

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0541.15
(322 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
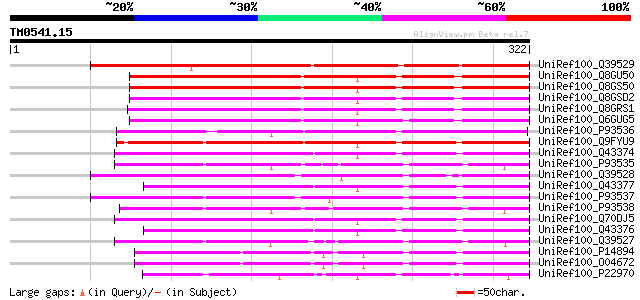
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q39529 Agglutinin II precursor [Cladrastis lutea] 213 6e-54
UniRef100_Q8GU50 Lectin precursor [Pterocarpus angolensis] 208 1e-52
UniRef100_Q8GS50 Lectin precursor [Pterocarpus angolensis] 208 1e-52
UniRef100_Q8GSD2 Lectin precursor [Pterocarpus angolensis] 206 7e-52
UniRef100_Q8GRS1 Lectin precursor [Pterocarpus angolensis] 205 2e-51
UniRef100_Q6GUG5 Lectin [Pterocarpus rotundifolius] 201 2e-50
UniRef100_P93536 Bark lectin II precursor [Sophora japonica] 201 3e-50
UniRef100_Q9FYU9 Lectin [Sophora flavescens] 196 9e-49
UniRef100_Q43374 Mannose/glucose-binding lectin precursor [Arach... 187 3e-46
UniRef100_P93535 Seed lectin precursor [Sophora japonica] 183 5e-45
UniRef100_Q39528 Agglutinin I precursor [Cladrastis lutea] 183 6e-45
UniRef100_Q43377 Mannose/glucose-binding lectin precursor [Arach... 182 8e-45
UniRef100_P93537 Bark lectin I precursor [Sophora japonica] 182 8e-45
UniRef100_P93538 Bark lectin precursor [Sophora japonica] 181 2e-44
UniRef100_Q70DJ5 Putative lectin [Arachis hypogaea] 178 2e-43
UniRef100_Q43376 Mannose/glucose-binding lectin precursor [Arach... 176 6e-43
UniRef100_Q39527 Lectin-related protein precursor [Cladrastis lu... 175 1e-42
UniRef100_P14894 Concanavalin A precursor [Canavalia gladiata] 171 2e-41
UniRef100_O04672 Lectin [Canavalia brasiliensis] 170 4e-41
UniRef100_P22970 Anti-H(O) lectin I [Cytisus sessilifolius] 169 9e-41
>UniRef100_Q39529 Agglutinin II precursor [Cladrastis lutea]
Length = 290
Score = 213 bits (542), Expect = 6e-54
Identities = 120/276 (43%), Positives = 170/276 (61%), Gaps = 9/276 (3%)
Query: 51 SQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN 110
S N T I+ LLA + L+L+ SDSLSF+F F PD R ++L GDA ++
Sbjct: 4 SNTNLLQTKKPISLPLLAFITLFLMLLNRVNSSDSLSFTFDNFRPDQRDLILQGDAKISS 63
Query: 111 G--TLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPH-GD 167
G +LQLTK D G P S G + +Y +HL D T R+A F+T FTFV++ + GD
Sbjct: 64 GGDSLQLTKTDTSGKPVRGSVGRALYYTPLHLWDSSTNRLASFQTTFTFVLSSPTNNPGD 123
Query: 168 GFTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKNSIVAVEFDSF-SNSFDPNFPHIG 226
G FF+A + P G GG LGLF + A N S N IVAVEFD+F +N++DP+ HIG
Sbjct: 124 GIAFFIAPPETTIPPGSSGG-LLGLFSPDNALNNSLNQIVAVEFDTFVNNNWDPSHRHIG 182
Query: 227 IDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPI 286
ID+NTI+S+ T+ W + G++ A+ISY+S ++ L+VV YP Q + + Y +
Sbjct: 183 IDVNTIKSSATVRWQ---RENGSLATAQISYNSDTKKLSVVSSYPNTQANEDY-TVSYDV 238
Query: 287 DIESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
D+++ L EWV +GFS +TG + + H+ILSW+FNS L
Sbjct: 239 DLKTELPEWVRVGFSGSTGGYVQNHNILSWTFNSNL 274
>UniRef100_Q8GU50 Lectin precursor [Pterocarpus angolensis]
Length = 260
Score = 208 bits (530), Expect = 1e-52
Identities = 112/253 (44%), Positives = 153/253 (60%), Gaps = 13/253 (5%)
Query: 75 LLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAF 134
