

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0490.3
(1199 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
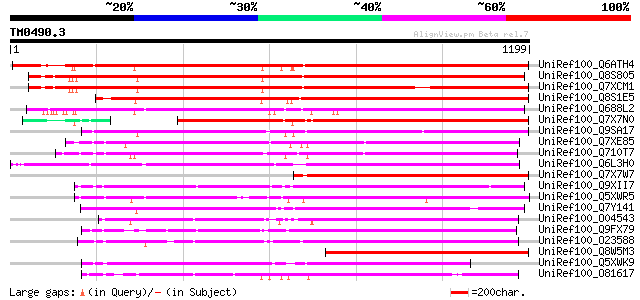
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q6ATH4 Putative polyprotein [Oryza sativa] 1101 0.0
UniRef100_Q8S805 Putative copia-type polyprotein [Oryza sativa] 1095 0.0
UniRef100_Q7XCM1 Putative gag-pol polyprotein [Oryza sativa] 1060 0.0
UniRef100_Q8S1E5 Putative gag/pol polyprotein [Oryza sativa] 1052 0.0
UniRef100_Q688L2 Putative polyprotein [Oryza sativa] 992 0.0
UniRef100_Q7X7N0 OSJNBa0035I04.5 protein [Oryza sativa] 830 0.0
UniRef100_Q9SA17 F28K20.17 protein [Arabidopsis thaliana] 789 0.0
UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa] 713 0.0
UniRef100_Q710T7 Gag-pol polyprotein [Populus deltoides] 663 0.0
UniRef100_Q6L3H0 Putative receptor kinase [Solanum demissum] 645 0.0
UniRef100_Q7X7W7 OSJNBa0061G20.2 protein [Oryza sativa] 638 0.0
UniRef100_Q9XII7 Putative retroelement pol polyprotein [Arabidop... 627 e-178
UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Sol... 625 e-177
UniRef100_Q7Y141 Putative polyprotein [Oryza sativa] 621 e-176
UniRef100_O04543 F20P5.25 protein [Arabidopsis thaliana] 615 e-174
UniRef100_Q9FX79 Putative retroelement polyprotein [Arabidopsis ... 611 e-173
UniRef100_O23588 Retrotransposon like protein [Arabidopsis thali... 602 e-170
UniRef100_Q8W5M3 Putative gag-pol polyprotein [Oryza sativa] 600 e-169
UniRef100_Q5XWK9 Gag-pol polyprotein-like [Solanum tuberosum] 598 e-169
UniRef100_O81617 F8M12.17 protein [Arabidopsis thaliana] 596 e-168
>UniRef100_Q6ATH4 Putative polyprotein [Oryza sativa]
Length = 1480
Score = 1101 bits (2847), Expect = 0.0
Identities = 601/1305 (46%), Positives = 786/1305 (60%), Gaps = 165/1305 (12%)
Query: 7 ASQTALHTSASRPPASVDSSQQRPTYR-SDQGHSNHRYGSGQNRNYQGGSGKPRKKGGSR 65
A+ ++ + +S A+ SSQ R S+ G N R G+N GG+G+ GG
Sbjct: 226 AAVSSASSCSSGTCATTTSSQPRGGGGGSNNGKHNQRRNGGRNNGGGGGNGRQNNSGGGS 285
Query: 66 YPGSPGSSGSSAASPPPWRPPPQASWNPWGWDPPPGWAPSPWGMPPCPYPTSQWSRPMGV 125
Y S + WRPP S
Sbjct: 286 YENSAST----------WRPPSNPS----------------------------------- 300
Query: 126 PQQPGVLGPRPQAYTATAS-----PAP--------------TDIAAAMHTMSLTPPDTPW 166
G+LGP PQA+TA AS P P + AA++ +S+ P +PW
Sbjct: 301 -NSAGLLGPYPQAHTAFASSYVSPPTPGWPPMPQVQPNWDQAGLVAALNQLSVNSP-SPW 358
Query: 167 YMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNHV 226
+DTGA+SH +++ G L + SH + VG+G IP+ G + +P AL ++
Sbjct: 359 VLDTGATSHMSSTDGILVTRLPNSHTF--ITVGNGHTIPVICRGTSFLPIGTTRFALKNI 416
Query: 227 LHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRSSP- 285
L P +++NL+ VRQ T DN S FD FGFSV D T +LRCNS GDLY + + P
Sbjct: 417 LVAPSLVRNLLFVRQFTRDNKCSFEFDEFGFSVKDLPTRRVILRCNSRGDLYTLPTTVPA 476
Query: 286 -----FAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPFS 340
F +S++WH+RLGHP+ +A+ L ++ C S ++ +C +C LGKH RL FS
Sbjct: 477 ITAHSFLAKSSTLWHHRLGHPSPAAVQTLHKLAILSCTRS-NNKLCHACHLGKHTRLSFS 535
Query: 341 SSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYETFTTL 400
S+ T PF+++H D+WTSPVLS +G +YY++ LDD T F WTFP+ KS V++
Sbjct: 536 KSSSSTSSPFELVHCDVWTSPVLSLSGFKYYLVVLDDFTHFCWTFPLRHKSDVHQHLLEF 595
Query: 401 ATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERKIRTI 460
+KTQFS I+C Q DNG ++ N + + + G++ R SCP+TS QNGKAER +RTI
Sbjct: 596 VAYVKTQFSLPIRCFQADNGTKFVNHATTSFFASRGIVLRLSCPYTSPQNGKAERVLRTI 655
Query: 461 NNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSHLRVFG 520
N IRTLL +S+PPS+W AL ATYLLN P +++N P QLL+++ P YS+L+VFG
Sbjct: 656 NKSIRTLLIQASMPPSYWAEALATATYLLNRRPSTSVRNSIPYQLLHNKLPDYSNLQVFG 715
Query: 521 CLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETRFPFA 580
CLC+P + T +KL PRS P VFLGY +H+G++C D+S R++ ISRHV+FDE FPFA
Sbjct: 716 CLCYPNLSAMTSHKLSPRSAPYVFLGYSASHKGFRCLDISTRRLYISRHVVFDEKTFPFA 775
Query: 581 DL------------------------------SLTPAPSYECFTEDLPPSLIHHWQTVSS 610
+ S+TP+P E D S +Q +
Sbjct: 776 AIPQDASSFDFLLQGFSIAVAPSSEVERPRFSSMTPSPEVEQPIPDDDTSGTELFQLLPG 835
Query: 611 RPPDPPVQP--SSPTDS-----------------------SPPLSALVSPTASPSPLPLP 645
+P SP D+ PP +++V P S P
Sbjct: 836 LRSSAAGRPLAGSPVDARLPGGCANDAANGSSSSNLSPVMDPPAASVVRPAPSEGPTTSL 895
Query: 646 PVP----------PAP-----PVR-----------------TMTTRSMHGISKPKKPFSL 673
P PAP P+R TM TRS G +P + F+
Sbjct: 896 ISPYRHTYLRRSQPAPTAIHRPIRASRAFHSATDQQQQTGHTMVTRSQTGHLRPIQRFTY 955
Query: 674 SVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIF 733
+ + D +SP+P N + AL+DPNW++A +E+ AL+ +NTW LVPRP NV+ WIF
Sbjct: 956 TATHD--VVSPVPSNYRSALADPNWRAATANEYKALVDNNTWRLVPRPPGANVVTGKWIF 1013
Query: 734 RHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQL 793
+HK S+G R+KAR V G SQ G+D DETFSPVVKPATI VLSIA SRSWPIHQL
Sbjct: 1014 KHKFHSDGTLARHKARWVVRGYSQQHGIDYDETFSPVVKPATIHVVLSIAASRSWPIHQL 1073
Query: 794 DVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSI 853
DV+NAFLHG+L ET+Y P GF DP P+ VC L+KSLYGLKQAPRAWYQRFA Y+ +
Sbjct: 1074 DVKNAFLHGNLEETVYYQQPSGFVDPSAPNAVCLLQKSLYGLKQAPRAWYQRFATYIRQL 1133
Query: 854 GFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLS 913
GF S S+ SLF+++ G +IAY+LLYVDDIIL ASS L A L SEFAM DLG L
Sbjct: 1134 GFTSSASNTSLFVYKDGDNIAYLLLYVDDIILTASSATLLHHITARLHSEFAMTDLGDLH 1193
Query: 914 YFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASL 973
+FLGI+VTR + GLFLSQ Y +++ RAGM+ C+ +ATPVD + KLS++ G P D +
Sbjct: 1194 FFLGISVTRSSDGLFLSQRQYVVDLLKRAGMSECHSTATPVDARSKLSATDGAPVSDPTE 1253
Query: 974 YRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPS 1033
YRSL GALQYLT TRP+++YAVQQVCL MH P H+ +KR+LRY++GTL GLHL +
Sbjct: 1254 YRSLVGALQYLTLTRPELAYAVQQVCLFMHDPREPHLALIKRILRYIKGTLHIGLHLGTT 1313
Query: 1034 PVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANV 1093
PV++L +Y+DADW GCPD+ RSTSGYCVFLGDNL+SWSSKRQ T+SRSSAEAEYR VA+
Sbjct: 1314 PVDSLTAYSDADWAGCPDSHRSTSGYCVFLGDNLVSWSSKRQTTVSRSSAEAEYRAVAHA 1373
Query: 1094 VSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVR 1153
++E CW+R LL ELH L+ AT+V+CDNVSA+Y++ NPVHH+RTKHIE+DIHFVREKV
Sbjct: 1374 IAECCWLRQLLSELHISLTSATVVYCDNVSAVYMAANPVHHRRTKHIEIDIHFVREKVAL 1433
Query: 1154 GQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPASTAG 1198
GQ RVLH+PS HQ ADI TKGLP LF +FRSSL V + PASTAG
Sbjct: 1434 GQVRVLHLPSSHQFADIMTKGLPVQLFTEFRSSLCVRDNPASTAG 1478
>UniRef100_Q8S805 Putative copia-type polyprotein [Oryza sativa]
Length = 1803
Score = 1095 bits (2831), Expect = 0.0
Identities = 579/1183 (48%), Positives = 753/1183 (62%), Gaps = 50/1183 (4%)
Query: 44 GSGQNRNYQGGSGKPRKKGGSRYPGSPGSSGSSAASPPPWRPPPQASWNPWGWDPPPGWA 103
G+G + +G+PR KG + G G + PP P G D P
Sbjct: 229 GTGSSSGTSDTTGQPRPKGRGKRRGRGGGA------PPGGAPSTPGGGAGAGHDGQPR-P 281
Query: 104 PSPWGMPPCPYPTSQWSRPMGVPQQPGVLGPRP-----QAYTAT--------ASPAPTDI 150
P+PWG P W P P GVLGPRP QA TA ASP
Sbjct: 282 PAPWGYNPWTGFVQAWPFPFRAPGA-GVLGPRPPFQAQQAMTAQHLLPALPPASPGVHST 340
Query: 151 AA-----------AMHTMSLTPPDTP-WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIV 198
A + + TPP W++DTGAS+H +++ G L L + + V
Sbjct: 341 GAWDNSALYSALQSAGVATTTPPSAADWFLDTGASAHMSSTPGILAHPRPLP-FSSCITV 399
Query: 199 GSGQGIPIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFS 258
G+G +P+ + T IPT L L++VL +P +IKNLISV++LT DNNVS+ FDP GFS
Sbjct: 400 GNGAKLPVTHTASTHIPTSSTDLHLHNVLVSPPLIKNLISVKKLTRDNNVSIEFDPTGFS 459
Query: 259 VSDFKTGMPLLRCNSLGDLYPVTRSSPFAGLASSV-----WHNRLGHPASSALNHLRNNK 313
+ D +T + LRC+S GDLYP+ SP A A+S WH RLGHP S++L+ + +
