLKEANLVGIDDDSKPVVEAIGYGHIGDGNLHLNVAVREYSKEIESV 475
Query: 491 IEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKV 550
+EPFVYEW NGSISAEHGLG K N I YSK+ ++LM +K DPN I+NPYK
Sbjct: 476 LEPFVYEWIQSKNGSISAEHGLGFQKKNYIGYSKNDIEIRLMKDLKKHYDPNGIMNPYKY 535
Query: 551 L 551
+
Sbjct: 536 I 536
>UniRef100_Q6FP63 Candida glabrata strain CBS138 chromosome J complete sequence
[Candida glabrata]
Length = 543
Score = 526 bits (1354), Expect = e-148
Identities = 258/480 (53%), Positives = 352/480 (72%), Gaps = 13/480 (2%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQ---DEEKLSTVNTDWMHKYEGSSKLLLQPWN 141
V+R+ RF L DD+ YF+ IL + +++ + L+ N DWM KY+G SKLLL+P
Sbjct: 64 VKRDGRFKSLAKDDIDYFKSILSETELLEGNDSNDSLAAYNEDWMRKYKGQSKLLLKPKT 123
Query: 142 TDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVC 201
+QVS+I+KYCN LAVVPQGGNTGLVGGS+PVFDE+I+SL+++NK+ FD VSGI C
Sbjct: 124 VEQVSKIIKYCNDNRLAVVPQGGNTGLVGGSIPVFDEIILSLANLNKIREFDPVSGIFKC 183
Query: 202 EAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEA 261
+AG ILE +L+ G+I PLDLGAKGSC +GG V+TNAGGLRL+RYGSLHGSVLG+E
Sbjct: 184 DAGVILEAANEYLEKNGYIFPLDLGAKGSCHVGGVVATNAGGLRLLRYGSLHGSVLGLEV 243
Query: 262 VLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKD 321
VL +G +++ + +LRKDNTGYD+K LFIGSEG++GI+T V+IL PPK NV L+ +D
Sbjct: 244 VLPNGKIVNSMHSLRKDNTGYDIKQLFIGSEGTIGIITGVSILAPPKPKFTNVCFLSLED 303
Query: 322 YNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSD 381
Y QK+ +A+++L EI+SA+EF+D++S++L +HLEG PL H FYVLIET+GS+
Sbjct: 304 YESVQKVFVKARQELSEIISAYEFMDNKSLSLTKDHLEGVPFPLEDEHPFYVLIETSGSN 363
Query: 382 ESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSI 441
+ D KLEAFL +ME L++DGV+AQD + ++ W+ RE I E+ G VYKYD+S+
Sbjct: 364 KEHDDAKLEAFLELAMEEGLVTDGVVAQDETELANLWQWREMIPESSQSNGGVYKYDVSL 423
Query: 442 PLENLYNLVEEMRSR------LGNA----ANVVGYGHLGDGNLHLNISVPQYDDKILSQI 491
PL+++Y+LVE + R LG A + +GYGH+GDGNLHLN++V +Y ++ +
Sbjct: 424 PLKSMYSLVEAVNDRLTQHGLLGEAPKPVVSAIGYGHIGDGNLHLNVAVREYTKEVEKCL 483
Query: 492 EPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVL 551
EPFVYE+ +H+GS+SAEHGLG K N I YSK+ E ++++ SIKN DPN ILNPYK +
Sbjct: 484 EPFVYEFIQRHHGSVSAEHGLGFQKKNYIQYSKTPEEIEMIKSIKNQYDPNSILNPYKYI 543
>UniRef100_UPI000004DF7D UPI000004DF7D UniRef100 entry
Length = 527
Score = 525 bits (1352), Expect = e-147
Identities = 268/478 (56%), Positives = 342/478 (71%), Gaps = 14/478 (2%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQ 144
VQR+ +F +L D+ YF+ +L + +++ DE+ L N DWM KY G S+L+L+P T+Q
Sbjct: 49 VQRDAKFKQLESQDIEYFKSVLPENSIITDEDDLLFFNEDWMRKYRGQSQLVLKPKTTEQ 108
Query: 145 VSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAG 204
V+ ILKYCN LAVVPQGGNTGLVGGS P+FDE+I+SLS+MNK+ SFD VSGIL +AG
Sbjct: 109 VASILKYCNDNKLAVVPQGGNTGLVGGSNPIFDEIIISLSAMNKIRSFDPVSGILKVDAG 168
Query: 205 CILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLA 264
ILE +L +G+I PLDLGAKGSC +GGNV+ NAGGLRL+RYGSLHGSVLG+EAVL
Sbjct: 169 VILETADQYLAEQGYIFPLDLGAKGSCHVGGNVACNAGGLRLLRYGSLHGSVLGLEAVLP 228
Query: 265 DGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNC 324
DGTV + + +LRKDNTGYDLK LFIGSEG+LGI+T V+IL P + + NVA LA Y
Sbjct: 229 DGTVYNSMHSLRKDNTGYDLKQLFIGSEGTLGIITGVSILCPSRPQAQNVAFLAVSSYEA 288
Query: 325 CQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPT-LHDFYVLIETTGSDES 383
QK+ +A+++L EILSAFEF+D+ S L HL G +P+ + FYVLIET+GS++
Sbjct: 289 VQKVFVQARKELQEILSAFEFMDNTSQKLTAKHL-GLEHPIESGDFPFYVLIETSGSNKE 347
Query: 384 SDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIPL 443