+L+ DSLSF FPTF D + ++ GDA T N +QLTK D GNP + G F
Sbjct: 1 MLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQTKNNAVQLTKTDSNGNPVASTVGRILF 60
Query: 135 YGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPEGGGGGGFLGLF 193
VHL +K + RVA+F+++F+F + +G DG FF+A D P G GGG LGLF
Sbjct: 61 SAQVHLWEKSSSRVANFQSQFSFSLKSPLSNGADGIAFFIAPPDTTIP-SGSGGGLLGLF 119
Query: 194 DEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGA 249
TA NTS N ++AVEFD+F SN++DPN+PHIGID+N+IRS T+ W D+ G
Sbjct: 120 APGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYPHIGIDVNSIRSVKTVKW---DRRDGQ 176
Query: 250 IQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAE 309
N ++++ +R+L VV Y G + Y +D+ S L EWV +GFSA +GE +
Sbjct: 177 SLNVLVTFNPSTRNLDVVATYSDGTRY----EVSYEVDVRSVLPEWVGVGFSAASGEQYQ 232
Query: 310 THSILSWSFNSTL 322
TH++ SWSF STL
Sbjct: 233 THTLESWSFTSTL 245
>UniRef100_Q8GS50 Lectin precursor [Pterocarpus angolensis]
Length = 260
Score = 208 bits (530), Expect = 1e-52
Identities = 112/253 (44%), Positives = 153/253 (60%), Gaps = 13/253 (5%)
Query: 75 LLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAF 134
+L+ DSLSF FPTF D + ++ GDA T N +QLTK D GNP + G F
Sbjct: 1 MLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQTKNNAVQLTKTDSNGNPVASTVGRILF 60
Query: 135 YGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPEGGGGGGFLGLF 193
VHL +K + RVA+F+++F+F + +G DG FF+A D P G GGG LGLF
Sbjct: 61 SAQVHLWEKSSSRVANFQSQFSFSLKSPLSNGADGIAFFIAPPDTTIP-SGSGGGLLGLF 119
Query: 194 DEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGA 249
TA NTS N ++AVEFD+F SN++DPN+PHIGID+N+IRS T+ W D+ G
Sbjct: 120 APGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYPHIGIDVNSIRSVKTVKW---DRRDGQ 176
Query: 250 IQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAE 309
N ++++ +R+L VV Y G + Y +D+ S L EWV +GFSA +GE +
Sbjct: 177 SLNVLVTFNPSTRNLDVVATYSDGTRY----EVSYEVDVRSVLPEWVRVGFSAASGEQYQ 232
Query: 310 THSILSWSFNSTL 322
TH++ SWSF STL
Sbjct: 233 THTLESWSFTSTL 245
>UniRef100_Q8GSD2 Lectin precursor [Pterocarpus angolensis]
Length = 260
Score = 206 bits (524), Expect = 7e-52
Identities = 111/253 (43%), Positives = 152/253 (59%), Gaps = 13/253 (5%)
Query: 75 LLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAF 134
+L+ DSLSF FPTF D + ++ GDA N +QLTK D GNP + G F
Sbjct: 1 MLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQIKNNAVQLTKTDSNGNPVASTVGRILF 60
Query: 135 YGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPEGGGGGGFLGLF 193
VHL +K + RVA+F+++F+F + +G DG FF+A D P G GGG LGLF
Sbjct: 61 SAQVHLWEKSSSRVANFQSQFSFSLKSPLSNGADGIAFFIAPPDTTIP-SGSGGGLLGLF 119
Query: 194 DEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGA 249
TA NTS N ++AVEFD+F SN++DPN+PHIGID+N+IRS T+ W D+ G
Sbjct: 120 APGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYPHIGIDVNSIRSVKTVKW---DRRDGQ 176
Query: 250 IQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAE 309
N ++++ +R+L VV Y G + Y +D+ S L EWV +GFSA +GE +
Sbjct: 177 SLNVLVTFNPSTRNLDVVATYSDGTRY----EVSYEVDVRSVLPEWVRVGFSAASGEQYQ 232
Query: 310 THSILSWSFNSTL 322
TH++ SWSF STL
Sbjct: 233 THTLESWSFTSTL 245