Sbjct: 460 IKDLQTQVVKLRCDSPGDLYPLRLPSPHALSATSSPSVEHWHLRLGHPGSASLSKVLGSF 519
Query: 314 LIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVL 373
C S C +C +G +VRLPF SS+ TL PF ++H+D+WTSP+ S +G++YYV+
Sbjct: 520 DFQCNKSAPHH-CSACHVGTNVRLPFHSSSSQTLFPFQLVHTDVWTSPIYSNSGYKYYVV 578
Query: 374 FLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCD 433
FLDD T ++WTFP+ KS+V+ T + TQF + LQ DNG+EYD+ +
Sbjct: 579 FLDDFTHYIWTFPVRNKSEVFHTVRSFFAYAHTQFGLPVLALQTDNGKEYDSYALRSLLS 638
Query: 434 ANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILP 493
+G + R SCP++S QNGKAER +RTIN+ +RT+L HS+ P SFW ALQ A +L+N P
Sbjct: 639 LHGAVLRLSCPYSSQQNGKAERILRTINDCVRTMLVHSAAPLSFWAEALQTAMHLINRRP 698
Query: 494 RKTLQNDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRG 553
+ + P QLL P+Y HLRVFGCLC+P + +KL PRS CVF+GYP +HRG
Sbjct: 699 CRATGSLKPYQLLLGAPPTYDHLRVFGCLCYPNTIATAPHKLSPRSLACVFIGYPADHRG 758
Query: 554 YKCYDLSHRKILISRHVIFDETRFPFADL-----SLTPAPSYECFTEDLPPSLIHHWQT- 607
Y+CYD+ R++ SRHV F E FPF D S P P + T L P+ H T
Sbjct: 759 YRCYDMVSRRVFTSRHVTFVEDVFPFRDAPSPRPSAPPPPDHGDDTIVLLPAPAQHVVTP 818
Query: 608 VSSRPPDPPVQPSSPTDSSPPLSALVSPTASP-SPLPLPPVPPAPPVRTMTTRSMHGISK 666
V + P P SP S+P +A A P SP P +PP MTTR+ GISK
Sbjct: 819 VGTAPAHDAASPPSPASSTPSSAAPAHDVAPPPSPETSSPASASPPRHAMTTRARAGISK 878
Query: 667 PKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNV 726
P ++++ + ++SP P + + AL DPNW++AMQ+EF+AL+ + TW LVPRP +
Sbjct: 879 PNPRYAMTAT---STLSPTPSSVRVALRDPNWRAAMQAEFDALLANRTWTLVPRPPGARI 935
Query: 727 IRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSR 786
I W+F+ K ++G ++YKAR V G +Q GVD ETFSPVVKPATIRTVL++ S+
Sbjct: 936 ITGKWVFKTKLHADGSLDKYKARWVVRGFNQRPGVDFGETFSPVVKPATIRTVLTLISSK 995
Query: 787 SWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRF 846
WP HQLDV NAFLHG L E + P GF D P VC L +SLYGL+QAPRAW++RF
Sbjct: 996 QWPAHQLDVSNAFLHGHLQERVLCQQPTGFEDAARPADVCLLSRSLYGLRQAPRAWFKRF 1055
Query: 847 ADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAM 906
AD+ +S+GF S +D SLF+ R+GSD AY+LLYVDD+IL ASS L + + L +EF +
Sbjct: 1056 ADHATSLGFVQSRADPSLFVLRRGSDTAYLLLYVDDMILSASSSSLLQRIIDRLQAEFKV 1115
Query: 907 KDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGT 966
KD+GPL YFLGI V R A G LSQS YAT+++ RAGMA+C ATP D K KLSS G
Sbjct: 1116 KDMGPLKYFLGIEVQRTADGFVLSQSKYATDVLERAGMANCKAVATPADAKPKLSSDEGP 1175
Query: 967 PCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTY 1026
+D+S YRS+ GALQYLT TRPDI+YAVQQVCLHMHAP H+ LKR+LRY++GT +
Sbjct: 1176 LFQDSSWYRSIAGALQYLTLTRPDIAYAVQQVCLHMHAPREAHVTLLKRILRYIKGTAAF 1235
Query: 1027 GLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAE 1086
GLHL S TL +++DADW GCPDTRRSTSG+C+FLGD+LISWSSKRQ T+SRSSAEAE
Sbjct: 1236 GLHLRASTSPTLTAFSDADWAGCPDTRRSTSGFCIFLGDSLISWSSKRQTTVSRSSAEAE 1295
Query: 1087 YRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHF 1146
YRGVAN V+E W+R LL ELH + QAT+ +CDN+S++Y+S NPVHH+RTKHIE+DIHF
Sbjct: 1296 YRGVANAVAECTWLRQLLGELHCRVPQATIAYCDNISSVYMSKNPVHHKRTKHIELDIHF 1355
Query: 1147 VREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV 1189
VREKV G+ RVL +PS HQ AD+FTKGLP +F++FR+SL V
Sbjct: 1356 VREKVALGELRVLPIPSAHQFADVFTKGLPSSMFNEFRASLCV 1398
>UniRef100_Q7XCM1 Putative gag-pol polyprotein [Oryza sativa]
Length = 1417
Score = 1060 bits (2742), Expect = 0.0
Identities = 569/1192 (47%), Positives = 742/1192 (61%), Gaps = 82/1192 (6%)
Query: 44 GSGQNRNYQGGSGKPRKKGGSRYPGSPGSSGSSAASPPPWRPPPQASWNPWGWDPPPGWA 103
G+G + +G+PR KG + G G + PP P G D P
Sbjct: 229 GTGSSSGTSDTTGQPRPKGRGKRRGRGGGA------PPGGAPSTPGGGAGAGHDGQPR-P 281
Query: 104 PSPWGMPPCPYPTSQWSRPMGVPQQPGVLGPRP-----QAYTAT--------ASPAPTDI 150
P+PWG P W P P GVLGPRP QA TA ASP
Sbjct: 282 PAPWGYNPWTGFVQAWPFPFRAPGA-GVLGPRPPFQAQQAMTAQHLLPALPPASPGVHST 340
Query: 151 AA-----------AMHTMSLTPPDTP-WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIV 198
A + + TPP W++DTGAS+H +++ G L L + + V
Sbjct: 341 GAWDNSALYSALQSAGVATTTPPSAADWFLDTGASAHMSSTPGILAHPRPLP-FSSCITV 399
Query: 199 GSGQGIPIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFS 258
G+G +P+ + T IPT L L++VL +P +IKNLISV++LT DNNVS+ FDP GFS
Sbjct: 400 GNGAKLPVTHTASTHIPTSSTDLHLHNVLVSPPLIKNLISVKKLTRDNNVSIEFDPTGFS 459
Query: 259 VSDFKTGMPLLRCNSLGDLYPVTRSSPFAGLASSV-----WHNRLGHPASSALNHLRNNK 313
+ D +T + LRC+S GDLYP+ SP A A+S WH RLGHP S++L+ + +
Sbjct: 460 IKDLQTQVVKLRCDSPGDLYPLRLPSPHALSATSSPSVEHWHLRLGHPGSASLSKVLGSF 519
Query: 314 LIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVL 373
C S C +C +G +VRLPF SS+ TL PF ++H+D+WTSP+ S +G++YYV+
Sbjct: 520 DFQCNKSAPHH-CSACHVGTNVRLPFHSSSSQTLFPFQLVHTDVWTSPIYSNSGYKYYVV 578
Query: 374 FLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCD 433
FLDD T ++WTFP+ KS+V+ T + TQF + LQ DNG+EYD+ +
Sbjct: 579 FLDDFTHYIWTFPVRNKSEVFHTVRSFFAYAHTQFGLPVLALQTDNGKEYDSYALRSLLS 638
Query: 434 ANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILP 493
+G + R SCP++S QNGKAER +RTIN+ +RT+L HS+ P SFW ALQ AT+L+N P
Sbjct: 639 LHGAVLRLSCPYSSQQNGKAERILRTINDYVRTMLVHSAAPLSFWAEALQTATHLINRRP 698
Query: 494 RKTLQNDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRG 553
+ + +P QLL P+Y HLRVFGCLC+P + +KL PRS CVF+GYP +HRG
Sbjct: 699 CRATGSLTPYQLLLGAPPTYDHLRVFGCLCYPNTIATAPHKLSPRSLACVFIGYPADHRG 758
Query: 554 YKCYDLSHRKILISRHVIFDETRFPFADL-----SLTPAPSYECFTEDLPPSLIHHWQT- 607
Y+CYD+ R++ SRHV F E FPF D S P P + T L P+ H T
Sbjct: 759 YRCYDMVSRRVFTSRHVTFVEDVFPFRDAPSPRPSAPPPPDHGDDTIVLLPAPAQHVVTP 818
Query: 608 VSSRPPDPPVQPSSPTDSSPPLSALVSPTASP-SPLPLPPVPPAPPVRTMTTRSMHGISK 666
V + P P SP S+P +A A P SP P +PP MTTR+ GISK
Sbjct: 819 VGTAPAHDAASPPSPASSTPSSAAPAHDVAPPPSPETSSPASASPPRHAMTTRARAGISK 878
Query: 667 PKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNV 726
P ++++ + ++SP P + + AL DPNW++AMQ+EF+AL+ + TW LVPRP +
Sbjct: 879 PNPRYAMTAT---STLSPTPSSVRAALRDPNWRAAMQAEFDALLANRTWTLVPRPPGARI 935
Query: 727 IRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSR 786
I W+F+ K ++G ++YKAR V G +Q GVD ETFSPVVKPATIRTVL++ S+
Sbjct: 936 ITGKWVFKTKLHADGSLDKYKARWVVRGFNQRPGVDFGETFSPVVKPATIRTVLTLISSK 995
Query: 787 SWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRF 846
WP HQLDV NAFLHG L E + P GF D P VC L +SLYGL+QAPRAW++RF
Sbjct: 996 QWPAHQLDVSNAFLHGHLQERVLCQQPTGFEDAARPADVCLLSRSLYGLRQAPRAWFKRF 1055
Query: 847 ADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAM 906
AD+ +S+GF S +D SLF+ R+GSD AY+LLYVDD+IL ASS L + + L +EF +
Sbjct: 1056 ADHATSLGFVQSRADPSLFVLRRGSDTAYLLLYVDDMILSASSSSLLQRIIDRLQAEFKV 1115
Query: 907 KDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGT 966
KD+GPL YFLGI V R A G LSQS YAT
Sbjct: 1116 KDMGPLKYFLGIEVQRTADGFVLSQSKYAT------------------------------ 1145
Query: 967 PCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTY 1026
+D+S YRS+ GALQYLT TRPDI+YAVQQVCLHMHAP H+ LKR+LRY++GT +
Sbjct: 1146 --DDSSWYRSIAGALQYLTLTRPDIAYAVQQVCLHMHAPREAHVTLLKRILRYIKGTAAF 1203
Query: 1027 GLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAE 1086
GLHL S TL +++DADW GCPDTRRSTSG+C+FLGD+LISWSSKRQ T+SRSSAEAE
Sbjct: 1204 GLHLRASTSPTLTAFSDADWAGCPDTRRSTSGFCIFLGDSLISWSSKRQTTVSRSSAEAE 1263
Query: 1087 YRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHF 1146
YRGVAN V+E W+R LL ELH + QAT+ +CDN+S++Y+S NPVHH+RTKHIE+DIHF
Sbjct: 1264 YRGVANAVAECTWLRQLLGELHCRVPQATIAYCDNISSVYMSKNPVHHKRTKHIELDIHF 1323
Query: 1147 VREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPASTAG 1198