D +KLE FL ++ME L+ DG++AQD Q S W RE I EA G VYKYD+SIPL
Sbjct: 348 HDDEKLETFLGNAMEEGLVDDGIIAQDEAQIQSLWSWRESIPEATTIGGGVYKYDVSIPL 407
Query: 444 ENLYNLVEEMRSRLGNAA------------NVVGYGHLGDGNLHLNISVPQYDDKILSQI 491
+LY LVE++ +RL +A +GYGH+GDGNLHLN+SV +Y +I + +
Sbjct: 408 ADLYGLVEDINTRLNDAGIASLDDESKLVLAALGYGHIGDGNLHLNVSVRKYSPEIETIL 467
Query: 492 EPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYK 549
EPFVYEW +K NGSISAEHGLG K N I YSK+ V+L+ IK DPN I+NPYK
Sbjct: 468 EPFVYEWIAKKNGSISAEHGLGFQKKNYIGYSKNEIEVKLIKEIKQHYDPNGIMNPYK 525
>UniRef100_UPI00003380C5 UPI00003380C5 UniRef100 entry
Length = 542
Score = 523 bits (1348), Expect = e-147
Identities = 266/463 (57%), Positives = 341/463 (73%), Gaps = 4/463 (0%)
Query: 91 FSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILK 150
FS + ++D+ +F+ +L + V D + L + N DW+ GSS+LLL+P +++VSQILK
Sbjct: 80 FSRVTEEDLAFFRSVLPGR-AVTDPDLLESSNQDWLKSVRGSSELLLRPRTSEEVSQILK 138
Query: 151 YCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENI 210
YCN R+LAV PQGGNTGLVGGSVPV DE+I+S + M + SFD VSGIL C+AGC+LE++
Sbjct: 139 YCNHRNLAVNPQGGNTGLVGGSVPVHDEIILSTALMKDIRSFDSVSGILTCQAGCVLEDL 198
Query: 211 ISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLD 270
+L++ IMPLDLGAKGSC IGGNV+TNAGGLRL+RYGSLHG+VLG+E VLADG VLD
Sbjct: 199 SLYLEDRDHIMPLDLGAKGSCHIGGNVATNAGGLRLLRYGSLHGTVLGLEVVLADGRVLD 258
Query: 271 MLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQ 330
L TLRKDNTGYDLK LFIGSEG+LG++T V+IL P K SV VA L C + K Q
Sbjct: 259 CLATLRKDNTGYDLKQLFIGSEGTLGVITAVSILCPRKPRSVKVAFLGCDTFEQLLKTFQ 318
Query: 331 EAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDESSDKQKLE 390
+ LGEILSA+EFLDS+ M L+ HL+ NP+ FY+++ET+GSD + D++KL
Sbjct: 319 LCRAMLGEILSAYEFLDSECMRLLNTHLQ-LSNPIRDCR-FYIVMETSGSDPTHDEEKLH 376
Query: 391 AFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIPLENLYNLV 450
+FL ++ + L+++G +A + ++ + W LRE ITEAL G YKYD+S+P+E LY LV
Sbjct: 377 SFLEAAVTSSLVTEGTVATEDSKIKALWALRERITEALTHHGYTYKYDVSLPVEQLYQLV 436
Query: 451 EEMRSRL-GNAANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAE 509
+MR L G A +VVGYGH+GDGNLHLNIS D +L+ IEPFVYEWT++H GS+SAE
Sbjct: 437 TDMRGHLQGRAKSVVGYGHVGDGNLHLNISSQAKDPALLAAIEPFVYEWTARHRGSVSAE 496
Query: 510 HGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
HGLGL K N I YSKS + V LM +K LLDP ILNPYK LP
Sbjct: 497 HGLGLKKRNYIYYSKSSQAVALMGQLKALLDPKGILNPYKTLP 539
>UniRef100_Q6CCU5 Similar to sp|P46681 Saccharomyces cerevisiae Actin interacting
protein 2 [Yarrowia lipolytica]
Length = 541
Score = 519 bits (1336), Expect = e-145
Identities = 261/473 (55%), Positives = 341/473 (71%), Gaps = 13/473 (2%)
Query: 91 FSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILK 150
++ L++ D+ +F+ IL + +V D E + NTDWM K+ G ++L+L+P + +QVS+IL
Sbjct: 70 YARLSEADIEHFKTILPENGLVTDVEDIEMFNTDWMRKFRGQTQLVLKPTSAEQVSKILS 129
Query: 151 YCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENI 210
YCN + LAVVPQGGNTGLVGGSVP++DE+I+SLS+MNKV SFD VSGILV +AG ILE
Sbjct: 130 YCNDKKLAVVPQGGNTGLVGGSVPLYDEIILSLSNMNKVRSFDNVSGILVADAGVILEAA 189
Query: 211 ISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLD 270
+L +GF+ PLDLGAKGSC IGGNV+TNAGGLRL+RYGSLHGSVLG+E VLA+G V++
Sbjct: 190 DMYLAEQGFMFPLDLGAKGSCHIGGNVATNAGGLRLLRYGSLHGSVLGLEVVLANGKVVN 249
Query: 271 MLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQ 330
ML LRKDNTGYDLK LFIGSEG+LG++T V+IL P + ++ NVA LA + Y Q+
Sbjct: 250 MLSKLRKDNTGYDLKQLFIGSEGTLGVITAVSILCPQRPAAKNVAYLAVESYEKVQQAFI 309
