

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0590b.4
(283 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
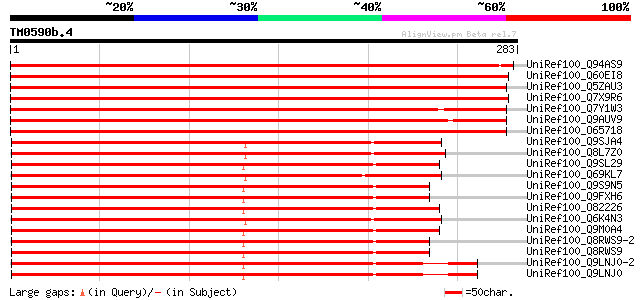
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q94AS9 Cyclic nucleotide-gated ion channel 4 [Arabidop... 505 e-142
UniRef100_Q60EI8 Putative cyclic nucleotide gated ion channel [O... 465 e-130
UniRef100_Q5ZAU3 Putative cyclic nucleotide and calmodulin-regul... 464 e-129
UniRef100_Q7X9R6 Putative cyclic nucleotide and calmodulin-regul... 456 e-127
UniRef100_Q7Y1W3 Putative cyclic nucleotide and calmodulin-regul... 336 5e-91
UniRef100_Q9AUV9 Putative cyclic nucleotide and calmodulin-regul... 335 1e-90
UniRef100_O65718 Cyclic nucleotide-gated ion channel 2 [Arabidop... 322 5e-87
UniRef100_Q9SJA4 Putative cyclic nucleotide-gated ion channel 14... 223 4e-57
UniRef100_Q8L7Z0 Probable cyclic nucleotide-gated ion channel 17... 221 2e-56
UniRef100_Q9SL29 Putative cyclic nucleotide-gated ion channel 15... 220 3e-56
UniRef100_Q69KL7 Putative cyclic nucleotide and calmodulin-regul... 215 1e-54
UniRef100_Q9S9N5 Putative cyclic nucleotide-gated ion channel 7 ... 215 1e-54
UniRef100_Q9FXH6 Putative cyclic nucleotide-gated ion channel 8 ... 213 5e-54
UniRef100_O82226 Probable cyclic nucleotide-gated ion channel 6 ... 212 8e-54
UniRef100_Q6K4N3 Cyclic nucleotide-gated calmodulin-binding ion ... 211 2e-53
UniRef100_Q9M0A4 Putative cyclic nucleotide-gated ion channel 9 ... 211 2e-53
UniRef100_Q8RWS9-2 Splice isoform 2 of Q8RWS9 [Arabidopsis thali... 209 5e-53
UniRef100_Q8RWS9 Probable cyclic nucleotide-gated ion channel 5 ... 209 5e-53
UniRef100_Q9LNJ0-2 Splice isoform 2 of Q9LNJ0 [Arabidopsis thali... 209 9e-53
UniRef100_Q9LNJ0 Probable cyclic nucleotide-gated ion channel 10... 209 9e-53
>UniRef100_Q94AS9 Cyclic nucleotide-gated ion channel 4 [Arabidopsis thaliana]
Length = 694
Score = 505 bits (1300), Expect = e-142
Identities = 239/281 (85%), Positives = 263/281 (93%), Gaps = 1/281 (0%)
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL ATTSKKQAM L+MRN+EWWM+KR LP G RQRVRNYERQRW AMRGVDEC+M++NL
Sbjct: 415 VFLHATTSKKQAMHLKMRNIEWWMKKRHLPIGFRQRVRNYERQRWAAMRGVDECEMVQNL 474
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P+GLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSL+FTKGETI +EGD V+RML
Sbjct: 475 PEGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLIFTKGETIQKEGDAVQRML 534
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
FVVRGHLQSSQ LRDGVKS C LGPGNFSGDELLSWCLRRPF+ERLPPSS+TLVTLETTE
Sbjct: 535 FVVRGHLQSSQLLRDGVKSCCMLGPGNFSGDELLSWCLRRPFVERLPPSSSTLVTLETTE 594
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
AFGL+AEDV+YVTQHFRYTF+ EKVKRSARYYSPGWRTWAAVA+QLAWRRYKHRL+LTSL
Sbjct: 595 AFGLDAEDVKYVTQHFRYTFVNEKVKRSARYYSPGWRTWAAVAVQLAWRRYKHRLTLTSL 654
Query: 241 SFIRPRRPASRCTSMEEDRLRLYTALLTSPKPNDQEEYDDF 281
SFIRPRRP SRC S+ ED+LRLY A+LTSPKPN +++DD+