VREKV G+ RVL +PS HQ AD+FTKGLP +F++FR+SL VG+ A+TAG
Sbjct: 1324 VREKVALGELRVLPIPSAHQFADVFTKGLPSSMFNEFRASLCVGDTTAATAG 1375
>UniRef100_Q8S1E5 Putative gag/pol polyprotein [Oryza sativa]
Length = 1090
Score = 1052 bits (2721), Expect = 0.0
Identities = 551/1056 (52%), Positives = 692/1056 (65%), Gaps = 74/1056 (7%)
Query: 199 GSGQGIPIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFS 258
G G +P G GY + P ++NL+SVRQ T DN S+ FD FGFS
Sbjct: 39 GGGGALPGGGGGYQGVANP---------------VRNLLSVRQFTRDNKCSIEFDEFGFS 83
Query: 259 VSDFKTGMPLLRCNSLGDLYPVTRSSPFAGL------ASSVWHNRLGHPASSALNHLRNN 312
V D +T +LRCNS G+LY + ++P + +S++WH RLGHP +A++ LRN
Sbjct: 84 VKDLQTRRVILRCNSRGELYTLPAATPSSAAHGLLATSSTLWHCRLGHPGPAAIHGLRNI 143
Query: 313 KLIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYV 372
I C +S+C +C LGKH RLPF +S+ T PF+++H D+WTSPV+ST+G +YY+
Sbjct: 144 ASISCNKI-DTSLCHACQLGKHTRLPFHNSSSRTSVPFELVHCDVWTSPVMSTSGFKYYL 202
Query: 373 LFLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYC 432
+ LDD + F WTF + KS V+ + TQF +K Q DNGRE+ N + +
Sbjct: 203 VVLDDFSHFCWTFLLRLKSDVHRHIVEFVEYVSTQFGLPLKSFQADNGREFVNTAITTFL 262
Query: 433 DANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNIL 492
+ G R SCP+TS QNGKAER +RTINN IRTLL +S+PPS+W AL ATYLLN
Sbjct: 263 ASRGTQLRLSCPYTSPQNGKAERMLRTINNSIRTLLIQASMPPSYWAEALATATYLLNRR 322
Query: 493 PRKTLQNDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHR 552
P ++ P QLL+ P +SHLRVFGCLC+P + T +KL PRST CVFLGYP +H+
Sbjct: 323 PSSSIHQSLPFQLLHRTIPDFSHLRVFGCLCYPNLSATTPHKLSPRSTACVFLGYPTSHK 382
Query: 553 GYKCYDLSHRKILISRHVIFDETRFPFA----------------------------DLSL 584
GY+C DLS +I+ISRHV+FDE++FPFA L
Sbjct: 383 GYRCLDLSTHRIIISRHVVFDESQFPFAATPPAASSFDFLLQGLSPADAPSLEVEQPRPL 442
Query: 585 TPAPSYECFTEDLP-PS--LIHHWQTVSSRPPD--PPVQPSSPTDSSPPLSAL-VSPTAS 638
T APS E LP PS L TV+S P P+ +S D++PP SA S S
Sbjct: 443 TVAPSTEVEQPYLPLPSRRLSAGTVTVASEAPSAGAPLVGTSSADATPPGSATRASTIVS 502
Query: 639 P-----SPLPLPPVPP-----------APPVRTMTTRSMHGISKPKKPFSLSVSIDDPSI 682
P + P+ VPP AP +M TRS G +P L+ + +
Sbjct: 503 PFRHVYTRRPVTTVPPSSSTAVTNAVAAPQPHSMVTRSQSGSLRPVD--RLTYTATQAAA 560
Query: 683 SPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGL 742
SP+P N AL+DPNW++AM E+ L+ + TW LV RP N+ WIF+HK S+G
Sbjct: 561 SPVPANYHSALADPNWRAAMADEYKELVDNGTWRLVSRPPRANIATGKWIFKHKFHSDGS 620
Query: 743 FERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHG 802
RYKAR V G SQ G+D DETFSPVVK ATIR VLSIA SR+WPIHQLDV+NAFLHG
Sbjct: 621 LARYKARWVVRGYSQQHGIDYDETFSPVVKLATIRVVLSIAASRAWPIHQLDVKNAFLHG 680
Query: 803 DLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDH 862
L ET+Y P GF DP PD VC L+KSLYGLKQAPRAWYQRFA Y+ +GF S SD
Sbjct: 681 HLKETVYCQQPSGFVDPTAPDAVCLLQKSLYGLKQAPRAWYQRFATYIRQMGFMPSASDT 740
Query: 863 SLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTR 922
SLF+++ G IAY+LLYVDDIIL AS+ L + A L SEFAM DLG L +FLGI+V R
Sbjct: 741 SLFVYKDGDRIAYLLLYVDDIILTASTTTLLQQLTARLHSEFAMTDLGDLHFFLGISVKR 800
Query: 923 HAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQ 982
GLFLSQ YA +++ RAGMA C+ ++TPVDT KLS++ G P D S YRS+ GALQ
Sbjct: 801 SPDGLFLSQRQYAVDLLQRAGMAECHSTSTPVDTHAKLSATDGLPVADPSAYRSIAGALQ 860
Query: 983 YLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYT 1042
YLT TRPD++YAVQQVCL MH P H+ +KR+LRYV+G+L+ GLH+ P+++L +Y+
Sbjct: 861 YLTLTRPDLAYAVQQVCLFMHDPREPHLALVKRILRYVKGSLSIGLHIGSGPIQSLTAYS 920
Query: 1043 DADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRN 1102
DADW GCP++RRSTSGYCV+LGDNL+SWSSKRQ T+SRSSAEAEYR VA+ V+E CW+R
Sbjct: 921 DADWAGCPNSRRSTSGYCVYLGDNLVSWSSKRQTTVSRSSAEAEYRAVAHAVAECCWLRQ 980
Query: 1103 LLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVP 1162
LL ELH P++ AT+V+CDNVSA+Y++ NPVHH+RTKHIE+DIHFVREKV GQ RVL+VP
Sbjct: 981 LLQELHVPIASATIVYCDNVSAVYMTANPVHHRRTKHIEIDIHFVREKVALGQVRVLYVP 1040
Query: 1163 SRHQIADIFTKGLPRLLFDDFRSSLSVGEPPASTAG 1198
S HQ ADI TKGLP LF DFRSSL V PPA+TAG
Sbjct: 1041 SSHQFADIMTKGLPVQLFTDFRSSLCVRAPPAATAG 1076
>UniRef100_Q688L2 Putative polyprotein [Oryza sativa]
Length = 1679
Score = 992 bits (2565), Expect = 0.0
Identities = 588/1313 (44%), Positives = 752/1313 (56%), Gaps = 170/1313 (12%)
Query: 39 SNHRYGSGQNRNYQGGSGKP----RKKGGSRYPGSPGSSGSSA-----------ASPPPW 83
+N +G+G QGG GK R++GG+ G G G ++ A W
Sbjct: 374 ANPAHGTGYGAGNQGGGGKGNSNRRRRGGNGGGGGGGGGGGNSNQGGGSTVPAQAQGAQW 433
Query: 84 R-----PPPQASWNPWG-----WDPP-------PGWAPSPW------GMPPCPYPTSQWS 120
R P PQ NPW W P P APSP+ G P P +
Sbjct: 434 RAGSQWPSPQ---NPWAGTIHMWPGPLGRGVLGPRPAPSPFAGAALAGPSGGPLPGQVYP 490
Query: 121 RPM------GVPQQP-----GVLGP-RPQAYTATASP------APTD-----IAAAMHTM 157
P+ G+ QQ G GP +P A P AP+ +A + +T
Sbjct: 491 APVPYQTAAGLGQQAQLGGVGQFGPSQPPTSVALQQPLGWFGGAPSQWDQASLAGSFNTT 550
Query: 158 SLTPPDT-PWYMDTGASSHTAASQGTLT-SYSNLSHLNQKLIVGSGQGIPIQGSGYTSIP 215
+L P T WYMDTGA++H + G L+ S+ + ++VG+G IP+ G++ I
Sbjct: 551 TLHQPATNDWYMDTGATAHMTSDTGILSLSHPPNPNSPSHIVVGNGSTIPVTSIGHSKIC 610
Query: 216 TPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLG 275
P+ L +L +P IIKNLISVR+ DN SV FDPFGFSV D +T + R NS G
Sbjct: 611 HPNCSFTLRDILCSPAIIKNLISVRRFVIDNWCSVEFDPFGFSVKDLRTRTVIARFNSSG 670
Query: 276 DLYPVTRSSP-----FAGLASS----VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVC 326
LY + + P A LA+S +WH RLGH ALN L ++ + VC
Sbjct: 671 PLYSLHHALPPPPAATALLANSSTDLLWHRRLGHLGHDALNRLA--AVVPMTRGDLTGVC 728
Query: 327 DSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFP 386
+C LG+HVRLPF+SS F++ H DLWTSPV+S +G +YY++ LDD + ++WTFP
Sbjct: 729 HACQLGRHVRLPFASSTSRASTNFELFHCDLWTSPVVSASGFKYYLVILDDCSHYVWTFP 788
Query: 387 ISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHT 446
+ KS + T + +KTQF+ NI+ +QCDNGRE+DN + + NG+ R SCPHT
Sbjct: 789 LRFKSDTFTTLSHFFAYVKTQFATNIRSIQCDNGREFDNSAARTFFLTNGVHLRMSCPHT 848
Query: 447 SSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLL 506
S QNGKAER +R++NN++R++L + +P SFW AL AT+L+N P KTL +P L
Sbjct: 849 SPQNGKAERILRSLNNIVRSMLFQAKLPGSFWVEALHTATHLINRHPTKTLDRHTPHFAL 908
Query: 507 YHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILI 566
Y PSYSHLRVFGC C+P + T +KL PRST CVFLGYP+ H+GY+C+D +++I
Sbjct: 909 YGTHPSYSHLRVFGCKCYPNLSATTPHKLAPRSTMCVFLGYPLYHKGYRCFDPLSNRVII 968
Query: 567 SRHVIFDETRFPFADLSLTPAPSYEC-FTEDLP------------PSLIHHWQTVSS--- 610
SRHV+FDE FPF +L+ + + + F ED P++ QT SS
Sbjct: 969 SRHVVFDEHSFPFTELTNGVSNATDLDFLEDFTAPAQAPIGATRRPAVAPTTQTASSPMV 1028
Query: 611 ------------RPPDPPVQPSSPTDSSPPLS---ALVSPTASPSPLPLPPVPPAPPVRT 655
RP P PSSP P S AL+ P AS SP P P PAP T
Sbjct: 1029 HGLERPPPCSPTRPVSTPGGPSSPDSRLGPPSPTPALIGP-ASTSPGP-PSAGPAPSAST 1086
Query: 656 MTTRS-MHGISKPKKPFSLSVSIDDPSISPLPHNPKQALS-------------------- 694
S +++P P L +D P + P PH P++ S
Sbjct: 1087 CPASSTWETVARPPTPPGLP-RLDGPHLPPAPHVPRRLRSVRATGAPTPLSGLEISPVVN 1145
Query: 695 ----DPNWKSAMQSEFNAL-IRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFER---- 745
KS + L + + LVP+ + +W +++ N L
Sbjct: 1146 DHVMTTRAKSGHHKPVHRLNLHAAPLSLVPKTYRAALADPLWRAAMEEEYNALLANRTWD 1205
Query: 746 ----------------YKARLVGDGR-------------SQIAGVDCDETFSPVVKPATI 776
+K + DG +Q GVD DETFSPVVKPAT+
Sbjct: 1206 LVPRPAGVNVVTGKWIFKHKFHADGSLDRYKARWVLRGFTQRPGVDFDETFSPVVKPATV 1265
Query: 777 RTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLK 836
RTVLS+A+SR WP+HQLDV+NAFLHG L ET+Y P GF D PD VC L KSLYGLK
Sbjct: 1266 RTVLSLAVSRDWPVHQLDVKNAFLHGTLQETVYCTQPPGFVDSAKPDMVCCLNKSLYGLK 1325
Query: 837 QAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSF 896