>UniRef100_Q8GRS1 Lectin precursor [Pterocarpus angolensis]
Length = 272
Score = 205 bits (521), Expect = 2e-51
Identities = 111/254 (43%), Positives = 152/254 (59%), Gaps = 13/254 (5%)
Query: 74 LLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISA 133
L+L+ DSLSF FPTF D + ++ GDA N +QLTK D GNP + G
Sbjct: 12 LMLLNKAYSQDSLSFGFPTFPSDQKNLIFQGDAQIKNNAVQLTKTDSNGNPVASTVGRIL 71
Query: 134 FYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPEGGGGGGFLGL 192
F VHL +K + RVA+F+++F+F + +G DG FF+A D P G GGG GL
Sbjct: 72 FSAQVHLWEKSSSRVANFQSQFSFSLKSPLSNGADGIAFFIAPPDTTIP-SGSGGGLPGL 130
Query: 193 FDEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQG 248
F TA NTS N ++AVEFD+F SN++DPN+PHIGID+N+IRS T+ W D+ G
Sbjct: 131 FAPGTAQNTSANQVIAVEFDTFYAQDSNTWDPNYPHIGIDVNSIRSVKTVKW---DRRDG 187
Query: 249 AIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFA 308
N ++++ +R+L VV Y G + Y +D+ S L EWV +GFSA +GE
Sbjct: 188 QSLNVLVTFNPSTRNLDVVATYSDGTRY----EVSYEVDVRSVLPEWVRVGFSAASGEQY 243
Query: 309 ETHSILSWSFNSTL 322
+TH++ SWSF STL
Sbjct: 244 QTHTLESWSFTSTL 257
>UniRef100_Q6GUG5 Lectin [Pterocarpus rotundifolius]
Length = 249
Score = 201 bits (511), Expect = 2e-50
Identities = 113/253 (44%), Positives = 149/253 (58%), Gaps = 13/253 (5%)
Query: 75 LLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSPHSSGISAF 134
+L+ SDS F F FD D R ++ GDA N LQLTK D GNP + G +
Sbjct: 1 MLLNKAYSSDSFPFGFFNFDQDERNLIYQGDARAQNNVLQLTKTDSNGNPVRSTVGRILY 60
Query: 135 YGAVHLSDKKTGRVADFRTEFTFVVNHSQPH-GDGFTFFLASLDFEFPEGGGGGGFLGLF 193
V L +K T RVA+F+++F+ ++ S + DG FF+A D P G GGG LGLF
Sbjct: 61 TAQVRLWEKSTNRVANFQSQFSLHLSSSLSNPADGIAFFIAPPDTTIP-SGSGGGLLGLF 119
Query: 194 DEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGA 249
TA NTS N ++AVEFD+F SN++DPN+ HIGID+N+IRSA T+ W D G
Sbjct: 120 APGTAQNTSANQVLAVEFDTFYAQDSNTWDPNYQHIGIDVNSIRSARTVRWERRD---GE 176
Query: 250 IQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAE 309
N ++Y+ +R L VV YP GQ + Y +D+ S L EWV +GFSA +GE +
Sbjct: 177 TLNVLVTYNPSTRTLDVVATYPDGQRY----EVSYEVDVRSVLPEWVRVGFSAASGEQYQ 232
Query: 310 THSILSWSFNSTL 322
THS+ SWSF STL
Sbjct: 233 THSLESWSFTSTL 245
>UniRef100_P93536 Bark lectin II precursor [Sophora japonica]
Length = 266
Score = 201 bits (510), Expect = 3e-50
Identities = 118/259 (45%), Positives = 156/259 (59%), Gaps = 14/259 (5%)
Query: 67 LATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTNGTLQLTKIDQYGNPSP 126
L + F L L+ SDSLSF+F F PD R ++L GDA+ +GTLQLTK D G
Sbjct: 1 LVFITFFLTLLNMVNSSDSLSFTFNNFGPDQRDLILQGDAHIPSGTLQLTKTDSSG---- 56
Query: 127 HSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNH--SQPHGDGFTFFLASLDFEFPEGG 184
G + +Y VHL D + GR+A F T F+FV+ + GDG FF+A + P
Sbjct: 57 --VGRALYYLPVHLWDSRRGRLASFETSFSFVITSQGTDDPGDGIAFFIAPPETTIPP-R 113
Query: 185 GGGGFLGLFDEETAYNTSKNSIVAVEFDSFSN-SFDPNFPHIGIDINTIRSAVTIPWSVN 243
GGFLGLF ETA N+S N +VAVEFD+F N +DP++ HIGID+N+I+S+ W
Sbjct: 114 SSGGFLGLFSPETALNSSLNPVVAVEFDTFINEDWDPSYWHIGIDVNSIKSSAAARW--- 170
Query: 244 DQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNID-AIKYPIDIESYLSEWVVIGFSA 302