Query: 331 EAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDESSDKQKLE 390
AK+ LGEILSAFE +D +S V H + R P+ T F++LIET GS++ D KLE
Sbjct: 310 AAKKDLGEILSAFELMDGRSQGWVQQHCDLTR-PVDTESPFFILIETAGSNKEHDDAKLE 368
Query: 391 AFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIPLENLYNLV 450
AFL ME E++ DGV+A D Q + W RE ITE + + G VYKYD+SIPL LYN+V
Sbjct: 369 AFLEDVMEQEIVCDGVVASDETQFQNLWAWRERITEVIGKEGGVYKYDISIPLPQLYNIV 428
Query: 451 EEMRSRLGN------------AANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEW 498
+++R+ + +V+GYGH+GDGNLHLNI +Y ++I +EP++YEW
Sbjct: 429 DDLRAVFESKDMISKTDASKPVVDVIGYGHVGDGNLHLNIMTREYSERISQVLEPWIYEW 488
Query: 499 TSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVL 551
SK +GSISAEHGLGL+K + I YSK + V LM +KN DPNHILNPYK +
Sbjct: 489 VSKVHGSISAEHGLGLIKKHFIQYSKDEDNVYLMKQLKNAYDPNHILNPYKYI 541
>UniRef100_UPI00003C14D1 UPI00003C14D1 UniRef100 entry
Length = 574
Score = 518 bits (1333), Expect = e-145
Identities = 251/448 (56%), Positives = 330/448 (73%), Gaps = 10/448 (2%)
Query: 114 DEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSV 173
D +L T N DWM+KY G S+L+L+P T +VS+I+KYC+S ++AVVPQGGNTGLVGG V
Sbjct: 125 DPNELDTFNNDWMNKYHGKSRLVLKPKTTKEVSKIMKYCHSNNIAVVPQGGNTGLVGGGV 184
Query: 174 PVFDEVIVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQI 233
PVFDEV++ L +N++ SFD+V+G LVC+AGCILE++ +++ +G++MPLDLGAKGSC I
Sbjct: 185 PVFDEVVLQLGGLNQIRSFDEVAGTLVCDAGCILESLDNYIAEKGYMMPLDLGAKGSCHI 244
Query: 234 GGNVSTNAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEG 293
GGNV+TNAGGLR +RYGSLHG+VLG+E VL DGT++D L TLRKDNTG+DLK LFIGSEG
Sbjct: 245 GGNVATNAGGLRFLRYGSLHGNVLGLEVVLPDGTIMDNLSTLRKDNTGFDLKQLFIGSEG 304
Query: 294 SLGIVTKVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNL 353
++G++T V+I+TP + S+VNVA+ A Y Q + + K+ GEILSAFEF D S ++
Sbjct: 305 TIGVITGVSIITPRRPSAVNVAVFALDSYEAVQTVFGQVKKHCGEILSAFEFFDQTSFDV 364
Query: 354 VINHLEG-ARNPLPTLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDIN 412
V H R+P H FYVLIET+GS++ D +KL+ L H ME +I DGVLAQD
Sbjct: 365 VQKHATAHVRDPFEARHPFYVLIETSGSNKDHDDEKLQVLLEHLMEEGMIQDGVLAQDET 424
Query: 413 QASSFWRLREGITEALMRVGAVYKYDLSIPLENLYNLVEEMRSRL---------GNAANV 463
Q W LRE I EA ++G V+KYDLS+P++ +Y LV +MR R + V
Sbjct: 425 QLQGLWALRESIPEAAGKMGKVFKYDLSMPIDKMYKLVLDMRQRFEEHGIHGQDKDVGAV 484
Query: 464 VGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYS 523
GYGH+GDGNLH+NI YD+KI + IEP++YEWTS+ GS+SAEHGLGLMKA +I YS
Sbjct: 485 CGYGHIGDGNLHINIVAKSYDEKIENIIEPYIYEWTSQVKGSVSAEHGLGLMKATKIGYS 544
Query: 524 KSLETVQLMASIKNLLDPNHILNPYKVL 551
K+ E+++ M IK+L DP ILNPYK +
Sbjct: 545 KTPESIEYMKKIKHLFDPKGILNPYKYI 572
>UniRef100_Q8N465 Hypothetical protein MGC25181 [Homo sapiens]
Length = 521
Score = 513 bits (1322), Expect = e-144
Identities = 266/469 (56%), Positives = 340/469 (71%), Gaps = 5/469 (1%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQ 144
VQR P FS ++ D+ F+ I+ VV D E L N DW+ G SK+LL+P +++
Sbjct: 54 VQRLP-FSTVSKQDLAAFERIV-PGGVVTDPEALQAPNVDWLRTLRGCSKVLLRPRTSEE 111
Query: 145 VSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAG 204
VS IL++C+ R+LAV PQGGNTG+VGGSVPVFDE+I+S + MN+V+SF VSGILVC+AG
Sbjct: 112 VSHILRHCHERNLAVNPQGGNTGMVGGSVPVFDEIILSTARMNRVLSFHSVSGILVCQAG 171
Query: 205 CILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLA 264
C+LE + +++ FIMPLDLGAKGSC IGGNV+TNAGGLR +RYGSLHG+VLG+E VLA
Sbjct: 172 CVLEELSRYVEERDFIMPLDLGAKGSCHIGGNVATNAGGLRFLRYGSLHGTVLGLEVVLA 231