QAPRAWY RF ++ SIGF + SD SLFI +G+D Y+LLYVDDI+L ASS L
Sbjct: 1326 QAPRAWYSRFTTFLQSIGFVEAKSDTSLFILHRGNDTVYLLLYVDDIVLTASSRTLLHWT 1385
Query: 897 MALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDT 956
++ L EF+MKDLG L +FLG++VTR++ GL LSQ Y +I+ RAGMA C P TPVDT
Sbjct: 1386 ISALQGEFSMKDLGALHHFLGVSVTRNSAGLVLSQRQYCIDILERAGMADCKPCNTPVDT 1445
Query: 957 KQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRV 1016
KLSSS G P D + +RSL GALQYLTFTRPDISYAVQQVCLHMH P H+ ALKR+
Sbjct: 1446 TAKLSSSDGPPVADPTDFRSLAGALQYLTFTRPDISYAVQQVCLHMHDPREPHLAALKRI 1505
Query: 1017 LRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQP 1076
L Y+RG++ GLH+ S L Y+DADW GCPDTRRSTSGY VFLGDNL+SWSSKRQ
Sbjct: 1506 LHYIRGSVDLGLHIQRSSACDLAVYSDADWAGCPDTRRSTSGYAVFLGDNLVSWSSKRQH 1565
Query: 1077 TLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQR 1136
T+SRSSAEAEYR VAN V+E W+R LL ELH P S+ATLV+CDNVSA+YLS NPV HQR
Sbjct: 1566 TVSRSSAEAEYRAVANAVAEVTWLRQLLQELHSPPSRATLVYCDNVSAVYLSSNPVQHQR 1625
Query: 1137 TKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV 1189
TKH+E+D+HFVRE+V G RVLHVP+ Q ADIFTKGLP +F +FRSSL+V
Sbjct: 1626 TKHVEIDLHFVRERVAVGAVRVLHVPTTSQYADIFTKGLPTPVFTEFRSSLNV 1678
>UniRef100_Q7X7N0 OSJNBa0035I04.5 protein [Oryza sativa]
Length = 1246
Score = 830 bits (2144), Expect = 0.0
Identities = 444/840 (52%), Positives = 541/840 (63%), Gaps = 41/840 (4%)
Query: 388 SKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTS 447
S KS V+ TQF +KCLQ DNG E+ N + + G + R SCP+TS
Sbjct: 415 SLKSDVHRHIVEFVEYAHTQFGHRLKCLQADNGTEFVNHNTTSFLAGRGSLLRLSCPYTS 474
Query: 448 SQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLY 507
QNGKAER IRT+NN IRTLL +S+PPS+W L ATYLLN P ++ N P QLL+
Sbjct: 475 PQNGKAERMIRTLNNSIRTLLLQASMPPSYWAEGLATATYLLNRRPSSSVNNSIPFQLLH 534
Query: 508 HRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILIS 567
+ P+YS LRVFGCLC+P + +KL P S CVFLGYP +H+GY C ++S R+I+IS
Sbjct: 535 RKIPNYSMLRVFGCLCYPNLSATAAHKLAPYSAACVFLGYPSSHKGYCCLNISTRRIIIS 594
Query: 568 RHVIFDETRFPFADLSLTPAPSYECFTEDLP-PSLIHHWQTVSSRPPDPPVQPSSPTDSS 626
HVIFDET+FPF+ A S + +D P PS+I +P P +
Sbjct: 595 CHVIFDETQFPFSG-DPVDASSLDFLLQDAPAPSVIAPSLAGVEQPHLPHAPFPVNVEQR 653
Query: 627 PPLSALVSPTASPSPLPLPPVPPA-----------------------------PPVRTMT 657
P A P+ LP P A P V
Sbjct: 654 LPTGA---PSTKDEHLPYYVQPAAHCGQDDGKFHTAGCHVSIFLKQGTIVTSYPAVHVFF 710
Query: 658 TRSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWEL 717
T S HG + P P + DD + S H + P +AM EF ALI + TW L
Sbjct: 711 TTSRHGGAIPV-PLPTTTGADDSAPS---HRCGCTHTSP---AAMAEEFKALIDNGTWRL 763
Query: 718 VPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIR 777
VPRP NV+ WIF+HK S+G R+KAR V G SQ G+D DETFSPVVKP+TIR
Sbjct: 764 VPRPPGANVVTGKWIFKHKFHSDGSLARHKARWVVHGYSQQHGIDYDETFSPVVKPSTIR 823
Query: 778 TVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQ 837
VLSIA SRSWPIHQLDV+NAFLHG L ET+Y GF DP PD VC L++SLYGLKQ
Sbjct: 824 VVLSIATSRSWPIHQLDVKNAFLHGTLDETVYCQQSSGFIDPAAPDAVCLLQRSLYGLKQ 883
Query: 838 APRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFM 897
APRAWYQRFA Y+ +GF S SD SLFI R G +AY+LLYVDDIIL+ASS +L
Sbjct: 884 APRAWYQRFATYIRQLGFTPSASDTSLFILRDGDRLAYLLLYVDDIILMASSAELLCHIT 943
Query: 898 ALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTK 957
A L +EFAM DLG L +FLGI+V R LFLSQ YA +++ RAGM+ +P+ATPVD +
Sbjct: 944 ARLHTEFAMTDLGDLHFFLGISVRRTPDDLFLSQRQYAVDLLQRAGMSEYHPTATPVDAR 1003
Query: 958 QKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVL 1017
KLS+S G P D + YRSL ALQYLT TRP+I+YAVQQVC MH PH H+ +KR++
Sbjct: 1004 AKLSASEGAPVADPTEYRSLADALQYLTLTRPEIAYAVQQVCRFMHDPHEPHLALVKRIM 1063
Query: 1018 RYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPT 1077
RY++G+L+ GLH+ PV +L +Y+DADW GCPD+RRSTSGYC+FLGDNL+SWSSKRQ T
Sbjct: 1064 RYIKGSLSAGLHIGTGPVGSLTAYSDADWAGCPDSRRSTSGYCIFLGDNLVSWSSKRQTT 1123
Query: 1078 LSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRT 1137
+SRSSAEAEYR VA+ ++E C +R LL ELH P+S T+V CDNVSA Y++ NPVHH+ T
Sbjct: 1124 VSRSSAEAEYRAVAHAIAECCCLRQLLQELHAPISTTTVVFCDNVSAAYMTANPVHHRCT 1183
Query: 1138 KHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPASTA 1197
KHIE+DIHFVREKV GQ RVLHVPS HQ D TKGLP LF DFRSSL V +PPA+TA
Sbjct: 1184 KHIEIDIHFVREKVALGQVRVLHVPSTHQFPDFMTKGLPVQLFTDFRSSLCVRDPPAATA 1243
Score = 61.2 bits (147), Expect = 2e-07
Identities = 57/218 (26%), Positives = 80/218 (36%), Gaps = 60/218 (27%)
Query: 31 TYRSDQGHSNHRYGSGQNRNYQGGSGKPRKKGGSRYPGSP--GSSGSSAASPPPWRPPPQ 88
T S G +R G G N GK + +R G G G PWR P
Sbjct: 243 TCHSSGGQQQNRAGGGNGGNGSKNKGKNGGRNNNRNSGGKCGGGFGGRPGQQQPWRAP-- 300
Query: 89 ASWNPWGWDPPPGWAPSPWGMPPCPYPTSQWSRPMGVPQQPGVLGPRPQAYTA----TAS 144
G+LGP PQA+TA +S
Sbjct: 301 ---------------------------------------SAGILGPYPQAHTAFTGPYSS 321
Query: 145 PAP----------TDIAAAMHTMSLTPPDTPWYMDTGASSHTAASQGTLTSYSNLSHLNQ 194
P P + AA++ M+L P++ W ++TGA+SH ++ G L + L +
Sbjct: 322 PPPAGSVQPAWDQAGLVAALNGMTLHNPNS-WVLNTGATSHMSSIDGILV--TRLPNSQT 378
Query: 195 KLIVGSGQGIPIQGSGYTSIPTPHKPLALNHVLHTPRI 232
+ VGSG IPI G + +P L HVL P +
Sbjct: 379 YITVGSGHSIPIVCRGTSFLPIGTTRFDLRHVLVAPSL 416
>UniRef100_Q9SA17 F28K20.17 protein [Arabidopsis thaliana]
Length = 1415
Score = 789 bits (2037), Expect = 0.0
Identities = 435/1064 (40%), Positives = 599/1064 (55%), Gaps = 45/1064 (4%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W+ D+ A++H +S L S + + ++VG G +PI +G T+I + + + LN
Sbjct: 322 WHPDSAATAHVTSSTNGLQSATEYEG-DDAVLVGDGTYLPITHTGSTTIKSSNGKIPLNE 380
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRSSP 285
VL P I K+L+SV +L D V FD + D +T + LY V +
Sbjct: 381 VLVVPNIQKSLLSVSKLCDDYPCGVYFDANKVCIIDLQTQKVVTTGPRRNGLY-VLENQE 439
Query: 286 FAGLASS--------VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRL 337
F L S+ VWH+RLGH S AL HL+N+K I SR+S VC+ C +GK RL
Sbjct: 440 FVALYSNRQCAATEEVWHHRLGHANSKALQHLQNSKAIQINKSRTSPVCEPCQMGKSSRL 499
Query: 338 PFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYET 396
PF S L P D +H DLW SPV+S G +YY +F+DD++ + W +P+ KS+
Sbjct: 500 PFLISDSRVLHPLDRIHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSWFYPLHNKSEFLSV 559
Query: 397 FTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERK 456
F + L++ Q + IK Q D G E+ ++ + +G+ R SCP+T QNG AERK
Sbjct: 560 FISFQKLVENQLNTKIKVFQSDGGGEFVSNKLKTHLSEHGIHHRISCPYTPQQNGLAERK 619
Query: 457 IRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSHL 516
R + + ++L HS P FW + A Y++N LP L+N SP + L+ P YS L
Sbjct: 620 HRHLVELGLSMLFHSHTPQKFWVESFFTANYIINRLPSSVLKNLSPYEALFGEKPDYSSL 679
Query: 517 RVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETR 576
RVFG C+P NK PRS CVFLGY ++GY+C+ K+ ISR+VIF+E+
Sbjct: 680 RVFGSACYPCLRPLAQNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGKVYISRNVIFNESE 739
Query: 577 FPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQP----SSPTDSSPPLSAL 632
PF + + P Y L+ WQ P P S P D + +
Sbjct: 740 LPFKEKYQSLVPQYST-------PLLQAWQHNKISEISVPAAPVQLFSKPIDLNTYAGSQ 792
Query: 633 VS-------PTASPSPLPLPPVPPAPPV----------RTMTTRSMHGISKPKKPFSLSV 675
V+ PT++ P A + MTTRS GI KP ++L
Sbjct: 793 VTEQLTDPEPTSNNEGSDEEVNPVAEEIAANQEQVINSHAMTTRSKAGIQKPNTRYALIT 852
Query: 676 SIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRH 735
S + + P A+ P W A+ E N + +TW LVP D+N++ W+F+
Sbjct: 853 SRMNTAE---PKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNILSSKWVFKT 909
Query: 736 KKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDV 795