Sbjct: 655 SFIRPRRPLSRCASLGEDKLRLYAAILTSPKPN-PDDFDDY 694
>UniRef100_Q60EI8 Putative cyclic nucleotide gated ion channel [Oryza sativa]
Length = 691
Score = 465 bits (1197), Expect = e-130
Identities = 216/278 (77%), Positives = 248/278 (88%)
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL+ATTSKKQAMQ R+R LEWWM + +P G RQRVR +ERQRW A RGVDECQ++R+L
Sbjct: 414 VFLNATTSKKQAMQTRLRGLEWWMEHKGVPHGFRQRVRQFERQRWAATRGVDECQIVRDL 473
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P+GLRRDIKYHLCLDLVRQVPLF HMDDLVLENICDRVKSL+F KGE I REGDPV+RML
Sbjct: 474 PEGLRRDIKYHLCLDLVRQVPLFHHMDDLVLENICDRVKSLIFPKGEIIVREGDPVQRML 533
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
F+VRGHLQ SQ +R+G S+CTLGPGNFSGDELLSWC+RRPF+ERLP SS+TLVT E+TE
Sbjct: 534 FIVRGHLQCSQVMRNGATSWCTLGPGNFSGDELLSWCMRRPFMERLPASSSTLVTAESTE 593
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
AFGLEA DV+YVTQHFRYTF +KV+RSARYYS GWRTWAAVA+QLAWRRYKHR +L SL
Sbjct: 594 AFGLEAGDVKYVTQHFRYTFTSDKVRRSARYYSHGWRTWAAVAVQLAWRRYKHRKTLASL 653
Query: 241 SFIRPRRPASRCTSMEEDRLRLYTALLTSPKPNDQEEY 278
SFIRPRRP SRC+S+ E++LRLYTA+LTSPKPN +++
Sbjct: 654 SFIRPRRPLSRCSSLGEEKLRLYTAILTSPKPNQDDDF 691
>UniRef100_Q5ZAU3 Putative cyclic nucleotide and calmodulin-regulated ion channel
[Oryza sativa]
Length = 666
Score = 464 bits (1194), Expect = e-129
Identities = 216/277 (77%), Positives = 249/277 (88%)
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL+A TSKKQAMQ R+R +EWWM++++LPQ R RVR +ERQRW A RGVDEC+++R+L
Sbjct: 387 VFLNAATSKKQAMQTRLRGVEWWMKRKKLPQSFRHRVRQHERQRWAATRGVDECRIVRDL 446
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P+GLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVF KGE I REGDPV+RML
Sbjct: 447 PEGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFPKGEIIVREGDPVQRML 506
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
F+VRGHLQSSQ LR G S CTLGPGNFSGDELLSWC+RRPF+ERLP SS+TLVT+E+TE
Sbjct: 507 FIVRGHLQSSQVLRTGATSCCTLGPGNFSGDELLSWCMRRPFLERLPASSSTLVTMESTE 566
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
AFGLEA DV+YVTQHFRYTF ++V+RSARYYS GWRTWAAVA+QLAWRRYKHR +L SL
Sbjct: 567 AFGLEAADVKYVTQHFRYTFTNDRVRRSARYYSHGWRTWAAVAVQLAWRRYKHRKTLASL 626
Query: 241 SFIRPRRPASRCTSMEEDRLRLYTALLTSPKPNDQEE 277
SFIRPRRP SRC+S+ E++LRLYTA+LTSPKPN ++
Sbjct: 627 SFIRPRRPLSRCSSLGEEKLRLYTAILTSPKPNPNQD 663
>UniRef100_Q7X9R6 Putative cyclic nucleotide and calmodulin-regulated ion channel
protein [Hordeum vulgare var. distichum]
Length = 587
Score = 456 bits (1173), Expect = e-127
Identities = 210/278 (75%), Positives = 246/278 (87%)
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL+ATTSKKQAM R+R+LEWWM+++ LPQ R RVR +ERQRW A RGVDECQ++R+L
Sbjct: 310 VFLNATTSKKQAMHTRLRSLEWWMKRKDLPQSYRHRVRQFERQRWAATRGVDECQIVRDL 369
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P+ LRRDIKYHLCLDLVRQVPLF HMDDLVLEN+CDRV+SL++ KGETI REGDPV+RM+
Sbjct: 370 PEALRRDIKYHLCLDLVRQVPLFHHMDDLVLENMCDRVRSLIYPKGETIVREGDPVQRMV 429