Query: 265 DGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNC 324
DGTVLD L +LRKDNTGYDLK LFIGSEG+LGI+T V+IL PPK +VNVA L C +
Sbjct: 232 DGTVLDCLTSLRKDNTGYDLKQLFIGSEGTLGIITTVSILCPPKPRAVNVAFLGCPGFAE 291
Query: 325 CQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDESS 384
+ K LGEILSAFEF+D+ M LV HL A +P+ FYVLIET+GS+
Sbjct: 292 VLQTFSTCKGMLGEILSAFEFMDAVCMQLVGRHLHLA-SPVQE-SPFYVLIETSGSNAGH 349
Query: 385 DKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIPLE 444
D +KL FL H++ + L++DG +A D + W LRE ITEAL R G VYKYDLS+P+E
Sbjct: 350 DAEKLGHFLEHALGSGLVTDGTMATDQRKVKMLWALRERITEALSRDGYVYKYDLSLPVE 409
Query: 445 NLYNLVEEMRSRLG-NAANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHN 503
LY++V ++R+RLG +A +VVGYGHLGDGNLHLN++ + +L+ +EP VYEWT+
Sbjct: 410 RLYDIVTDLRARLGPHAKHVVGYGHLGDGNLHLNVTAEAFSPSLLAALEPHVYEWTAGQQ 469
Query: 504 GSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
GS+SAEHG+G K + + YSK +QLM +K LLDP ILNPYK LP
Sbjct: 470 GSVSAEHGVGFRKRDVLGYSKPPGALQLMQQLKALLDPKGILNPYKTLP 518
>UniRef100_UPI0000456F51 UPI0000456F51 UniRef100 entry
Length = 521
Score = 512 bits (1318), Expect = e-143
Identities = 265/469 (56%), Positives = 340/469 (71%), Gaps = 5/469 (1%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQDEEKLSTVNTDWMHKYEGSSKLLLQPWNTDQ 144
V+R P FS ++ D+ F+ I+ VV D E L N DW+ G SK+LL+P +++
Sbjct: 54 VRRLP-FSTVSKQDLAAFERIV-PGGVVTDPEALQAPNVDWLRTLRGCSKVLLRPRTSEE 111
Query: 145 VSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCEAG 204
VS IL++C+ R+LAV PQGGNTG+VGGSVPVFDE+I+S + MN+V+SF VSGILVC+AG
Sbjct: 112 VSHILRHCHERNLAVNPQGGNTGMVGGSVPVFDEIILSTARMNRVLSFHSVSGILVCQAG 171
Query: 205 CILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAVLA 264
C+LE + +++ FIMPLDLGAKGSC IGGNV+TNAGGLR +RYGSLHG+VLG+E VLA
Sbjct: 172 CVLEELSRYVEERDFIMPLDLGAKGSCHIGGNVATNAGGLRFLRYGSLHGTVLGLEVVLA 231
Query: 265 DGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDYNC 324
DGTVLD L +LRKDNTGYDLK LFIGSEG+LGI+T V+IL PPK +VNVA L C +
Sbjct: 232 DGTVLDCLTSLRKDNTGYDLKQLFIGSEGTLGIITTVSILCPPKPRAVNVAFLGCPGFAE 291
Query: 325 CQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDESS 384
+ K LGEILSAFEF+D+ M LV HL A +P+ FYVLIET+GS+
Sbjct: 292 VLQTFSTCKGMLGEILSAFEFMDAVCMQLVGRHLHLA-SPVQE-SPFYVLIETSGSNAGH 349
Query: 385 DKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIPLE 444
D +KL FL H++ + L++DG +A D + W LRE ITEAL R G VYKYDLS+P+E
Sbjct: 350 DAEKLGHFLEHALGSGLVTDGTMATDQRKVKMLWALRERITEALSRDGYVYKYDLSLPVE 409
Query: 445 NLYNLVEEMRSRLG-NAANVVGYGHLGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHN 503
LY++V ++R+RLG +A +VVGYGHLGDGNLHLN++ + +L+ +EP VYEWT+
Sbjct: 410 RLYDIVTDLRARLGPHAKHVVGYGHLGDGNLHLNVTAEAFSPSLLAALEPHVYEWTAGQQ 469
Query: 504 GSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYKVLP 552
GS+SAEHG+G K + + YSK +QLM +K LLDP ILNPYK LP
Sbjct: 470 GSVSAEHGVGFRKRDVLGYSKPPGALQLMQQLKALLDPKGILNPYKTLP 518
>UniRef100_Q6CL48 Kluyveromyces lactis strain NRRL Y-1140 chromosome F of strain NRRL
Y- 1140 of Kluyveromyces lactis [Kluyveromyces lactis]
Length = 543
Score = 511 bits (1317), Expect = e-143
Identities = 257/479 (53%), Positives = 344/479 (71%), Gaps = 14/479 (2%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQ--DEEKLSTVNTDWMHKYEGSSKLLLQPWNT 142
++R F +L +D+ YF+ IL Q+ +++ + E L+ N DWM KY G SKL+L+P T
Sbjct: 63 IKRLESFKKLGKEDLDYFKTILSQQEILEANENEDLAFYNEDWMRKYRGQSKLVLRPKTT 122
Query: 143 DQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGILVCE 202