K +G ++ KARLV G Q GVD ETFSPVV+ ATIR VL ++ S+ WPI QLDV
Sbjct: 910 KLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSKGWPIKQLDV 969
Query: 796 QNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGF 855
NAFLHG+L E ++M+ P GF DP P +VCRL K++YGLKQAPRAW+ F++++ GF
Sbjct: 970 SNAFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTFSNFLLDYGF 1029
Query: 856 RHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYF 915
S SD SLF+ Q I Y+LLYVDDI+L S L + + L + F+MKDLGP YF
Sbjct: 1030 VCSKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSMKDLGPPRYF 1089
Query: 916 LGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYR 975
LGI + +A GLFL Q+ YAT+I+ +AGM+ CNP TP+ Q+L + + + + +R
Sbjct: 1090 LGIQIEDYANGLFLHQTAYATDILQQAGMSDCNPMPTPL--PQQLDNLNSELFAEPTYFR 1147
Query: 976 SLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPV 1035
SL G LQYLT TRPDI +AV +C MH+P T LKR+LRY++GT+ GL + +
Sbjct: 1148 SLAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGMGLPIKRNST 1207
Query: 1036 ETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVS 1095
TL +Y+D+D GC +TRRST+G+C+ LG NLISWS+KRQPT+S SS EAEYR +
Sbjct: 1208 LTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAEYRALTYAAR 1267
Query: 1096 ESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQ 1155
E WI LL +L P T V+CDN+SA+YLS NP H R+KH + D H++RE+V G
Sbjct: 1268 EITWISFLLRDLGIPQYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHYIREQVALGL 1327
Query: 1156 ARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV-GEPPASTAG 1198
H+ + Q+AD+FTK LPR F D RS L V G P S G
Sbjct: 1328 IETQHISATFQLADVFTKSLPRRAFVDLRSKLGVSGSPTPSLRG 1371
>UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa]
Length = 1688
Score = 713 bits (1840), Expect = 0.0
Identities = 431/1112 (38%), Positives = 594/1112 (52%), Gaps = 84/1112 (7%)
Query: 130 GVLGPRPQAYTATASPAPTDIAAAMHTMSLTP-PDTPWYMDTGASSHTAASQGTLTSYSN 188
G+LGP P A A+ S T +TP PW +D+GAS H + LTS
Sbjct: 157 GILGPYPCAGIASCSSIST----------VTPIASQPWILDSGASFHMSFDDSWLTSCRL 206
Query: 189 LSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNV 248
+ + V + G + + SI +P + +V P++ NLISV QLT D N
Sbjct: 207 VKN---GATVHTANGTLCKVTHQGSISSPQ--FTVPNVSLVPKLSMNLISVGQLT-DTNC 260
Query: 249 SVSFDPFGFSVSDFKTG--------------MPLLRCNSLGDLYPVTRS--SPFAGLASS 292
V FD V D TG + +L SL T S SP A
Sbjct: 261 FVGFDDTSCFVQDRHTGAVIGTGHRQKRSCGLYILDSLSLPSSSTNTPSVYSPMCSTACK 320
Query: 293 V---WHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRP 349
WH+RLGH S L L N ++ P ++ VC C LGK V+LP+ SS + RP
Sbjct: 321 SFPQWHHRLGHLCGSRLATLINQGVLGSVPVDTTFVCKGCKLGKQVQLPYPSSTSRSSRP 380
Query: 350 FDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQF 408
FD++HSD+W SP S GH YYV+F+DD++ + W + + +SQ+ + + A +I TQF
Sbjct: 381 FDLVHSDVWGKSPFPSKGGHNYYVIFVDDYSRYTWIYFMKHRSQLISIYQSFAQMIHTQF 440
Query: 409 SANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLL 468
S+ I+ + D+G EY +++F + + G + + SCP +QNG AERK R I RTLL
Sbjct: 441 SSAIRIFRSDSGGEYMSNAFREFLVSQGTLPQLSCPGAHAQNGVAERKHRHIIETARTLL 500
Query: 469 AHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSHLRVFGCLCFPLFP 528
S VP FW A+ A YL+N+ P +LQ SP ++L+ P Y HLRVFGC C+ L
Sbjct: 501 IASFVPAHFWAEAISTAVYLINMQPSSSLQGRSPGEVLFGSPPRYDHLRVFGCTCYVLLA 560
Query: 529 SATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETR-FPFADLSLTPA 587
KL +S CVFLGY + H+GY+CYD S R+I ISR V FDE + F ++ + +
Sbjct: 561 PRERTKLTAQSVECVFLGYSLEHKGYRCYDPSARRIRISRDVTFDENKPFFYSSTNQPSS 620
Query: 588 PSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDSSPPLSALVSPTASPSPLPLPPV 647
P LPP I +++ S P P P P+ SP P+ SPSP+ PP
Sbjct: 621 PENSISFLYLPP--IPSPESLPSSPITPSPSPIPPSVPSPTYVPPPPPSPSPSPVSPPPS 678
Query: 648 --------PPAPPVRTMTTRSMHGISKPKKPFSL----------SVSIDDPSISPL---- 685
P P T+ T H +PK P + S+DD S +P
Sbjct: 679 HIPASSSPPHVPSTITLDTFPFHYSRRPKIPNESQPSQPTLEDPTCSVDDSSPAPRYNLR 738
Query: 686 --------------------PHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVN 725
P ++A+ P+WK AM E AL R+NTW++VP P
Sbjct: 739 ARDALRAPNRDDFVVGVVFEPSTYQEAIVLPHWKLAMSEELAALERTNTWDVVPLPSHAV 798
Query: 726 VIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALS 785
I C W+++ K +S+G ERYKARLV G Q G D DETF+PV T+RT++++A +
Sbjct: 799 PITCKWVYKVKTKSDGQVERYKARLVARGFQQAHGRDYDETFAPVAHMTTVRTLIAVAAT 858
Query: 786 RSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQR 845
RSW I Q+DV+NAFLHGDLHE +YMH P G P P +V RL+++LYGLKQAPRAW+ R
Sbjct: 859 RSWTISQMDVKNAFLHGDLHEEVYMHPPPGVEAP--PGHVFRLRRALYGLKQAPRAWFAR 916
Query: 846 FADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFA 905
F+ V + GF S D +LFI +LLYVDD+++ + L+ +F
Sbjct: 917 FSSVVLAAGFSPSDHDPALFIHTSSRGRTLLLLYVDDMLITGDDLEYIAFVKGKLSEQFM 976
Query: 906 MKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSG 965
M DLGPLSYFLGI VT G +LSQ Y +++A++G+ + TP++ +L S+ G
Sbjct: 977 MSDLGPLSYFLGIEVTSTVDGYYLSQHRYIEDLLAQSGLTDSRTTTTPMELHVRLRSTDG 1036
Query: 966 TPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLT 1025
TP +D S YR L G+L YLT TRPDI+YAV + + AP + H L RVLRY+RGT T
Sbjct: 1037 TPLDDPSRYRHLVGSLVYLTVTRPDIAYAVHILSQFVSAPISVHYGHLLRVLRYLRGTTT 1096
Query: 1026 YGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEA 1085
L S L +++D+ W P RRS +GYC+FLG +L++W SK+Q +SRSS EA
Sbjct: 1097 QCLFYAASSPLQLRAFSDSTWASDPIDRRSVTGYCIFLGTSLLTWKSKKQTAVSRSSTEA 1156
Query: 1086 EYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIH 1145
E R +A SE W+R LL + T + CDN AI ++ +P+ H+ TKHI +D
Sbjct: 1157 ELRALATTTSEIVWLRWLLADFGVSCDVPTPLLCDNTGAIQIANDPIKHELTKHIGVDAS 1216
Query: 1146 FVREKVVRGQARVLHVPSRHQIADIFTKGLPR 1177
F R + + +VPS Q+AD FTK R
Sbjct: 1217 FTRSHCQQSTIALHYVPSELQVADFFTKAQTR 1248
>UniRef100_Q710T7 Gag-pol polyprotein [Populus deltoides]
Length = 1382
Score = 663 bits (1711), Expect = 0.0
Identities = 419/1101 (38%), Positives = 585/1101 (53%), Gaps = 53/1101 (4%)
Query: 105 SPWGMPPCPYPTSQWSRPMGVPQQPGVLGPRPQAYTATASPA--PTDIAAAMHTMSLTPP 162
SP G P + T+ + P G P L + Q + + A + I H+ S
Sbjct: 281 SPQGYKPPHHNTAAVASP-GSITDPNTLAEQFQKFLSLQPQAMSASSIGQLPHSSSGIS- 338
Query: 163 DTPWYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLA 222
+ W +D+GAS H + + TS S LS + V + G P+ +G S+ T H L+
Sbjct: 339 HSEWVLDSGASHHMSPDSSSFTSVSPLSSIP----VMTADGTPMPLAGVGSVVTLH--LS 392
Query: 223 LNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLY---- 278
L +V P++ NL S+ Q+ + V F V D ++ + LY
Sbjct: 393 LPNVYLIPKLKLNLASIGQICDSGDYLVMFSGSFCCVQDLQSQKLIGTGRRENGLYILDE 452
Query: 279 ---PVTRSSPFA-------GLASS---VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSV 325
PV ++ L+SS +WH+RLGH +SS L L + + + S
Sbjct: 453 LKVPVVVAATTVDLSFFRLSLSSSSFYLWHSRLGHVSSSRLRFLASTGALGNLKTCDISD 512
Query: 326 CDSCVLGKHVRLPFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWT 384
C C L K LPF+ S ++ PFD++HSD+W SPV + G RYYV F+DDHT + W
Sbjct: 513 CSGCKLAKFSALPFNRSTSVSSSPFDLIHSDVWGPSPVSTKGGSRYYVSFIDDHTRYCWV 572
Query: 385 FPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCP 444
+ + +S+ +E + LIKTQ SA IKC +CD G EY ++ F + +G I + SC
Sbjct: 573 YLMKHRSEFFEIYAAFRALIKTQHSAVIKCFRCDLGGEYTSNKFCQMLALDGTIHQTSCT 632
Query: 445 HTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQ 504
T QNG AERK R I R+LL + V FW A+ A L+N +P SP +
Sbjct: 633 DTPEQNGVAERKHRHIVETARSLLLSAFVLSEFWGEAVLTAVSLINTIPSSHSSGLSPFE 692
Query: 505 LLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKI 564
LY P YS RVFGC F L P NKL RS CVFLGY +GY+C+D +K+