++ G A ISY+S S+ L+VV YP +D + Y ID+ + L EWV IGFSA
Sbjct: 171 ERKSGRKFTAHISYNSSSKKLSVVSSYPNTNCLVRVDYTVSYDIDLTTVLPEWVRIGFSA 230
Query: 303 TTGEFAETHSILSWSFNST 321
+TG E HSILSWSF+S+
Sbjct: 231 STGYKIEEHSILSWSFSSS 249
>UniRef100_Q9FYU9 Lectin [Sophora flavescens]
Length = 284
Score = 196 bits (497), Expect = 9e-49
Identities = 113/262 (43%), Positives = 163/262 (62%), Gaps = 16/262 (6%)
Query: 67 LATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDAN-TTNGTLQLTKIDQYGNPS 125
+AT++ L+ + +DSLSF+F FDP+ +L GDA+ T+N LQLTK G P
Sbjct: 17 IATIV--LMSLRGVNSADSLSFTFSDFDPNGEDLLFQGDAHVTSNNILQLTKTSN-GVPL 73
Query: 126 PHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPH-GDGFTFFLASLDFEFPEGG 184
++ G + F +HL +K T R++ F + FTFV+ Q + DGF FF+A D PE G
Sbjct: 74 QNTVGRALFSTPIHLWEKSTNRLSSFESTFTFVLTSPQSNPADGFAFFIAPPDTTIPE-G 132
Query: 185 GGGGFLGLFDEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPW 240
GG LGLF E A N N +VAVEFD+F SNS+DPN+ HIGID+N I+S+ T+ W
Sbjct: 133 SDGGLLGLFSPENALNPKANQVVAVEFDTFYDKSSNSWDPNYVHIGIDVNQIKSSATVRW 192
Query: 241 SVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGF 300
D+ +G I A+I+Y++ +R+L+VV YP G + Y +D+ + L E+V +GF
Sbjct: 193 ---DRKEGVIGTARINYNAATRNLSVVSSYPGGSQDY---VVSYVVDLRTKLPEFVRVGF 246
Query: 301 SATTGEFAETHSILSWSFNSTL 322
SA+TG+ + HSI SW F+S+L
Sbjct: 247 SASTGQQYQVHSIRSWFFSSSL 268
>UniRef100_Q43374 Mannose/glucose-binding lectin precursor [Arachis hypogaea]
Length = 280
Score = 187 bits (476), Expect = 3e-46
Identities = 108/264 (40%), Positives = 154/264 (57%), Gaps = 15/264 (5%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPD-VRVILLNGDAN-TTNGTLQLTKIDQYGN 123
LL+ LLL+ DSLSFS+ F+ D R ++L GDA + + +QLTK+D G
Sbjct: 10 LLSIATIFLLLLNKAHSLDSLSFSYNNFEQDDERNLILQGDATFSASKGIQLTKVDDNGT 69
Query: 124 PSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPE 182
P+ + G V L +K T R+ +F+ +F+FV+ +G DG FF+A+ D E P+
Sbjct: 70 PAKSTVGRVLHSTQVRLWEKSTNRLTNFQAQFSFVIKSPIDNGADGIAFFIAAPDSEIPK 129
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTI 238
GG LGLFD TA N S N ++AVEFD+F SN +DPN+ HIGID+N+I+SA T
Sbjct: 130 NSAGGT-LGLFDPSTAQNPSANQVLAVEFDTFYAQDSNGWDPNYQHIGIDVNSIKSAATT 188
Query: 239 PWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVI 298
W ++ G N +SY + S++L V YP GQ + Y +D+ YL EW +
Sbjct: 189 KW---ERRNGQTLNVLVSYDANSKNLQVTASYPDGQRYQ----VSYNVDLRDYLPEWGSV 241
Query: 299 GFSATTGEFAETHSILSWSFNSTL 322
GFSA +G+ ++H + SWSF STL
Sbjct: 242 GFSAASGQQYQSHELQSWSFTSTL 265
>UniRef100_P93535 Seed lectin precursor [Sophora japonica]
Length = 292
Score = 183 bits (465), Expect = 5e-45
Identities = 112/263 (42%), Positives = 157/263 (59%), Gaps = 17/263 (6%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNP 124
L ++ F LLL+ ++ LSFSFP F + +LL GDA ++ G LQLT ++ G P
Sbjct: 21 LAISITFFLLLLNKVNSAEILSFSFPKFASNQEDLLLQGDALVSSKGELQLTTVEN-GVP 79
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNH--SQPHGDGFTFFLASLDFEFPE 182
+S+G + +Y VH+ DK TGRVA F T F+FVV + DG FFLA + +
Sbjct: 80 IWNSTGRALYYAPVHIWDKSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPPNNQIQ- 138
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSV 242