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
F+VRGHLQ SQ+LR+G S C LGPGNF+GDELL WCLRRPF ERLP SS+TLVTLE+TE
Sbjct: 430 FIVRGHLQCSQELRNGGTSCCMLGPGNFTGDELLPWCLRRPFAERLPASSSTLVTLESTE 489
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
FGLEA DV+YVTQHFRYTF E+V+RSARYYSPGWRTWAAVA+QLAWRRYKHR +L+SL
Sbjct: 490 VFGLEAADVKYVTQHFRYTFTNERVRRSARYYSPGWRTWAAVAVQLAWRRYKHRKTLSSL 549
Query: 241 SFIRPRRPASRCTSMEEDRLRLYTALLTSPKPNDQEEY 278
SFIRPRRP SRC+S+ E++LRLYTA+L SPKPN + +
Sbjct: 550 SFIRPRRPLSRCSSLGEEKLRLYTAILASPKPNQDDGF 587
>UniRef100_Q7Y1W3 Putative cyclic nucleotide and calmodulin-regulated ion channel
protein [Hordeum vulgare var. distichum]
Length = 415
Score = 336 bits (861), Expect = 5e-91
Identities = 161/278 (57%), Positives = 211/278 (74%), Gaps = 4/278 (1%)
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL A ++K+ MQLR R++EWWMR+RQLP LRQRVR YER+RW A+ G +E +MI++L
Sbjct: 141 VFLHAVLARKRKMQLRFRDMEWWMRRRQLPSRLRQRVRKYERERWAAITGDEEMEMIKDL 200
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P+GLRRDIK +LCL+LV+QVPLF MD+L+L+NICDR++ LVF GE + REGDPV+RM+
Sbjct: 201 PEGLRRDIKRYLCLELVKQVPLFHGMDELILDNICDRLRPLVFCGGEKVIREGDPVQRMV 260
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
FV++G L+S+Q L GV + C LG G+F GDELLSWCLRRPF++RLP SSAT +E +
Sbjct: 261 FVLQGKLRSTQPLTKGVVAECVLGAGSFLGDELLSWCLRRPFVDRLPASSATFECVEAAQ 320
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
AF L+A D+RY+T+HFRY F +K+KR+ARYYS WRTWAAV +QLAWRRY+ R+ T+
Sbjct: 321 AFCLDAPDLRYITEHFRYKFANDKLKRTARYYSSNWRTWAAVNVQLAWRRYRARMMATA- 379
Query: 241 SFIRPRRPASRCTSMEED-RLRLYTALLTSPKPNDQEE 277
+ P PA + D RLR Y A+ S +P+D E
Sbjct: 380 --VLPPPPAGAAGPEDGDRRLRHYAAMFMSLRPHDHLE 415
>UniRef100_Q9AUV9 Putative cyclic nucleotide and calmodulin-regulated ion channel
protein [Oryza sativa]
Length = 782
Score = 335 bits (858), Expect = 1e-90
Identities = 162/277 (58%), Positives = 210/277 (75%), Gaps = 2/277 (0%)
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL A ++K+ MQLR R++EWWMR+RQLP LRQRVR YER+RW A+ G +E +MI++L
Sbjct: 508 VFLHAVLARKRKMQLRFRDMEWWMRRRQLPSRLRQRVRKYERERWAAITGDEEMEMIKDL 567
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P+GLRRDIK +LCL+LV+QVPLF MDDL+L+NICDR++ LVF+ GE + REGDPV+RM+
Sbjct: 568 PEGLRRDIKRYLCLELVKQVPLFHGMDDLILDNICDRLRPLVFSSGEKVIREGDPVQRMV 627
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
FV++G L+S+Q L GV + C LG GNF GDELLSWCLRRP ++RLP SSAT +ET +
Sbjct: 628 FVLQGKLRSTQPLAKGVVATCMLGAGNFLGDELLSWCLRRPSLDRLPASSATFECVETAQ 687
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
AF L+A D+R++T+ FRY F EK+KR+ARYYS WRTWAAV IQLAWRRYK R +
Sbjct: 688 AFCLDAPDLRFITEQFRYKFANEKLKRTARYYSSNWRTWAAVNIQLAWRRYKARTTTDLA 747
Query: 241 SFIRPRRPASRCTSMEEDRLRLYTALLTSPKPNDQEE 277
S +P P++ + RLR Y A+ S +P+D E
Sbjct: 748 SAAQP--PSAGGPDDGDRRLRHYAAMFMSLRPHDHLE 782
>UniRef100_O65718 Cyclic nucleotide-gated ion channel 2 [Arabidopsis thaliana]