Sbjct: 693 KLYGHVPDYSSFRVFGCTYFVLHPHVERNKLSSRSAICVFLGYGEGKKGYRCFDPITQKL 752
Query: 565 LISRHVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVS--SRPPDPPVQPSSP 622
+S HV+F E PF + T T L S + H S S P S
Sbjct: 753 YVSHHVVFLE-HIPFFSIPST--------THSLTKSDLIHIDPFSEDSGNDTSPYVRSIC 803
Query: 623 TDSSPPLSALVSPT-----ASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSI 677
T +S L+S T +S +P + PP +++ R + P +S S
Sbjct: 804 THNSAGTGTLLSGTPEASFSSTAPQASSEIVDPPPRQSIRIRKSTKL--PDFAYSCYSSS 861
Query: 678 DDPSISPL-----PHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWI 732
++ + P + K+A+ DP + AM E +AL +++TW+LVP P +V+ C W+
Sbjct: 862 FTSFLAYIHCLFEPSSYKEAILDPLGQQAMDEELSALHKTDTWDLVPLPPGKSVVGCRWV 921
Query: 733 FRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQ 792
++ K S+G ERYKARLV G SQ G+D +ETF+P+ K TIRT++++A R W I Q
Sbjct: 922 YKIKTNSDGSIERYKARLVAKGYSQQYGMDYEETFAPIAKMTTIRTLIAVASIRQWHISQ 981
Query: 793 LDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSS 852
LDV+NAFL+GDL E +YM P G H YVC+LKK+LYGLKQAPRAW+++F+ +SS
Sbjct: 982 LDVKNAFLNGDLQEEVYMAPPPGIS--HDSGYVCKLKKALYGLKQAPRAWFEKFSIVISS 1039
Query: 853 IGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPL 912
+GF S+ D +LFI + + LYVDD+I+ D LA F MKDLG L
Sbjct: 1040 LGFVSSSHDSALFIKCTDAGRIILSLYVDDMIITGDDIDGISVLKTELARRFEMKDLGYL 1099
Query: 913 SYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDAS 972
YFLGI V G LSQS Y I+ RA + TP++ + SSS G P D +
Sbjct: 1100 RYFLGIEVAYSPRGYLLSQSKYVANILERARLTDNKTVDTPIEVNARYSSSDGLPLIDPT 1159
Query: 973 LYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYP 1032
LYR++ G+L YLT T PDI+YAV V + +P T H A+ R+LRY+RGT+ L L
Sbjct: 1160 LYRTIVGSLVYLTITHPDIAYAVHVVSQFVASPTTIHWAAVLRILRYLRGTVFQSLLLSS 1219
Query: 1033 SPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVAN 1092
+ L +Y+DAD G P R+S +G+C+FLGD+LISW SK+Q +S+SS EAEY +A+
Sbjct: 1220 TSSLELRAYSDADHGSDPTDRKSVTGFCIFLGDSLISWKSKKQSIVSQSSTEAEYCAMAS 1279
Query: 1093 VVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVV 1152
E W R LL ++ S T ++CDN S+I ++ N V H+RTKHIE+D H R +
Sbjct: 1280 TTKEIVWSRWLLADMGISFSHLTPMYCDNQSSIQIAHNSVFHERTKHIEIDCHLTRHHLK 1339
Query: 1153 RGQARVLHVPSRHQIADIFTK 1173
G + VPS QIAD FTK
Sbjct: 1340 HGTIALPFVPSSLQIADFFTK 1360
>UniRef100_Q6L3H0 Putative receptor kinase [Solanum demissum]
Length = 1358
Score = 645 bits (1664), Expect = 0.0
Identities = 426/1196 (35%), Positives = 595/1196 (49%), Gaps = 71/1196 (5%)
Query: 1 MMLSGPASQTALHTSASRPPASVDSSQQRPTYRSDQGHSNHRYGSGQNRNYQGGSGKPRK 60
+ L+ P S H S P +VDSS + Q Y S +NR G GKPR
Sbjct: 200 LRLAAPPS----HKVVSSP--TVDSS-----ILASQTFEKRTYQSMENRRGGGRFGKPRS 248
Query: 61 KGGSRY-PGSPGSSGSSAASPPP-WRPPPQASWNPWGWDPPPGWAPSPWGMPPCPYPTSQ 118
K + PG PPP + P +N + + P P S
Sbjct: 249 KCSHCHKPGHTRDICYILHGPPPSYDPIVLKEYNEFLRNRASKQTSPPVAYGAQPNQPSN 308
Query: 119 WSRPMGVPQQPGVLGPRPQAYTA--TASPAPTDIAAAMHTMSLTPPDTP---WYMDTGAS 173
+ + + L R T+ S A D++ A ++ + + W MD+GAS
Sbjct: 309 -NAHIAQTEYDEFLQYRANKQTSPQVVSVAQPDVSVAGNSFACVSQSSTLGTWVMDSGAS 367
Query: 174 SHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNHVLHTPRII 233
H + ++ L S++ + + GI + G P + L+ VL+ P
Sbjct: 368 DHISGNKSLL---SDIVYSQSLPAITLANGIQTKPKGVGKAK-PLSSVTLDSVLYVPGSP 423
Query: 234 KNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRSSPFAGLASS- 292
NL SV +LT + S++F F + D TG + + LY +T S+ A + +
Sbjct: 424 FNLASVSRLTKALHCSITFFDDFFLMQDRSTGQMIGTGHESQGLYYLTSSNSLAACSITD 483
Query: 293 ---VWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRP 349
+ H RLGH + S L K++ S S+ C+SC LGKH R FS S
Sbjct: 484 SPDLIHKRLGHSSLSKLQ-----KMVPSLSSLSTLDCESCQLGKHTRATFSRSTEGRSES 538
Query: 350 -FDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQ 407
F ++HSD+W S V ST G RY+V F+DD++ W F + +S+++ F + I+ Q
Sbjct: 539 IFSLVHSDIWGPSRVSSTLGFRYFVSFIDDYSKCTWVFLMKDRSELFSIFKSFFAEIQNQ 598
Query: 408 FSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTL 467
F +I+ + DN EY + F + G+I + +CP+T QNG AERK R + RTL
Sbjct: 599 FGVSIRTFRSDNALEYLSSQFREFMTHQGIIHQTTCPYTPQQNGVAERKNRHLIETARTL 658
Query: 468 LAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYS-HLRVFGCLCFPL 526
L S+VP FW A+ + YL+N +P ++QN P +L+ + Y RVFG CF
Sbjct: 659 LLESNVPLRFWGDAVLTSCYLINRMPSSSIQNQVPHSILFPQSHLYPIPPRVFGSTCFVH 718
Query: 527 FPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETRFPFADLSLTP 586
+ +KL PR+ CVFLGY +GY+CY + L+S V F E++ P+ S P
Sbjct: 719 NLAPGKDKLAPRALKCVFLGYSRVQKGYRCYSHDLHRYLMSADVTFFESQ-PYYTSSNHP 777
Query: 587 APSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDSSPPLSALVSPTASPSPLPLPP 646
S + P TV+S P V P T P LV + +P P P
Sbjct: 778 DVSMVLPIPQVLPVPTFVESTVTST--SPVVVPPLLTYHRRPRPTLVPDDSCHAPDPAPT 835
Query: 647 VPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEF 706
PP + PL +ALS W+ AM E
Sbjct: 836 ADLPPPSQ-----------------------------PLALQKGEALSHSGWRQAMVDEM 866
Query: 707 NALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDET 766
+AL +S TWELV P + + C W++ K +G +R KARLV G +QI G+D +T
Sbjct: 867 SALHKSGTWELVSLPAGKSTVGCRWVYAVKIGPDGQVDRLKARLVAKGYTQIFGLDYSDT 926
Query: 767 FSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGF-RDPHHPDYV 825
F+PV K A++R LS+A R WP+HQLD++NAFLHGDL E +YM P GF V
Sbjct: 927 FAPVAKIASVRLFLSMAAVRHWPLHQLDIKNAFLHGDLEEEVYMEQPPGFVAQGESSSLV 986
Query: 826 CRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQG-SDIAYILLYVDDII 884
CRL++SLYGLKQ+PRAW+ +F+ + G S +DHS+F S Y+++YVDDI+
Sbjct: 987 CRLRRSLYGLKQSPRAWFGKFSTVIQEFGMTRSGADHSVFYRHSAPSRCIYLVVYVDDIV 1046
Query: 885 LVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGM 944
+ + D L F KDLG L YFLGI V + G+ +SQ YA +I+ GM
Sbjct: 1047 ITGNDQDGITDLKQHLFKHFQTKDLGRLKYFLGIEVAQSRSGIVISQRKYALDILEETGM 1106
Query: 945 ASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHA 1004
C P TP+D KL G P + YR L G L YLT TRPDIS+ V V M +
Sbjct: 1107 MGCRPVDTPMDPNVKLLPGQGEPLSNPERYRRLVGKLNYLTVTRPDISFPVSVVSQFMTS 1166
Query: 1005 PHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLG 1064
P H A+ R+LRY++ GL E ++ YTDADW G P RRSTSGYCV +G
Sbjct: 1167 PCDSHWEAVVRILRYIKSAPGKGLLFEDQGHEHIIGYTDADWAGSPSDRRSTSGYCVLVG 1226
Query: 1065 DNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSA 1124
NL+SW SK+Q ++RSSAE+EYR +A E WI+ LL EL F + CDN +A
Sbjct: 1227 GNLVSWKSKKQNVVARSSAESEYRAMATATCELVWIKQLLGELKFGKVDKMELVCDNQAA 1286
Query: 1125 IYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGL--PRL 1178
++++ NPV H+RTKHIE+D HFVREK++ G V S Q+ADIFTK L PR+
Sbjct: 1287 LHIASNPVFHERTKHIEIDCHFVREKILSGDIVTKFVKSNDQLADIFTKSLTCPRI 1342
>UniRef100_Q7X7W7 OSJNBa0061G20.2 protein [Oryza sativa]
Length = 542
Score = 638 bits (1646), Expect = 0.0
Identities = 317/543 (58%), Positives = 391/543 (71%), Gaps = 6/543 (1%)
Query: 656 MTTRSMHGISKPKKPFSLSVSIDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTW 715
M TR GI +P ++ + DD +P + L DP W +AMQ E+ AL + TW
Sbjct: 1 MVTRWKAGIVQPNPRYAHLATADD-----VPTTVRAVLRDPAWFAAMQDEYRALQDNGTW 55
Query: 716 ELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPAT 775
LVPRP +VI WIF++K ++G ER KAR V G +Q G+D D+TFSPVVKPAT
Sbjct: 56 ALVPRPRGAHVITGKWIFKNKFHADGTLERRKARWVARGFTQRPGLDFDKTFSPVVKPAT 115
Query: 776 IRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGL 835
IRTVL +A R WP+HQLDV+NAFLHGDL E Y H P GF DP PD VC L+KSLYGL
Sbjct: 116 IRTVLHLAAMRDWPVHQLDVKNAFLHGDLTEHFYCHQPAGFVDPSQPDAVCLLRKSLYGL 175
Query: 836 KQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKS 895