+QVS+ILKYCN ++LAVVPQGGNTGLVGGSVPVFDE+++SL+ +NK+ FD+VSGIL C+
Sbjct: 123 EQVSKILKYCNEQNLAVVPQGGNTGLVGGSVPVFDEIVLSLTQLNKIREFDEVSGILKCD 182
Query: 203 AGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLGVEAV 262
AG ILE FL G+I P+DLGAKGSC +GG V+TNAGGLRL+RYGSLHG+VLG+E V
Sbjct: 183 AGVILEAADMFLAERGYIFPMDLGAKGSCHVGGIVATNAGGLRLLRYGSLHGTVLGLEVV 242
Query: 263 LADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLACKDY 322
L +G +++ + LRKDNTGYDLK LFIGSEG++G+VT V+IL P + ++ NV+ L + Y
Sbjct: 243 LPNGEIVNSMNALRKDNTGYDLKQLFIGSEGTIGVVTGVSILCPIRPNAFNVSFLGVESY 302
Query: 323 NCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETTGSDE 382
QK+ +AK++LGEILSAFEF+D +S NL HL +PL H FY+LIET+GS +
Sbjct: 303 EAVQKVFVQAKKELGEILSAFEFMDLKSQNLTKIHLADMDHPLADDHPFYILIETSGSKK 362
Query: 383 SSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYDLSIP 442
D KLEAFL +ME+ L+ DGV+AQD + + W+ RE I E+ G VYKYD+S+P
Sbjct: 363 EHDDAKLEAFLESAMEDGLVVDGVVAQDETELKNLWQWREMIPESSQAGGGVYKYDVSLP 422
Query: 443 LENLYNLVEEMRSRLGN------------AANVVGYGHLGDGNLHLNISVPQYDDKILSQ 490
L++LY+LV+ +RL + VGYGH+GDGNLHLN++V +Y ++
Sbjct: 423 LKDLYSLVDAANARLEEHGLLSIDDESKPVISAVGYGHVGDGNLHLNVAVRRYTKEVEKC 482
Query: 491 IEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNPYK 549
+EPFVYE+ S NGSISAEHGLG K N I YSKS + ++++ +KNL D N ILNPYK
Sbjct: 483 LEPFVYEFVSSKNGSISAEHGLGFQKKNYIGYSKSEQEIKMIKQLKNLYDSNTILNPYK 541
>UniRef100_P46681 D-lactate dehydrogenase [cytochrome] 2, mitochondrial precursor
[Saccharomyces cerevisiae]
Length = 530
Score = 511 bits (1317), Expect = e-143
Identities = 257/494 (52%), Positives = 352/494 (71%), Gaps = 12/494 (2%)
Query: 70 STGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQ--DEEKLSTVNTDWMH 127
ST IQ + + V R+PRF +L DD+ YF+ IL ++ +++ + E LS N DWM
Sbjct: 37 STKIQTRLTSENYPDVHRDPRFKKLTSDDLNYFKSILSEQEILRASESEDLSFYNEDWMR 96
Query: 128 KYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMN 187
KY+G SKL+L+P + ++VS IL YCN +AVVPQGGNTGLVGGSVP+FDE+I+SL+++N
Sbjct: 97 KYKGQSKLVLRPKSVEKVSLILNYCNDEKIAVVPQGGNTGLVGGSVPIFDELILSLANLN 156
Query: 188 KVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLV 247
K+ FD VSGIL C+AG ILEN +++ + ++ PLDLGAKGSC +GG V+TNAGGLRL+
Sbjct: 157 KIRDFDPVSGILKCDAGVILENANNYVMEQNYMFPLDLGAKGSCHVGGVVATNAGGLRLL 216
Query: 248 RYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPP 307
RYGSLHGSVLG+E V+ +G +++ + ++RKDNTGYDLK LFIGSEG++GI+T V+ILT P
Sbjct: 217 RYGSLHGSVLGLEVVMPNGQIVNSMHSMRKDNTGYDLKQLFIGSEGTIGIITGVSILTVP 276
Query: 308 KLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPT 367
K + NV+ L+ + + QK+ A+++L EILSAFEF+D++S L + L+ A PL
Sbjct: 277 KPKAFNVSYLSVESFEDVQKVFVRARQELSEILSAFEFMDAKSQVLAKSQLKDAAFPLED 336
Query: 368 LHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEA 427
H FY+LIET+GS++ D KLE FL + ME +++DGV+AQD + + W+ RE I EA
Sbjct: 337 EHPFYILIETSGSNKDHDDSKLETFLENVMEEGIVTDGVVAQDETELQNLWKWREMIPEA 396
Query: 428 LMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNA----------ANVVGYGHLGDGNLHLN 477
G VYKYD+S+PL++LY+LVE +RL A +GYGH+GDGNLHLN
Sbjct: 397 SQANGGVYKYDVSLPLKDLYSLVEATNARLSEAELVGDSPKPVVGAIGYGHVGDGNLHLN 456
Query: 478 ISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKN 537
++V +Y+ I +EPFVYE+ S +GS+SAEHGLG K N I YSKS E V++M +K
Sbjct: 457 VAVREYNKNIEKTLEPFVYEFVSSKHGSVSAEHGLGFQKKNYIGYSKSPEEVKMMKDLKV 516
Query: 538 LLDPNHILNPYKVL 551
DPN ILNPYK +
Sbjct: 517 HYDPNGILNPYKYI 530
>UniRef100_Q66RH8 YDL178W [Saccharomyces cerevisiae]
Length = 530