Length = 726
Score = 322 bits (826), Expect = 5e-87
Identities = 155/277 (55%), Positives = 198/277 (70%)
Query: 1 VFLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNL 60
VFL A +KK+ MQ+R R++EWWM++RQLP LRQRVR +ERQRW A+ G DE ++I +L
Sbjct: 450 VFLHAVMAKKRKMQIRCRDMEWWMKRRQLPSRLRQRVRRFERQRWNALGGEDELELIHDL 509
Query: 61 PQGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRML 120
P GLRRDIK +LC DL+ +VPLF+ MDDL+L+NICDR K VF+K E I REGDPV+RM+
Sbjct: 510 PPGLRRDIKRYLCFDLINKVPLFRGMDDLILDNICDRAKPRVFSKDEKIIREGDPVQRMI 569
Query: 121 FVVRGHLQSSQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETTE 180
F++RG ++ Q L GV + TL PG + GDELLSWCLRRPF++RLPPSSAT V LE E
Sbjct: 570 FIMRGRVKRIQSLSKGVLATSTLEPGGYLGDELLSWCLRRPFLDRLPPSSATFVCLENIE 629
Query: 181 AFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTSL 240
AF L +ED+RY+T HFRY F E++KR+ARYYS WRTWAAV IQ+AWRR + R ++
Sbjct: 630 AFSLGSEDLRYITDHFRYKFANERLKRTARYYSSNWRTWAAVNIQMAWRRRRKRTRGENI 689
Query: 241 SFIRPRRPASRCTSMEEDRLRLYTALLTSPKPNDQEE 277
+ E RL Y A+ S +P+D E
Sbjct: 690 GGSMSPVSENSIEGNSERRLLQYAAMFMSIRPHDHLE 726
>UniRef100_Q9SJA4 Putative cyclic nucleotide-gated ion channel 14 [Arabidopsis
thaliana]
Length = 726
Score = 223 bits (568), Expect = 4e-57
Identities = 117/242 (48%), Positives = 158/242 (64%), Gaps = 3/242 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + +L+ R+ E WM R LPQ LR+RVR + + +W A RGVDE ++ +LP
Sbjct: 401 YLQSITVRLEEWRLKRRDTEEWMGHRLLPQNLRERVRRFVQYKWLATRGVDEETILHSLP 460
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
LRRDI+ HLCLDLVR+VPLF MDD +L+ IC+R+ S + T+G I REGDPV MLF
Sbjct: 461 ADLRRDIQRHLCLDLVRRVPLFAQMDDQLLDAICERLASSLSTQGNYIVREGDPVTEMLF 520
Query: 122 VVRGHLQSS--QDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+SS R G + TL PG+F G+ELL+W L LP S+ T+ LE
Sbjct: 521 IIRGKLESSTTNGGRTGFFNSITLRPGDFCGEELLAWALLPKSTVNLPSSTRTVRALEEV 580
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L+A D+++V FR K K++ + RYYS WRTWAA +Q+AWRRYK + S
Sbjct: 581 EAFALQAGDLKFVANQFRRLHSK-KLQHTFRYYSHQWRTWAACFVQVAWRRYKRKKLAKS 639
Query: 240 LS 241
LS
Sbjct: 640 LS 641
>UniRef100_Q8L7Z0 Probable cyclic nucleotide-gated ion channel 17 [Arabidopsis
thaliana]
Length = 720
Score = 221 bits (562), Expect = 2e-56
Identities = 116/244 (47%), Positives = 158/244 (64%), Gaps = 3/244 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + +L+ R+ E WMR RQLP+ LR RVR YE+ +W A RGVDE ++++LP
Sbjct: 401 YLQSLTVRLEEWRLKKRDTEEWMRHRQLPEELRNRVRRYEQYKWLATRGVDEEVLLQSLP 460
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
LRRDI+ HLCLDLVR+VP F MDD +L+ IC+R+ S + T+G + REGD + MLF
Sbjct: 461 TDLRRDIQRHLCLDLVRRVPFFSQMDDQLLDAICERLVSSLCTEGTYLVREGDLISEMLF 520
Query: 122 VVRGHLQSS--QDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+SS R G + L PG+F G+ELLSW L LP S+ T+ L
Sbjct: 521 IIRGRLESSTTNGGRTGFFNSIILRPGDFCGEELLSWALLPKSTLNLPSSTRTVRALVEV 580
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L AED+++V FR K K++ + R+YS WRTWAA IQ AWRRYK R+ +