G GGG LGLF + YN+S I+AV+FD+ N++DPN HIGID+N+I S T+ W
Sbjct: 139 -GPGGGHLGLF-HSSGYNSSYQ-IIAVDFDTHINAWDPNTRHIGIDVNSINSTKTVTWGW 195
Query: 243 NDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSA 302
+ G + N ISY + + LTV + YP Q + + A +D++S L EWV +GF+A
Sbjct: 196 QN---GEVANVLISYQAATETLTVSLTYPSSQTSYILSA---AVDLKSILPEWVRVGFTA 249
Query: 303 TTG---EFAETHSILSWSFNSTL 322
TG ++ ETH +LSWSF STL
Sbjct: 250 ATGLTTQYVETHDVLSWSFTSTL 272
>UniRef100_Q39528 Agglutinin I precursor [Cladrastis lutea]
Length = 293
Score = 183 bits (464), Expect = 6e-45
Identities = 114/284 (40%), Positives = 164/284 (57%), Gaps = 22/284 (7%)
Query: 51 SQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDAN-TT 109
S N+ T ++ LLA + L+L+ SDSLSF+F F P+ ++ DA+ ++
Sbjct: 4 SNTNFLETKKPLSLPLLAFITIYLMLLHRVNSSDSLSFTFNNFPPNSEDLIFQKDASISS 63
Query: 110 NGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTF-VVNHSQPHGDG 168
N TL+LT+I G P+ S G + +Y V L DK TGR+A F+T F+F + + +Q GDG
Sbjct: 64 NETLELTRISSSGQPATSSVGRALYYTPVRLWDKSTGRLASFKTTFSFAITSPTQDPGDG 123
Query: 169 FTFFLASLDFEFPEGGGGGGFLGLFDEETAYNTSKN---------SIVAVEFDSFSN-SF 218
F FF+A D G GGG LGLF+ N+S N IVAVEFD++ N
Sbjct: 124 FAFFIAPPD---TTPGYGGGLLGLFNGFNLRNSSNNGVAVNNQSAQIVAVEFDTYINGQC 180
Query: 219 DPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTN 278
DP + H+GID+N+I S W + G A+ISY+ S+ LT V YP N+T
Sbjct: 181 DPKYRHVGIDVNSITSLAYTQWQWQN---GVKATAQISYNPASQKLTAVTSYP---NSTP 234
Query: 279 IDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
+ + ID+++ L EWV +GFSA+TG+ E +SIL+WSF+S+L
Sbjct: 235 L-TVSLDIDLQTVLPEWVRVGFSASTGQNVERNSILAWSFSSSL 277
>UniRef100_Q43377 Mannose/glucose-binding lectin precursor [Arachis hypogaea]
Length = 254
Score = 182 bits (463), Expect = 8e-45
Identities = 103/246 (41%), Positives = 147/246 (58%), Gaps = 15/246 (6%)
Query: 84 DSLSFSFPTFDPD-VRVILLNGDAN-TTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLS 141
DSLSFS+ F+ D R ++L GDA + + +QLTK+D G P+ + G V L
Sbjct: 2 DSLSFSYNNFEQDDERNLILQGDAKFSASKGIQLTKVDDNGTPAKSTVGRVLHSTQVRLW 61
Query: 142 DKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYN 200
+K T R+ +F+ +F+FV+ +G DG FF+A+ D E P+ GG LGLFD +TA N
Sbjct: 62 EKSTNRLTNFQAQFSFVIKSPIDNGADGIAFFIAAPDSEIPKNSAGGT-LGLFDPQTAQN 120
Query: 201 TSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKIS 256
S N ++AVEFD+F SN +DPN+ HIGID+N+I+SA T W D G N ++
Sbjct: 121 PSANQVLAVEFDTFYAQDSNGWDPNYQHIGIDVNSIKSAATTKWERRD---GQTLNVLVT 177
Query: 257 YSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSW 316
Y + S++L V YP GQ + Y +D+ YL EW +GFSA +G+ ++H + SW
Sbjct: 178 YDANSKNLQVTASYPDGQRYQ----LSYRVDLRDYLPEWGRVGFSAASGQQYQSHELQSW 233
Query: 317 SFNSTL 322
SF STL
Sbjct: 234 SFTSTL 239
>UniRef100_P93537 Bark lectin I precursor [Sophora japonica]
Length = 293
Score = 182 bits (463), Expect = 8e-45
Identities = 116/285 (40%), Positives = 162/285 (56%), Gaps = 24/285 (8%)
Query: 51 SQLNYQPTMANITSELLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDAN-TT 109
S N+ T + +LA + L+L+ SDSLSF++ F P+ ++L DA+ T+