Score = 511 bits (1316), Expect = e-143
Identities = 257/494 (52%), Positives = 351/494 (71%), Gaps = 12/494 (2%)
Query: 70 STGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQ--DEEKLSTVNTDWMH 127
ST IQ + + V R+PRF +L DD+ YF+ IL ++ +++ + E LS N DWM
Sbjct: 37 STKIQTRLTSENYPDVHRDPRFKKLTSDDLNYFKSILSEQEILRASESEDLSFYNEDWMR 96
Query: 128 KYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMN 187
KY+G SKL+L+P + ++VS IL YCN +AVVPQGGNTGLVGGSVP+FDE+I+SL+++N
Sbjct: 97 KYKGQSKLVLRPKSVEKVSLILNYCNDEKIAVVPQGGNTGLVGGSVPIFDELILSLANLN 156
Query: 188 KVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLV 247
K+ FD VSGIL C+AG ILEN +++ + ++ PLDLGAKGSC +GG V+TNAGGLRL+
Sbjct: 157 KIRDFDPVSGILKCDAGVILENANNYVMEQNYMFPLDLGAKGSCHVGGVVATNAGGLRLL 216
Query: 248 RYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPP 307
RYGSLHGSVLG+E V+ +G ++ + ++RKDNTGYDLK LFIGSEG++GI+T V+ILT P
Sbjct: 217 RYGSLHGSVLGLEVVMPNGQIVSSMHSMRKDNTGYDLKQLFIGSEGTIGIITGVSILTVP 276
Query: 308 KLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPT 367
K + NV+ L+ + + QK+ A+++L EILSAFEF+D++S L + L+ A PL
Sbjct: 277 KPKAFNVSYLSVESFEDVQKVFVRARQELSEILSAFEFMDAKSQVLAKSQLKDAAFPLED 336
Query: 368 LHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEA 427
H FY+LIET+GS++ D KLE FL + ME +++DGV+AQD + + W+ RE I EA
Sbjct: 337 EHPFYILIETSGSNKDHDDSKLETFLENVMEEGIVTDGVVAQDETELQNLWKWREMIPEA 396
Query: 428 LMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNA----------ANVVGYGHLGDGNLHLN 477
G VYKYD+S+PL++LY+LVE +RL A +GYGH+GDGNLHLN
Sbjct: 397 SQANGGVYKYDVSLPLKDLYSLVEATNARLSEAELVGDSPKPVVGAIGYGHVGDGNLHLN 456
Query: 478 ISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKN 537
++V +Y+ I +EPFVYE+ S +GS+SAEHGLG K N I YSKS E V++M +K
Sbjct: 457 VAVREYNKNIEKTLEPFVYEFVSSKHGSVSAEHGLGFQKKNYIGYSKSPEEVKMMKDLKV 516
Query: 538 LLDPNHILNPYKVL 551
DPN ILNPYK +
Sbjct: 517 HYDPNGILNPYKYI 530
>UniRef100_UPI000023F053 UPI000023F053 UniRef100 entry
Length = 1091
Score = 509 bits (1310), Expect = e-142
Identities = 263/507 (51%), Positives = 346/507 (67%), Gaps = 23/507 (4%)
Query: 71 TGIQH-KCYASMPGLVQRNPRFSELNDDDVRYFQEILGQK----------NVVQDEEKLS 119
T QH K + M ++R+ RF+++ + V F+EILG D
Sbjct: 54 TRAQHGKLTSEMYPQLERDSRFAQVTPEHVARFREILGDNPSAIIDGVTSGTGVDAADFE 113
Query: 120 TVNTDWMHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEV 179
T N DWMHKY+G SKL+L+P T++VS +LKYCN + LAVVPQGGNTGLVGGSVPVFDE+
Sbjct: 114 TYNEDWMHKYKGQSKLVLRPGTTEEVSSVLKYCNEQRLAVVPQGGNTGLVGGSVPVFDEI 173
Query: 180 IVSLSSMNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVST 239
++S++ MN++ SFD+VSG LV +AGC+LE + ++L +G+I PLDLGAKGSC +GGNV+T
Sbjct: 174 VISMARMNEIRSFDEVSGSLVIDAGCVLEAVDTYLAKKGYIFPLDLGAKGSCHVGGNVAT 233
Query: 240 NAGGLRLVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVT 299
NAGGLRL+RYGSLHGSVLGVEAVL +GT+++ L TLRK+NTGYD+K LFIG+EG+LGI+T
Sbjct: 234 NAGGLRLLRYGSLHGSVLGVEAVLPNGTIINDLCTLRKNNTGYDIKQLFIGAEGTLGIIT 293
Query: 300 KVAILTPPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLE 359
K+AI P + +VNVA+ + Y Q +EAK++L EILSAFE +D +S +++ ++
Sbjct: 294 KIAIQCPQRSPAVNVAVFGLESYEKAQLAFREAKKQLSEILSAFELMDGRSQR-IVSEVK 352
Query: 360 GARNPLPTLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWR 419
G +PL + FY LIET+GS+ D KLEAFL M E+I+DGV+AQD Q + W
Sbjct: 353 GQEHPLEGEYPFYCLIETSGSNSEHDYAKLEAFLEDVMTREVIADGVVAQDETQLRNLWG 412
Query: 420 LREGITEALMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNA-----------ANVVGYGH 468
REGITE L G YKYD+SIPL +Y LVE+ +SR+ A +V+GYGH