Sbjct: 581 EAFALRAEDLKFVANQFRRLHSK-KLQHTFRFYSHHWRTWAACFIQAAWRRYKRRVMENN 639
Query: 240 LSFI 243
L+ I
Sbjct: 640 LTAI 643
>UniRef100_Q9SL29 Putative cyclic nucleotide-gated ion channel 15 [Arabidopsis
thaliana]
Length = 678
Score = 220 bits (561), Expect = 3e-56
Identities = 112/241 (46%), Positives = 156/241 (64%), Gaps = 3/241 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L +TT + + ++R + E WM RQLP LRQ VR Y++ +W A RGVDE ++ +LP
Sbjct: 391 YLQSTTMRLEEWRIRRTDTEQWMHHRQLPPELRQAVRKYDQYKWLATRGVDEEALLISLP 450
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
LRRDIK HLC DLVR+VPLF MD+ +L+ IC+R+K + T+G + REGDPV MLF
Sbjct: 451 LDLRRDIKRHLCFDLVRRVPLFDQMDERMLDAICERLKPALCTEGTFLVREGDPVNEMLF 510
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RGHL S + R G + C +GPG+F G+ELL+W L + LP S+ T+ +
Sbjct: 511 IIRGHLDSYTTNGGRTGFFNSCLIGPGDFCGEELLTWALDPRPVVILPSSTRTVKAICEV 570
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L+AED+++V FR K+ ++ R+YS WRTWAA IQ AWRR++ R T
Sbjct: 571 EAFALKAEDLQFVASQFRRLHTKQ-LRHKFRFYSHQWRTWAACFIQAAWRRHRKRKYKTE 629
Query: 240 L 240
L
Sbjct: 630 L 630
>UniRef100_Q69KL7 Putative cyclic nucleotide and calmodulin-regulated ion channel
[Oryza sativa]
Length = 685
Score = 215 bits (547), Expect = 1e-54
Identities = 115/242 (47%), Positives = 151/242 (61%), Gaps = 3/242 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T++ + +++ R+ E WMR RQLPQ LR+RVR + +W A RGVDE +++ LP
Sbjct: 392 YLQSITARVEEWRIKQRDTEEWMRHRQLPQKLRERVRRFVHYKWLATRGVDEESILKALP 451
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
LRRDIK HLCLDLV +VP F MD +L+ IC+R+ S + T G I REGDPV MLF
Sbjct: 452 ADLRRDIKRHLCLDLVCRVPFFSQMDGQLLDAICERLVSSLSTVGTYIVREGDPVTEMLF 511
Query: 122 VVRGHLQSS--QDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+SS R G + TL G+F G+ELL W L LP S+ T+ T+
Sbjct: 512 IIRGKLESSTTDGGRTGFFNSITLKTGDFCGEELLGWALVPKPTVNLPSSTRTVKTIVEV 571
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L AED+++V FR K++ + RYYS WRTWAA IQ AWRRYK R
Sbjct: 572 EAFALRAEDLKFVASQFR-RLHSRKLQHTFRYYSHHWRTWAACFIQAAWRRYKRRRLAKD 630
Query: 240 LS 241
LS
Sbjct: 631 LS 632
>UniRef100_Q9S9N5 Putative cyclic nucleotide-gated ion channel 7 [Arabidopsis
thaliana]
Length = 738
Score = 215 bits (547), Expect = 1e-54
Identities = 107/235 (45%), Positives = 156/235 (65%), Gaps = 3/235 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + M+++ R+ E WM R LPQ LR+RVR Y++ +W RGVDE ++++LP
Sbjct: 422 YLQSLTVRLEEMRIKRRDSEQWMHHRSLPQNLRERVRRYDQYKWLETRGVDEENIVQSLP 481
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCL+LVR+VPLF +MD+ +L+ IC+R+K +FT+ I REGDPV M+F
Sbjct: 482 KDLRRDIKRHLCLNLVRRVPLFANMDERLLDAICERLKPSLFTESTYIVREGDPVNEMMF 541
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+S + R G + L G+F G+ELL+W L LP S+ T+ L
Sbjct: 542 IIRGRLESVTTDGGRSGFFNRGLLKEGDFCGEELLTWALDPKAGSNLPSSTRTVKALTEV 601