Sbjct: 4 SNTNFLQTKKPLFLPILAFITIFLMLLNRVNSSDSLSFTYENFQPNPEDLILQRDASITS 63
Query: 110 NGTLQLTKIDQYGNPSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVV-NHSQPHGDG 168
N TL LT+ G P S G + +Y V L DK TGR+A F T F+FV+ + + GDG
Sbjct: 64 NETLLLTRTSN-GKPQKGSVGRALYYAPVRLWDKSTGRLASFETSFSFVITSPTTDPGDG 122
Query: 169 FTFFLASLDFEFPEGGGGGGFLGLFDEET----------AYNTSKNSIVAVEFDSFSN-S 217
FF+A D G GG LGLF+ T A++ S IVAVEFD++ N
Sbjct: 123 IAFFIAPPD---TTPGYTGGLLGLFNSSTVQSNSSDHGVAFHNSLPQIVAVEFDTYINGG 179
Query: 218 FDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTT 277
DPN+ H+GID+N+I+S T W+ + G A ISY+ VS+ LT V YP + T
Sbjct: 180 RDPNYRHVGIDVNSIKSVSTTKWTWRN---GVEATANISYNPVSQRLTAVSSYPNSEPIT 236
Query: 278 NIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSWSFNSTL 322
+ Y ID+++ L EWV +GFSA+TGE E +SILSWSF+S+L
Sbjct: 237 ----VHYDIDLKTVLPEWVRVGFSASTGENVEINSILSWSFSSSL 277
>UniRef100_P93538 Bark lectin precursor [Sophora japonica]
Length = 270
Score = 181 bits (459), Expect = 2e-44
Identities = 108/260 (41%), Positives = 157/260 (59%), Gaps = 17/260 (6%)
Query: 69 TLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNPSPH 127
++ F LLL+ ++ LSFSFP F + +LL GDA ++ G LQLT ++ G P +
Sbjct: 2 SITFFLLLLNKVNSAEILSFSFPKFVSNQEDLLLQGDALVSSEGELQLTTVEN-GVPVWN 60
Query: 128 SSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNH--SQPHGDGFTFFLASLDFEFPEGGG 185
S+G + +Y VH+ D TGRVA F T F+FVV + DG FFLA L+ + G
Sbjct: 61 STGRALYYAPVHIWDNSTGRVASFATSFSFVVKAPVASKSADGIAFFLAPLNNQIH--GA 118
Query: 186 GGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSVNDQ 245
GGG GLF+ + +S IVAVEFD+ +N++DPN HIGID+N+++S T+ W +
Sbjct: 119 GGGLYGLFNSSSY--SSSYQIVAVEFDTHTNAWDPNTRHIGIDVNSVKSTKTVTWGWEN- 175
Query: 246 PQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTG 305
G + N I+Y + + LTV + YP Q + + A +D++S L EWV +GF+ATTG
Sbjct: 176 --GEVANVLITYQAATEMLTVSLTYPSNQTSYILSA---AVDLKSILPEWVRVGFTATTG 230
Query: 306 ---EFAETHSILSWSFNSTL 322
++ ET+ +LSWSF STL
Sbjct: 231 LTTQYVETNDVLSWSFTSTL 250
>UniRef100_Q70DJ5 Putative lectin [Arachis hypogaea]
Length = 280
Score = 178 bits (452), Expect = 2e-43
Identities = 104/264 (39%), Positives = 151/264 (56%), Gaps = 15/264 (5%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDP-DVRVILLNGDAN-TTNGTLQLTKIDQYGN 123
LL+ LLL+ SLSF + F+ D R ++L GDA + + +QLTK+D G
Sbjct: 10 LLSIATIFLLLLNKAHSLGSLSFGYNNFEQGDERNLILQGDATFSASKGIQLTKVDDNGT 69
Query: 124 PSPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPE 182
P+ + G V L +K T R+ +F+ +F+FV+N +G DG FF+A+ D E P+
Sbjct: 70 PAKSTVGRVLHSTQVRLWEKSTNRLTNFQAQFSFVINSPIDNGADGIAFFIAAPDSEIPK 129
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTI 238
GG LGL D TA N S N ++AVEFD+F SN +DPN+ HIG D++ I+SA T
Sbjct: 130 NSAGGT-LGLSDPSTAQNPSANQVLAVEFDTFYAQDSNGWDPNYQHIGFDVDPIKSAATT 188
Query: 239 PWSVNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVI 298
W ++ G N +SY + S++L V YP GQ+ + Y +D+ YL EW +
Sbjct: 189 KW---ERRNGQTLNVLVSYDANSKNLQVTASYPDGQSYQ----VSYNVDLRDYLPEWGRV 241
Query: 299 GFSATTGEFAETHSILSWSFNSTL 322
GFSA +G+ ++H + SWSF STL
Sbjct: 242 GFSAASGQQYQSHGLQSWSFTSTL 265