Sbjct: 413 WREGITECLGHWGGTYKYDISIPLNEMYKLVEDTKSRMTEAGLLGDTPDHPVVDVLGYGH 472
Query: 469 LGDGNLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLET 528
+GD NLHLNI V +YD + +EP+VY W K NGSISAEHGLG+ K I YS+ T
Sbjct: 473 MGDSNLHLNIPVRRYDPAVEKALEPWVYGWIQKRNGSISAEHGLGIAKKKFIGYSRDDTT 532
Query: 529 VQLMASIKNLLDPNHILNPYKVLPHSL 555
+ +M IKNL DP+ LPH L
Sbjct: 533 IGVMKQIKNLFDPHSQQEAITTLPHML 559
>UniRef100_Q7RYX6 Hypothetical protein [Neurospora crassa]
Length = 551
Score = 506 bits (1304), Expect = e-142
Identities = 258/485 (53%), Positives = 337/485 (69%), Gaps = 19/485 (3%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQD-------EEKLSTVNTDWMHKYEGSSKLLL 137
++R+PR+ ++ + V YF+ +LG ++ V D + + N+DWM KY G +L+L
Sbjct: 68 LKRDPRYGQVTKEHVDYFKGLLGTESAVIDGVTNENATDDIEPFNSDWMRKYRGHCRLVL 127
Query: 138 QPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSG 197
+P +T++VS+ILKYCN LAVVPQGGNTGLVGGSVPVFDE+++++ MN +I FD+VSG
Sbjct: 128 KPSSTEEVSKILKYCNDNKLAVVPQGGNTGLVGGSVPVFDEIVLNMGRMNNIIEFDEVSG 187
Query: 198 ILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVL 257
IL EAG ILE + FL ++G+I PLDLGAKGSC IGGN+STNAGGLRL+RYGSLHG+ L
Sbjct: 188 ILTVEAGAILEVVDQFLASKGYIFPLDLGAKGSCHIGGNLSTNAGGLRLLRYGSLHGTTL 247
Query: 258 GVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALL 317
G+EAVL DGTV+D L LRK+NTGYDLK LFIGSEG++G++TK I P + + NVAL
Sbjct: 248 GIEAVLPDGTVVDDLCKLRKNNTGYDLKQLFIGSEGTIGVITKAVIQCPQRPKAQNVALF 307
Query: 318 ACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIET 377
+ Y Q+ +EAK L EILSAFE +D+ S LV + +PL + FY LIET
Sbjct: 308 GLESYEAAQQAFREAKGHLSEILSAFELMDAGSQALV-RQVTKKNSPLEGEYPFYCLIET 366
Query: 378 TGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKY 437
+GS+ D +KL+ FL ME ++ DG LAQD Q + W REGITEAL +G VYKY
Sbjct: 367 SGSNSDHDGEKLQTFLEDVMEKGIVVDGTLAQDETQVKALWSWREGITEALGHLGGVYKY 426
Query: 438 DLSIPLENLYNLVEEMRSRLGNAA-----------NVVGYGHLGDGNLHLNISVPQYDDK 486
D+SIPL +Y LVE+ ++R+ A VVGYGH+GD NLHLN+S ++D++
Sbjct: 427 DVSIPLPEMYQLVEDTKARVEEAGLLGDTDEHPVRAVVGYGHMGDSNLHLNVSTRRFDER 486
Query: 487 ILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILN 546
+ +EPFVYEW SK GSISAEHGLGL K I YS++ V LM +K+L DPN ILN
Sbjct: 487 VEKVLEPFVYEWISKRQGSISAEHGLGLAKKKYIGYSRTPTMVGLMKQLKDLYDPNAILN 546
Query: 547 PYKVL 551
PYK +
Sbjct: 547 PYKYI 551
>UniRef100_Q75CD0 ACL021Cp [Ashbya gossypii]
Length = 534
Score = 506 bits (1302), Expect = e-142
Identities = 258/497 (51%), Positives = 352/497 (69%), Gaps = 17/497 (3%)
Query: 68 GISTGIQHKCYASMPGLVQRNPRFSELNDDDVRYFQEILGQKNVVQ--DEEKLSTVNTDW 125
G +T H + P LV R+ R+ +L ++D+ +F+ IL ++ ++Q + E L+ N DW
Sbjct: 38 GYATHAAHLTADTYPTLV-RDARYKKLGEEDIAFFRGILSEQEILQAGEGEDLALYNEDW 96
Query: 126 MHKYEGSSKLLLQPWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSS 185
M KY G SKL+L+P +T QV+ I++YCN + LAVVPQGGNTGLVGGSVPVFDE+++SL+
Sbjct: 97 MRKYRGQSKLVLRPKSTQQVAAIIRYCNEQRLAVVPQGGNTGLVGGSVPVFDEIVLSLAQ 156
Query: 186 MNKVISFDKVSGILVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLR 245
+NKV FD VSGIL C+AG ILEN S+L G++ PLDLGAKGSC +GG V+TNAGGLR
Sbjct: 157 LNKVRDFDPVSGILKCDAGVILENADSYLMERGYLFPLDLGAKGSCHVGGLVATNAGGLR 216
Query: 246 LVRYGSLHGSVLGVEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILT 305
L+RYGSLHGSVLG+E VL +G VL+ + LRKDNTG+DLK LFIGSEG++G++T V+IL
Sbjct: 217 LLRYGSLHGSVLGLEVVLPNGEVLNSMDALRKDNTGFDLKQLFIGSEGTIGVITGVSILC 276
Query: 306 PPKLSSVNVALLACKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPL 365