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHR 234
EAF LEAE++++V FR ++ V+++ R+YS WRTWA+ IQ AWRRY R
Sbjct: 602 EAFALEAEELKFVASQFRRLHSRQ-VQQTFRFYSQQWRTWASCFIQAAWRRYSRR 655
>UniRef100_Q9FXH6 Putative cyclic nucleotide-gated ion channel 8 [Arabidopsis
thaliana]
Length = 753
Score = 213 bits (542), Expect = 5e-54
Identities = 107/235 (45%), Positives = 156/235 (65%), Gaps = 3/235 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + M+++ R+ E WM R LPQ LR+RVR Y++ +W RGVDE ++++LP
Sbjct: 428 YLQSLTVRLEEMRIKRRDSEQWMHHRSLPQNLRERVRRYDQYKWLETRGVDEENIVQSLP 487
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCL+LVR+VPLF +MD+ +L+ IC+R+K ++T+ I REGDPV MLF
Sbjct: 488 KDLRRDIKRHLCLNLVRRVPLFANMDERLLDAICERLKPSLYTESTYIVREGDPVNEMLF 547
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+S + R G + L G+F G+ELL+W L LP S+ T+ L
Sbjct: 548 IIRGRLESVTTDGGRSGFFNRGLLKEGDFCGEELLTWALDPKAGSNLPSSTRTVKALTEV 607
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHR 234
EAF LEAE++++V FR ++ V+++ R+YS WRTWAA IQ AWRR+ R
Sbjct: 608 EAFALEAEELKFVASQFRRLHSRQ-VQQTFRFYSQQWRTWAACFIQAAWRRHLRR 661
>UniRef100_O82226 Probable cyclic nucleotide-gated ion channel 6 [Arabidopsis
thaliana]
Length = 747
Score = 212 bits (540), Expect = 8e-54
Identities = 108/241 (44%), Positives = 155/241 (63%), Gaps = 3/241 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + M+++ R+ E WM R LP LR+RVR Y++ +W RGVDE +++NLP
Sbjct: 434 YLQSLTIRLEEMRVKRRDSEQWMHHRMLPPELRERVRRYDQYKWLETRGVDEENLVQNLP 493
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCL LVR+VPLF++MD+ +L+ IC+R+K +FT+ + REGDPV MLF
Sbjct: 494 KDLRRDIKRHLCLALVRRVPLFENMDERLLDAICERLKPCLFTEKSYLVREGDPVNEMLF 553
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+S + R G + L G+F GDELL+W L LP S+ T+ L
Sbjct: 554 IIRGRLESVTTDGGRSGFYNRSLLKEGDFCGDELLTWALDPKSGSNLPSSTRTVKALTEV 613
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L A+++++V FR ++ V+ + R+YS WRTWAA +Q AWRRY R L
Sbjct: 614 EAFALIADELKFVASQFRRLHSRQ-VQHTFRFYSQQWRTWAACFMQAAWRRYIKRKKLEQ 672
Query: 240 L 240
L
Sbjct: 673 L 673
>UniRef100_Q6K4N3 Cyclic nucleotide-gated calmodulin-binding ion channel-like [Oryza
sativa]
Length = 449
Score = 211 bits (537), Expect = 2e-53
Identities = 113/242 (46%), Positives = 153/242 (62%), Gaps = 3/242 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + +L+ R+ E WMR RQLP LR+RVR + + +W A RGV+E +++ LP
Sbjct: 149 YLQSITVRVEEWRLKQRDTEEWMRHRQLPHELRERVRRFIQYKWLATRGVNEESILQALP 208
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
LRRDIK HLCL LVR+VP F MD+ +L+ IC+R+ S + T+G I REGDPV MLF
Sbjct: 209 ADLRRDIKRHLCLGLVRRVPFFSQMDNQLLDAICERLVSSLCTQGTYIVREGDPVTEMLF 268
Query: 122 VVRGHLQSS--QDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+SS R G + TL G+F G+ELL W L LP S+ T+ L
Sbjct: 269 IIRGKLESSTTNGGRTGFFNSTTLKSGDFCGEELLGWALVPKPTVNLPSSTRTVKALIEV 328
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L+AED+++V FR K +++ + RYYS WRTWA+ IQ AWRRYK R