>UniRef100_Q43376 Mannose/glucose-binding lectin precursor [Arachis hypogaea]
Length = 254
Score = 176 bits (447), Expect = 6e-43
Identities = 99/246 (40%), Positives = 147/246 (59%), Gaps = 15/246 (6%)
Query: 84 DSLSFSFPTFDPD-VRVILLNGDAN-TTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLS 141
DSLSFS+ F+ D R ++L GDA + + +QLTK+D G P+ + G V L
Sbjct: 2 DSLSFSYNKFEQDDERNLILQGDATFSASKGIQLTKVDANGTPAKSTVGRVLHSTQVRLW 61
Query: 142 DKKTGRVADFRTEFTFVVNHSQPHG-DGFTFFLASLDFEFPEGGGGGGFLGLFDEETAYN 200
+K T R+ +F+ +F+FV+ G DG FF+A+ D + P+ GG LGLFD +TA N
Sbjct: 62 EKSTNRLTNFQAQFSFVIKSPNDIGADGIAFFIAAPDSQIPKNSAGGT-LGLFDPQTAQN 120
Query: 201 TSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAKIS 256
S N ++AVEFD+F SN +DPN+ HIGID+N+I+SA T W ++ G N ++
Sbjct: 121 PSANQVLAVEFDTFYAQDSNGWDPNYQHIGIDVNSIKSAATTKW---ERRNGQTLNVLVT 177
Query: 257 YSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFAETHSILSW 316
Y + S++L V YP GQ + Y +D+ +L EW +GFSA++G+ ++H + SW
Sbjct: 178 YDANSKNLQVTASYPDGQRYQ----VSYVVDLRDHLPEWGRVGFSASSGQQYQSHELQSW 233
Query: 317 SFNSTL 322
SF S L
Sbjct: 234 SFTSNL 239
>UniRef100_Q39527 Lectin-related protein precursor [Cladrastis lutea]
Length = 290
Score = 175 bits (444), Expect = 1e-42
Identities = 111/263 (42%), Positives = 151/263 (57%), Gaps = 18/263 (6%)
Query: 66 LLATLIFSLLLITTTVKSDSLSFSFPTFDPDVRVILLNGDANTTN-GTLQLTKIDQYGNP 124
L ++ F LLL+ ++LSF+F F + +LL GDA ++ G LQLT+++ G P
Sbjct: 20 LAISITFYLLLLNKVNSEEALSFTFTKFVSNQDELLLQGDALVSSKGELQLTRVEN-GQP 78
Query: 125 SPHSSGISAFYGAVHLSDKKTGRVADFRTEFTFVVN--HSQPHGDGFTFFLASLDFEFPE 182
PHS G + + VH+ D TG VA F T FTFVV + DG FFLA D +
Sbjct: 79 IPHSVGRALYSDPVHIWDSSTGSVASFVTSFTFVVEAPNENKTADGIAFFLAPPDTQVQS 138
Query: 183 GGGGGGFLGLFDEETAYNTSKNSIVAVEFDSFSNSFDPNFPHIGIDINTIRSAVTIPWSV 242
GG FLGLF+ + YN+S N I+AVEFD+FSNS+DP HIGID+N+I S T W
Sbjct: 139 LGG---FLGLFNS-SVYNSS-NQILAVEFDTFSNSWDPTARHIGIDVNSIESTRTATWGW 193
Query: 243 NDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSA 302
+ G + I+Y + + L + YP Q + + A +D++S L EWV +GFSA
Sbjct: 194 RN---GEVAIVLITYVAPAETLIASLTYPSSQTSYILSA---AVDLKSILPEWVRVGFSA 247
Query: 303 TTGE---FAETHSILSWSFNSTL 322
TG + ETH +LSWSF STL
Sbjct: 248 ATGRSAGYVETHDVLSWSFTSTL 270
>UniRef100_P14894 Concanavalin A precursor [Canavalia gladiata]
Length = 290
Score = 171 bits (434), Expect = 2e-41
Identities = 102/261 (39%), Positives = 152/261 (58%), Gaps = 27/261 (10%)
Query: 78 TTTVKSDSLSFSFPTFDPDVRVILLNGDANT-TNGTLQLTKIDQYGNPSPHSSGISAFYG 136
++T ++++L F F F D + ++L GDA T T+G L+LT++ G+P S G + FY
Sbjct: 29 SSTHETNALHFMFNQFSKDQKDLILQGDATTGTDGNLELTRVSSNGSPQGSSVGRALFYA 88
Query: 137 AVHLSDKKTGRVADFRTEFTFVVNHSQPH-GDGFTFFLASLDFEFPEGGGGGGFLGLF-- 193
VH+ + + VA F FTF++ H DG FF++++D P G G LGLF
Sbjct: 89 PVHIWES-SAVVASFDATFTFLIKSPDSHPADGIAFFISNIDSSIPSGSTGR-LLGLFPD 146
Query: 194 ----------DEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIPWS 241
D AYN ++IVAVE D++ N+ DPN+PHIGIDI ++RS T W+
Sbjct: 147 ANVIRNSTTIDFNAAYNA--DTIVAVELDTYPNTDIGDPNYPHIGIDIKSVRSKKTAKWN 204
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
+ + G + A I Y+SV + L+ VV YP G + T + Y +D+++ L EWV +G S