PP+ ++ NV LA ++Y Q++ +AK++LGEILSAFEF+D S + HL+G +P
Sbjct: 277 PPRPTAFNVCFLALENYARVQEVFIKAKKELGEILSAFEFMDFNSQYIAGQHLKGVAHPF 336
Query: 366 PTLHDFYVLIETTGSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGIT 425
+ FYVLIET GS++ D KLE FL +ME L+SDG LAQ + + W+ RE I
Sbjct: 337 SEKYPFYVLIETAGSNKEHDDLKLEQFLEGAMEEGLVSDGALAQGETEVRNLWQWREMIP 396
Query: 426 EALMRVGAVYKYDLSIPLENLYNLVEEMRSRLGNAANV-------------VGYGHLGDG 472
EA G VYKYD+S+PL+++++LV+ + RL A N+ +GYGH GDG
Sbjct: 397 EASASEGGVYKYDVSLPLKDMHSLVDAVNERL-TAQNLSDTEDASKPVVCALGYGHFGDG 455
Query: 473 NLHLNISVPQYDDKILSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLM 532
NLHLN++V +Y ++ + +EPFVYE+ + +GSISAEHGLG K N I YSKS + ++++
Sbjct: 456 NLHLNVAVREYTKQVEAALEPFVYEFVASKHGSISAEHGLGFQKKNYISYSKSPQEIKMI 515
Query: 533 ASIKNLLDPNHILNPYK 549
IK+ DPN ILNPYK
Sbjct: 516 KDIKHHYDPNAILNPYK 532
>UniRef100_UPI0000236145 UPI0000236145 UniRef100 entry
Length = 1217
Score = 505 bits (1300), Expect = e-141
Identities = 260/485 (53%), Positives = 337/485 (68%), Gaps = 18/485 (3%)
Query: 85 VQRNPRFSELNDDDVRYFQEILGQKNVVQD------EEKLSTVNTDWMHKYEGSSKLLLQ 138
++R+P F+E+ +DV+YF+++LG ++ V D + + N DWM KY G +KL+L+
Sbjct: 60 LKRSPEFAEITAEDVKYFKDLLGSESAVIDGVTTDATDDIEPFNGDWMRKYRGHTKLVLK 119
Query: 139 PWNTDQVSQILKYCNSRSLAVVPQGGNTGLVGGSVPVFDEVIVSLSSMNKVISFDKVSGI 198
P + ++VS++LKYCN + LAVVPQGGNTGLVGGSVPVFDE++++ S MNK+ SFD+ SG+
Sbjct: 120 PQSKEEVSKVLKYCNEKKLAVVPQGGNTGLVGGSVPVFDEIVINTSRMNKIRSFDEASGV 179
Query: 199 LVCEAGCILENIISFLDNEGFIMPLDLGAKGSCQIGGNVSTNAGGLRLVRYGSLHGSVLG 258
LV +AG ILE +L + PLDLGAKGSC IGGNV+TNAGGLRL+RYGSLHG+VLG
Sbjct: 180 LVVDAGVILEVADQYLAERHHLFPLDLGAKGSCHIGGNVATNAGGLRLLRYGSLHGTVLG 239
Query: 259 VEAVLADGTVLDMLKTLRKDNTGYDLKHLFIGSEGSLGIVTKVAILTPPKLSSVNVALLA 318
VEAVL DGT++D L TLRK+NTGYDLK LFIGSEG++GI+T V+IL PP+ +VNVA
Sbjct: 240 VEAVLPDGTIMDGLSTLRKNNTGYDLKQLFIGSEGTIGIITGVSILCPPRPKAVNVAYFG 299
Query: 319 CKDYNCCQKLLQEAKRKLGEILSAFEFLDSQSMNLVINHLEGARNPLPTLHDFYVLIETT 378
+ Y+ ++ EAK++L EILSAFE +D +S LV + G + PL + FY LIET+
Sbjct: 300 LESYDKVRQAFGEAKKQLSEILSAFELMDGRSQKLV-HASTGNKFPLEEEYPFYCLIETS 358
Query: 379 GSDESSDKQKLEAFLLHSMENELISDGVLAQDINQASSFWRLREGITEALMRVGAVYKYD 438
GS+ D +KLE FL M +++DGVLAQD Q S WR REGITEAL +G YKYD
Sbjct: 359 GSNAEHDMEKLETFLESVMGEGIVADGVLAQDETQFQSIWRWREGITEALSHLGGTYKYD 418
Query: 439 LSIPLENLYNLVEEMRSRLGN-----------AANVVGYGHLGDGNLHLNISVPQYDDKI 487
+SIPL LY LV++ R RL VVGYGH+GD NLHLNI+V QY+ +
Sbjct: 419 VSIPLPELYQLVDDCRERLTKMGFVGDDDSFPVRAVVGYGHMGDSNLHLNIAVRQYNKDV 478
Query: 488 LSQIEPFVYEWTSKHNGSISAEHGLGLMKANEILYSKSLETVQLMASIKNLLDPNHILNP 547
IEP+VYEW K NGSISAEHGLGL K I YS++ ++LM +K L DP
Sbjct: 479 EKAIEPWVYEWIQKRNGSISAEHGLGLAKKEFIGYSQNDTNLKLMKQLKELYDPTRGCVA 538
Query: 548 YKVLP 552
Y +P
Sbjct: 539 YSAMP 543
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 943,306,374
Number of Sequences: 2790947
Number of extensions: 40401129
Number of successful extensions: 103410
Number of sequences better than 10.0: 1637
Number of HSP's better than 10.0 without gapping: 1207
Number of HSP's successfully gapped in prelim test: 430
Number of HSP's that attempted gapping in prelim test: 99594
Number of HSP's gapped (non-prelim): 1896
length of query: 557
length of database: 848,049,833
effective HSP length: 133
effective length of query: 424
effective length of database: 476,853,882
effective search space: 202186045968
effective search space used: 202186045968
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0627.15