Sbjct: 329 EAFALQAEDLKFVANQFRRLHSK-RLQHTFRYYSHHWRTWASCFIQAAWRRYKRRKMARD 387
Query: 240 LS 241
LS
Sbjct: 388 LS 389
>UniRef100_Q9M0A4 Putative cyclic nucleotide-gated ion channel 9 [Arabidopsis
thaliana]
Length = 733
Score = 211 bits (537), Expect = 2e-53
Identities = 106/241 (43%), Positives = 155/241 (63%), Gaps = 3/241 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + M+++ R+ E WM R LP LR+RVR Y++ +W RGVDE +++NLP
Sbjct: 433 YLQSLTIRLEEMRVKRRDSEQWMHHRMLPPELRERVRRYDQYKWLETRGVDEENLVQNLP 492
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCL LVR+VPLF++MD+ +L+ IC+R+K ++T+ + REGDPV MLF
Sbjct: 493 KDLRRDIKRHLCLALVRRVPLFENMDERLLDAICERLKPCLYTESSYLVREGDPVNEMLF 552
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+S + R G + L G+F G+ELL+W L LP S+ T L
Sbjct: 553 IIRGRLESVTTDGGRSGFFNRSLLKEGDFCGEELLTWALDPKSGSNLPSSTRTAKALTEV 612
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L A+++++V FR ++ V+ + R+YS WRTWAA+ IQ AWRRY + L
Sbjct: 613 EAFALIADELKFVASQFRRLHSRQ-VQHTFRFYSQQWRTWAAIFIQAAWRRYVKKKKLEQ 671
Query: 240 L 240
L
Sbjct: 672 L 672
>UniRef100_Q8RWS9-2 Splice isoform 2 of Q8RWS9 [Arabidopsis thaliana]
Length = 710
Score = 209 bits (533), Expect = 5e-53
Identities = 108/235 (45%), Positives = 151/235 (63%), Gaps = 3/235 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + M+++ R+ E WM R LPQ LR+RVR Y++ +W RGVDE +++NLP
Sbjct: 411 YLQSLTIRLEEMRVKRRDSEQWMHHRMLPQDLRERVRRYDQYKWLETRGVDEEYLVQNLP 470
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCL LVR+VPLF+ MDD +L+ IC R+K +FT+ + REGDPV MLF
Sbjct: 471 KDLRRDIKRHLCLALVRRVPLFKSMDDKLLDAICMRLKPCLFTESTYLVREGDPVDEMLF 530
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+S + R G + L G F G+ELL+W L LP S+ T+ L
Sbjct: 531 IIRGRLESVTTDGGRSGFFNRSLLKEGEFCGEELLTWALDPKSGVNLPSSTRTVKALTEV 590
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHR 234
EAF L +E++++V FR ++ V+ + R+YS WRTWAA IQ AWRRY R
Sbjct: 591 EAFALTSEELKFVASQFRRLHSRQ-VQHTFRFYSHQWRTWAACFIQAAWRRYCKR 644
>UniRef100_Q8RWS9 Probable cyclic nucleotide-gated ion channel 5 [Arabidopsis
thaliana]
Length = 717
Score = 209 bits (533), Expect = 5e-53
Identities = 108/235 (45%), Positives = 151/235 (63%), Gaps = 3/235 (1%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L + T + + M+++ R+ E WM R LPQ LR+RVR Y++ +W RGVDE +++NLP
Sbjct: 418 YLQSLTIRLEEMRVKRRDSEQWMHHRMLPQDLRERVRRYDQYKWLETRGVDEEYLVQNLP 477
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCL LVR+VPLF+ MDD +L+ IC R+K +FT+ + REGDPV MLF
Sbjct: 478 KDLRRDIKRHLCLALVRRVPLFKSMDDKLLDAICMRLKPCLFTESTYLVREGDPVDEMLF 537
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
++RG L+S + R G + L G F G+ELL+W L LP S+ T+ L
Sbjct: 538 IIRGRLESVTTDGGRSGFFNRSLLKEGEFCGEELLTWALDPKSGVNLPSSTRTVKALTEV 597
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHR 234
EAF L +E++++V FR ++ V+ + R+YS WRTWAA IQ AWRRY R
Sbjct: 598 EAFALTSEELKFVASQFRRLHSRQ-VQHTFRFYSHQWRTWAACFIQAAWRRYCKR 651