KQAPRAW+QRF ++ +GF S SD+SLF+ R+G+D A++LLYVDDI+L ASS L +
Sbjct: 176 KQAPRAWFQRFGTHLHHLGFVSSKSDNSLFVLRRGTDEAHLLLYVDDIVLAASSQRLLQH 235
Query: 896 FMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVD 955
+ L EFAMKDLGP+ +FLGI V R A G FLSQ YA +++ RAG+A C P+ TP+D
Sbjct: 236 IINQLRVEFAMKDLGPVHFFLGIQVRRTADGFFLSQEQYAGDVLDRAGLADCKPAPTPID 295
Query: 956 TKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKR 1015
TK K+SS++G P D + YRS+ GALQYLT TRPD+SYAVQQVCLHMH+P H +KR
Sbjct: 296 TKAKVSSTTGQPYSDPTFYRSIVGALQYLTLTRPDLSYAVQQVCLHMHSPRDVHWTLVKR 355
Query: 1016 VLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQ 1075
+LRYVRGT GL L S +L +Y+D DW GCPDTRRSTSG+CVF GD+ SWSSKRQ
Sbjct: 356 ILRYVRGTTHKGLQLRRSSTPSLTAYSDVDWAGCPDTRRSTSGFCVFFGDSSESWSSKRQ 415
Query: 1076 PTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQ 1135
+SRSSAEAEYRGVAN +E CW+R+LL ELH L +ATLV+CDN+SA+YLS NP+HH
Sbjct: 416 SVVSRSSAEAEYRGVANAAAECCWLRHLLGELHVKLDKATLVYCDNISAVYLSKNPLHHG 475
Query: 1136 RTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPAS 1195
R KH+E+D+HFVREKV G RV H+P+R Q+ADI TK LP LF+DFRSSL + + A
Sbjct: 476 RAKHVELDVHFVREKVAVGDIRVAHIPTRQQLADIMTKRLPTALFEDFRSSLCIVD-DAP 534
Query: 1196 TAG 1198
TAG
Sbjct: 535 TAG 537
>UniRef100_Q9XII7 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1454
Score = 627 bits (1616), Expect = e-178
Identities = 371/1055 (35%), Positives = 557/1055 (52%), Gaps = 42/1055 (3%)
Query: 150 IAAAMHTMSLTPPDTPWYMDTGASSHTAASQGTLTSY--SNLSHLNQKLIVGSGQGIPIQ 207
+ A HT+S W +D+GA+ H + + +S S LS +N + +G + I
Sbjct: 419 LTVARHTLS----SATWVIDSGATHHVSHDRSLFSSLDTSVLSAVN----LPTGPTVKIS 470
Query: 208 GSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMP 267
G G + + + L +VL P NLIS+ LT D V FD + D G
Sbjct: 471 GVGTLKL---NDDILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRM 527
Query: 268 LLRCNSLGDLYPVTRS----SPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRSS 323
L + + +LY + S A + S+WH RLGH + L+ + ++ ++ S
Sbjct: 528 LGQGRRVANLYLLDVGDQSISVNAVVDISMWHRRLGHASLQRLDAISDSLGTTRHKNKGS 587
Query: 324 SVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTA-GHRYYVLFLDDHTDFL 382
C C L K +L F +S + FD+LH D+W + T G++Y++ +DDH+
Sbjct: 588 DFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHSRAT 647
Query: 383 WTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFS 442
W + + KS+V F ++ Q+ +K ++ DN E SF+ G++ S
Sbjct: 648 WMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPELKFTSFYA---EKGIVSFHS 704
Query: 443 CPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSP 502
CP T QN ERK + I N+ R L+ S VP S W + A +L+N P + L N +P
Sbjct: 705 CPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNKTP 764
Query: 503 TQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHR 562
++L P Y LR FGCLC+ +K QPRS C+FLGYP ++GYK DL
Sbjct: 765 YEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLESN 824
Query: 563 KILISRHVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSP 622
+ ISR+V F E FP A + + S + FT +P S +S P PS
Sbjct: 825 TVFISRNVQFHEEVFPLAKNPGSES-SLKLFTPMVPVSS----GIISDTTHSPSSLPSQI 879
Query: 623 TDSSPPLSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVS----ID 678
+D P +S S P L TM + + IS +S S I+
Sbjct: 880 SDLPPQIS---SQRVRKPPAHLNDYH----CNTMQSDHKYPISSTISYSKISPSHMCYIN 932
Query: 679 DPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQ 738
+ + P+P N +A W A+ +E A+ ++NTWE+ P + C W+F K
Sbjct: 933 NITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAVGCKWVFTLKFL 992
Query: 739 SNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNA 798
++G ERYKARLV G +Q G+D +TFSPV K TI+ +L ++ S+ W + QLDV NA
Sbjct: 993 ADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKKWFLKQLDVSNA 1052
Query: 799 FLHGDLHETIYMHHPLGFRDPHH----PDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIG 854
FL+G+L E I+M P G+ + + V RLK+S+YGLKQA R W+++F+ + S+G
Sbjct: 1053 FLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWFKKFSSSLLSLG 1112
Query: 855 FRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSY 914
F+ + DH+LF+ + +L+YVDDI++ ++S L F ++DLG L Y
Sbjct: 1113 FKKTHGDHTLFLKMYDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQRFKLRDLGDLKY 1172
Query: 915 FLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLY 974
FLG+ V R G+ + Q YA E++ GM +C P + P+ K+ G ED Y
Sbjct: 1173 FLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSVPMIPNLKMRKDDGDLIEDIEQY 1232
Query: 975 RSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSP 1034
R + G L YLT TRPDI++AV ++C AP T H+ A RVL+Y++GT+ GL S
Sbjct: 1233 RRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKGTVGQGLFYSASS 1292
Query: 1035 VETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVV 1094
TL + D+DW C D+RRST+ + +F+GD+LISW SK+Q T+SRSSAEAEYR +A
Sbjct: 1293 DLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSSAEAEYRALALAT 1352
Query: 1095 SESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRG 1154
E W+ LL+ L +++ D+ +AIY++ NPV H+RTKHI++D H VRE++ G
Sbjct: 1353 CEMVWLFTLLVSLQ-ASPPVPILYSDSTAAIYIATNPVFHERTKHIKLDCHTVRERLDNG 1411
Query: 1155 QARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSV 1189
+ ++LHV + Q+ADI TK L F+ +S +S+
Sbjct: 1412 ELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSI 1446
>UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Solanum tuberosum]
Length = 1476
Score = 625 bits (1611), Expect = e-177
Identities = 392/1090 (35%), Positives = 554/1090 (49%), Gaps = 70/1090 (6%)
Query: 151 AAAMHTMSLTPPDTPWYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSG 210
A + H S T + W +D+GA+ H ++ L ++SH K+ + +G + SG
Sbjct: 404 AGSSHCNSNTH-SSAWIVDSGATDHMVSNTTLLNHGLSVSHPG-KVQLPTGDSAVVTHSG 461
Query: 211 YTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLR 270
+ + + +VL P NL+SV +LT + N V F P F + D TG
Sbjct: 462 SSQLTGGD---VVKNVLCVPTFQFNLLSVSKLTKELNCCVIFFPDFFIIQDLFTGKVKEI 518
Query: 271 CNSLGDLYPV-------TRSSPFAGLA----SSVWHNRLGHPASSALNHLRNNKLIFCEP 319
+ LY T A + + +WH RLGH S L ++ +F P
Sbjct: 519 GEEINGLYITRPHQHHDTSKKTLAAIKGCEEAEMWHKRLGHIPMSVLRKIK----MFDSP 574
Query: 320 SRSS-SVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGH-RYYVLFLDD 377
+ CD C L + VRLPF S + FD++H D+W +T RY++ +DD
Sbjct: 575 QKLVLPSCDVCPLARQVRLPFPISQSRSENCFDLIHLDVWGPYKAATHNKMRYFLTVVDD 634
Query: 378 HTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGL 437
H+ + W F + KS V +I TQF IK + DNG E+ N ++G+
Sbjct: 635 HSRWTWIFLMHLKSDVSTVLQNFILMIDTQFGQKIKIFRSDNGTEFFNAQCDGLFKSHGI 694
Query: 438 IFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTL 497
+ + SCPHT QNG ER+ + I R L +P FW + A +++N +P L
Sbjct: 695 VHQSSCPHTPQQNGVVERRHKHILETARALRFQGHLPIRFWGECVLSAVHIINRIPSSVL 754
Query: 498 QNDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCY 557
N SP +L+Y R P S++RV GCLC L ST +GYK Y
Sbjct: 755 HNKSPFELMYKRSPDLSYMRVIGCLCHA-------TNLVNTST----------QKGYKLY 797
Query: 558 DLSHRKILISRHVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPV 617
DL H+ +SR ++F+E FPF +L F PP H + +P
Sbjct: 798 DLEHQHFFVSRDMVFNEAVFPFQSPALADPHDTPVFLAS-PPCSSHTEDADAVQPAIITS 856
Query: 618 QPSSPTDSSPPLSALVSPTASPSP-------LPLPPVPPAPPVRTMTTRSMHGISKPKKP 670
+ P S P SA+ P P PP+ + T T+RS H +
Sbjct: 857 EEIIPVASPP--SAVSDDHLHPPPERRRSYRTGKPPIWQKDFITTSTSRSNHCLYPISDN 914
Query: 671 FSLSVS-------IDDPSISPLPHNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCD 723
S I S+ P QA +D W AM+ E AL + TWE+V P
Sbjct: 915 IDYSCLSSTYQCYIASSSVETEPQFYYQAANDCRWVHAMKEEIQALEDNKTWEVVSLPKG 974