Sbjct: 205 MQN---GKVGTAHIIYNSVGKRLSAVVSYPNGDSAT----VSYDVDLDNVLPEWVRVGLS 257
Query: 302 ATTGEFAETHSILSWSFNSTL 322
A+TG + ET++ILSWSF S L
Sbjct: 258 ASTGLYKETNTILSWSFTSKL 278
>UniRef100_O04672 Lectin [Canavalia brasiliensis]
Length = 290
Score = 170 bits (431), Expect = 4e-41
Identities = 101/261 (38%), Positives = 152/261 (57%), Gaps = 27/261 (10%)
Query: 78 TTTVKSDSLSFSFPTFDPDVRVILLNGDANT-TNGTLQLTKIDQYGNPSPHSSGISAFYG 136
++T ++++L F F F D + ++L GDA T T+G L+LT++ G+P S G + FY
Sbjct: 29 SSTHETNALHFMFNQFSKDQKDLILQGDATTGTDGNLELTRVSSNGSPQGSSVGRALFYA 88
Query: 137 AVHLSDKKTGRVADFRTEFTFVVNHSQPH-GDGFTFFLASLDFEFPEGGGGGGFLGLF-- 193
VH+ + + VA F FTF++ H DG FF++++D P G G LGLF
Sbjct: 89 PVHIWES-SAVVASFEATFTFLIKSPDSHPADGIAFFISNIDSSIPSGSTGR-LLGLFPD 146
Query: 194 ----------DEETAYNTSKNSIVAVEFDSFSNSF--DPNFPHIGIDINTIRSAVTIPWS 241
D AYN ++IVAVE D++ N+ DP++PHIGIDI ++RS T W+
Sbjct: 147 ANVIRNSTTIDFNAAYNA--DTIVAVELDTYPNTDIGDPSYPHIGIDIKSVRSKKTAKWN 204
Query: 242 VNDQPQGAIQNAKISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFS 301
+ + G + A I Y+SV + L+ VV YP G + T + Y +D+++ L EWV +G S
Sbjct: 205 MQN---GKVGTAHIIYNSVGKRLSAVVSYPNGDSAT----VSYDVDLDNVLPEWVRVGLS 257
Query: 302 ATTGEFAETHSILSWSFNSTL 322
A+TG + ET++ILSWSF S L
Sbjct: 258 ASTGLYKETNTILSWSFTSKL 278
>UniRef100_P22970 Anti-H(O) lectin I [Cytisus sessilifolius]
Length = 244
Score = 169 bits (428), Expect = 9e-41
Identities = 104/250 (41%), Positives = 147/250 (58%), Gaps = 22/250 (8%)
Query: 83 SDSLSFSFPTFDPDVRVILLNGDAN-TTNGTLQLTKIDQYGNPSPHSSGISAFYGAVHLS 141
+D LSF+F F P+ IL G+A+ +T G LQ+TK+ + P+ S G + + VH+
Sbjct: 2 NDHLSFNFDKFVPNQNNILFQGEASVSTTGVLQVTKVSK---PATRSIGRALYAAPVHIW 58
Query: 142 DKKTGRVADFRTEFTFVVNHSQPHG---DGFTFFLASLDFEFPEGGGGGGFLGLFDEETA 198
D TGRVA F T F+FVV DG TFFLA + + P G G F GLF+ ++
Sbjct: 59 DSTTGRVASFETSFSFVVKDEPEKSNGVDGLTFFLAPANSQIPSGSSAGLF-GLFN--SS 115
Query: 199 YNTSKNSIVAVEFDSF----SNSFDPNFPHIGIDINTIRSAVTIPWSVNDQPQGAIQNAK 254
N S N I+AVEFD++ N +DP+F HIG+D+N+I+S T+ W D G + N
Sbjct: 116 DNKSSNQIIAVEFDTYFGKTYNPWDPDFKHIGVDVNSIKSIKTVKW---DWRNGEVANVV 172
Query: 255 ISYSSVSRDLTVVVHYPPGQNTTNIDAIKYPIDIESYLSEWVVIGFSATTGEFA--ETHS 312
I+Y + ++ LTV + YP Q T+NI + +D+++ L EWV +GFSA G A ETH
Sbjct: 173 ITYRAPTKSLTVSLSYPSDQ-TSNI--VTASVDLKAILPEWVSVGFSAGVGNAAEFETHD 229
Query: 313 ILSWSFNSTL 322
+LSW F S L
Sbjct: 230 VLSWYFTSNL 239
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 557,334,665
Number of Sequences: 2790947
Number of extensions: 23913439
Number of successful extensions: 59986
Number of sequences better than 10.0: 390
Number of HSP's better than 10.0 without gapping: 239
Number of HSP's successfully gapped in prelim test: 151
Number of HSP's that attempted gapping in prelim test: 58599
Number of HSP's gapped (non-prelim): 428
length of query: 322
length of database: 848,049,833
effective HSP length: 127
effective length of query: 195
effective length of database: 493,599,564
effective search space: 96251914980
effective search space used: 96251914980
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0541.15