>UniRef100_Q9LNJ0-2 Splice isoform 2 of Q9LNJ0 [Arabidopsis thaliana]
Length = 706
Score = 209 bits (531), Expect = 9e-53
Identities = 111/262 (42%), Positives = 160/262 (60%), Gaps = 16/262 (6%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L +TT +++ M++R R+ E WM R LP+ LR+R+R YE+ RW RGV+E ++RNLP
Sbjct: 388 YLESTTVREEEMRVRKRDAEQWMSHRMLPEDLRKRIRRYEQYRWQETRGVEEETLLRNLP 447
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCLDL+++VPLF+ MD+ +L+ +CDR++ +++T+ + REGDPV MLF
Sbjct: 448 KDLRRDIKRHLCLDLLKKVPLFEIMDEQLLDAVCDRLRPVLYTENSYVIREGDPVGEMLF 507
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
V+RG L S + R G + L +F G++LL W L P S+ T+ L
Sbjct: 508 VMRGRLVSATTNGGRSGFFNAVNLKASDFCGEDLLPWALDPQSSSHFPISTRTVQALTEV 567
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L AED++ V FR K+ ++ + R+YS WRTW+ IQ AWRRY
Sbjct: 568 EAFALTAEDLKSVASQFRRLHSKQ-LQHTFRFYSVQWRTWSVSFIQAAWRRY-------- 618
Query: 240 LSFIRPRRPASRCTSMEEDRLR 261
RR ++ EEDRLR
Sbjct: 619 -----CRRKLAKSLRDEEDRLR 635
>UniRef100_Q9LNJ0 Probable cyclic nucleotide-gated ion channel 10 [Arabidopsis
thaliana]
Length = 711
Score = 209 bits (531), Expect = 9e-53
Identities = 111/262 (42%), Positives = 160/262 (60%), Gaps = 16/262 (6%)
Query: 2 FLSATTSKKQAMQLRMRNLEWWMRKRQLPQGLRQRVRNYERQRWTAMRGVDECQMIRNLP 61
+L +TT +++ M++R R+ E WM R LP+ LR+R+R YE+ RW RGV+E ++RNLP
Sbjct: 393 YLESTTVREEEMRVRKRDAEQWMSHRMLPEDLRKRIRRYEQYRWQETRGVEEETLLRNLP 452
Query: 62 QGLRRDIKYHLCLDLVRQVPLFQHMDDLVLENICDRVKSLVFTKGETITREGDPVKRMLF 121
+ LRRDIK HLCLDL+++VPLF+ MD+ +L+ +CDR++ +++T+ + REGDPV MLF
Sbjct: 453 KDLRRDIKRHLCLDLLKKVPLFEIMDEQLLDAVCDRLRPVLYTENSYVIREGDPVGEMLF 512
Query: 122 VVRGHLQS--SQDLRDGVKSYCTLGPGNFSGDELLSWCLRRPFIERLPPSSATLVTLETT 179
V+RG L S + R G + L +F G++LL W L P S+ T+ L
Sbjct: 513 VMRGRLVSATTNGGRSGFFNAVNLKASDFCGEDLLPWALDPQSSSHFPISTRTVQALTEV 572
Query: 180 EAFGLEAEDVRYVTQHFRYTFMKEKVKRSARYYSPGWRTWAAVAIQLAWRRYKHRLSLTS 239
EAF L AED++ V FR K+ ++ + R+YS WRTW+ IQ AWRRY
Sbjct: 573 EAFALTAEDLKSVASQFRRLHSKQ-LQHTFRFYSVQWRTWSVSFIQAAWRRY-------- 623
Query: 240 LSFIRPRRPASRCTSMEEDRLR 261
RR ++ EEDRLR
Sbjct: 624 -----CRRKLAKSLRDEEDRLR 640
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.324 0.137 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 458,580,339
Number of Sequences: 2790947
Number of extensions: 17955533
Number of successful extensions: 49152
Number of sequences better than 10.0: 572
Number of HSP's better than 10.0 without gapping: 442
Number of HSP's successfully gapped in prelim test: 130
Number of HSP's that attempted gapping in prelim test: 48441
Number of HSP's gapped (non-prelim): 630
length of query: 283
length of database: 848,049,833
effective HSP length: 126
effective length of query: 157
effective length of database: 496,390,511
effective search space: 77933310227
effective search space used: 77933310227
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0590b.4