Query: 724 VNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIA 783
I C W+++ K +++G ER+KARLV G +Q G+D ETFSPVVK T+RTVL++A
Sbjct: 975 KKAIGCKWVYKIKYKASGEIERFKARLVAKGYNQKEGLDYQETFSPVVKMVTLRTVLTLA 1034
Query: 784 LSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFR-DPHHPDYVCRLKKSLYGLKQAPRAW 842
+S+ W I Q+DV NAFL GDL E +YM P GF+ D VCRL KSLYGLKQA R W
Sbjct: 1035 VSKGWDIQQMDVYNAFLQGDLIEEVYMQLPQGFQYDKTGDPKVCRLLKSLYGLKQASRQW 1094
Query: 843 YQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLAS 902
+ + + GF+ S D+SL + R I +L+YVDD+++ SS L +L +
Sbjct: 1095 NVKLTTALLAAGFQQSHLDYSLMLKRTADGIVIVLIYVDDLLITGSSLQLIDDAKQVLKA 1154
Query: 903 EFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKL-- 960
F +KDLG L YFLG+ R+A G+ + Q YA E+I+ G+ PS TPV+ KL
Sbjct: 1155 NFKIKDLGTLRYFLGMEFARNASGMLMHQRKYALELISDLGLGGSKPSVTPVELHLKLTT 1214
Query: 961 --------SSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLA 1012
SS + + D + Y+ L G L YLT TRPDIS+AVQ + MHAP HM A
Sbjct: 1215 REFDLHVGSSGADSLLADPTEYQRLVGRLLYLTITRPDISFAVQHLSQFMHAPKVSHMEA 1274
Query: 1013 LKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSS 1072
RV++YV+ GL++ +TL +Y DADWG C +TR+S +GY + G L+SW S
Sbjct: 1275 AIRVVKYVKQAPGLGLYMAVQTADTLQAYCDADWGSCINTRKSITGYMIQFGSALLSWKS 1334
Query: 1073 KRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPV 1132
K+QPT+SRSSAEAEYR +A+ V+E W+ L EL PLS ++CD+ +AI ++ NPV
Sbjct: 1335 KKQPTISRSSAEAEYRSLASTVAELVWLTGLFKELDMPLSLPVSLYCDSKAAIQIAANPV 1394
Query: 1133 HHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGE- 1191
H+RTKHI++D HF+REKV G + ++P++ Q ADI TKGL S L +
Sbjct: 1395 FHERTKHIDIDCHFIREKVQAGLVMIHYLPTQEQPADILTKGLSSAQHSYLVSKLGLKNI 1454
Query: 1192 --PPASTAGV 1199
PP+ G+
Sbjct: 1455 FIPPSLKGGI 1464
>UniRef100_Q7Y141 Putative polyprotein [Oryza sativa]
Length = 1335
Score = 621 bits (1601), Expect = e-176
Identities = 366/1040 (35%), Positives = 551/1040 (52%), Gaps = 43/1040 (4%)
Query: 163 DTPWYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLA 222
D W +D+G ++H AA + H K+ +G+G +G G ++ T P
Sbjct: 323 DDVWVIDSGCTNHMAADPNLFREMDSSYHA--KIHMGNGSIAQSEGKGTVAVQTADGPKF 380
Query: 223 LNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTR 282
+ VL P + +NL+S+ QL ++ +V F+ F + D K + + N + + R
Sbjct: 381 IKDVLLVPDLKQNLLSIGQLL-EHGYAVYFEDFSCKILDRKNNRLVAKINMEKNRNFLLR 439
Query: 283 SSPFAGLA-------SSVWHNRLGHPASSALNHLRNNKLIFCEP--SRSSSVCDSCVLGK 333
+ +A S +WH R+GH AL LR ++ P + S C+ CV GK
Sbjct: 440 MNHTTQMALRSEVDISDLWHKRMGHLNYRALKLLRTKGMVQGLPFITLKSDPCEGCVFGK 499
Query: 334 HVRLPFS-SSAIITLRPFDILHSDLWTS-PVLSTAGHRYYVLFLDDHTDFLWTFPISKKS 391
+R F S A P +++H+D+ P +S G+ Y++ F+DD+T +W + + +KS
Sbjct: 500 QIRASFPHSGAWRASAPLELVHADIVGKVPTISEGGNWYFITFIDDYTRMIWVYFLKEKS 559
Query: 392 QVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNG 451
E F +++ Q + IK L+ D GREY + F +YC+ G+ + + +++ QNG
Sbjct: 560 AALEIFKKFKAMVENQSNRKIKVLRSDQGREYISKEFEKYCENAGIRRQLTAGYSAQQNG 619
Query: 452 KAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDP 511
AERK RTIN+M ++L +P SFW A+ A Y+LN P K + N +P + Y + P
Sbjct: 620 VAERKNRTINDMANSMLQDKGMPKSFWAEAVNTAVYILNRSPTKAVTNRTPFEAWYGKKP 679
Query: 512 SYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVI 571
H+RVFGC+C+ P+ K +S C+F+GY +GY+ Y+L +KI+ISR I
Sbjct: 680 VIGHMRVFGCICYAQVPAQKRVKFDNKSDRCIFVGYADGIKGYRLYNLEKKKIIISRDAI 739
Query: 572 FDETR-FPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDSSPPLS 630
FDE+ + + + P T L +H V P P QPSSP SS
Sbjct: 740 FDESATWNWKSPEASSTPLLPTTTITLGQPHMHGTHEVEDHTPSP--QPSSPMSSSS--- 794
Query: 631 ALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDDPSISPL-PHNP 689
S +SPS P + P R RSM + + S + + S + P +
Sbjct: 795 --ASSDSSPSSEEQISTPESAPRRV---RSMVELLESTSQQRGSEQHEFCNYSVVEPQSF 849
Query: 690 KQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKAR 749
++A NW AM+ E + + ++NTWELV RP D VI W+++ K +G ++YKAR
Sbjct: 850 QEAEKHDNWIKAMEDEIHMIEKNNTWELVDRPRDREVIGVKWVYKTKLNPDGSVQKYKAR 909
Query: 750 LVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIY 809
LV G Q G+D ET++PV + TIRT++++A + W I+QLDV++AFL+G L E IY
Sbjct: 910 LVAKGFKQKPGIDYYETYAPVARLETIRTIIALAAQKRWKIYQLDVKSAFLNGYLDEEIY 969
Query: 810 MHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQ 869
+ P GF + V RLKK+LYGLKQAPRAWY + Y GF S S+ +L++ +
Sbjct: 970 VEQPEGFSVQGGENKVFRLKKALYGLKQAPRAWYSQIDKYFIQKGFAKSISEPTLYVNKT 1029
Query: 870 GSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFL 929
G+DI + LYVDD+I +S + + F + + M DLG L YFLG+ V + G+F+
Sbjct: 1030 GTDILIVSLYVDDLIYTGNSEKMMQDFKKDMMHTYEMSDLGLLHYFLGMEVHQSDEGIFI 1089
Query: 930 SQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRP 989
SQ YA I+ + M +C TP+ +K + G D ++YRSL G+L YLT TRP
Sbjct: 1090 SQRKYAENILKKFKMDNCKSVTTPLLPNEKQKARDGADKADPTIYRSLVGSLLYLTATRP 1149
Query: 990 DISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGC 1049
DI +A + +M +P + A KRVLRY++GT YG+ P L+ YTD+DW GC
Sbjct: 1150 DIMFAASLLSRYMSSPSQLNFTAAKRVLRYIKGTADYGIWYKPVKESKLIGYTDSDWAGC 1209
Query: 1050 PDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHF 1109
D +STSGY LG SAEAEY + VS+ W+R ++ +L
Sbjct: 1210 LDDMKSTSGYAFSLG-----------------SAEAEYVAASKAVSQVVWLRRIMEDLGE 1252
Query: 1110 PLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIAD 1169
Q T ++CD+ SAI +S NPV H RTKHI + H++RE V R + ++ + Q+AD
Sbjct: 1253 KQYQPTTIYCDSKSAIAISENPVSHDRTKHIAIKYHYIREAVDRQEVKLEFCRTDEQLAD 1312
Query: 1170 IFTKGLPRLLFDDFRSSLSV 1189
IFTK L + F R + V
Sbjct: 1313 IFTKALSKEKFVRDRELIGV 1332
>UniRef100_O04543 F20P5.25 protein [Arabidopsis thaliana]
Length = 1315
Score = 615 bits (1586), Expect = e-174
Identities = 368/1021 (36%), Positives = 533/1021 (52%), Gaps = 88/1021 (8%)
Query: 205 PIQGSGYTSIPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKT 264
PI GS + + L LN VL P+ NL+SV LT + FD + D
Sbjct: 312 PISGSVHLG-----RHLILNDVLFIPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDATR 366
Query: 265 GMPLLRCNSLGDLY---------PVTRSSPFAGLASS--VWHNRLGHPASSALNHLRNNK 313
+ + + +LY P T SS +S +WH RLGHP+ L + +
Sbjct: 367 ELMVGMGKQVANLYIVDLDSLSHPGTDSSITVASVTSHDLWHKRLGHPSVQKLQPMSSLL 426
Query: 314 LIFCEPSRSSSVCDSCVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTA-GHRYYV 372
+ + + C C + K LPF S + RPFD++H D W + T G+RY++
Sbjct: 427 SFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRYFL 486
Query: 373 LFLDDHTDFLWTFPISKKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYC 432
+DD++ W + + KS V T T+++ QF IK ++ DN E + F ++
Sbjct: 487 TIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELN---FTQFY 543
Query: 433 DANGLIFRFSCPHTSSQNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNIL 492
+ G++ SCP T QN ERK + I N+ R+L S +P S+W + A YL+N L
Sbjct: 544 HSKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINRL 603
Query: 493 PRKTLQNDSPTQLLYHRDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHR 552
P L++ P ++L P+Y H++VFGCLC+ +K PR+ C F+GYP +
Sbjct: 604 PAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSGFK 663
Query: 553 GYKCYDLSHRKILISRHVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRP 612
GYK DL I++SRHV+F E FPF L+ + F DL P+
Sbjct: 664 GYKLLDLETHSIIVSRHVVFHEELFPFLGSDLSQEE--QNFFPDLNPT------------ 709
Query: 613 PDPPVQPSS-----PTDSSPPLSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKP 667
PP+Q S P+DSS + L P+A+P+ P P V+T K
Sbjct: 710 --PPMQRQSSDHVNPSDSSSSVEIL--PSANPTNNV-----PEPSVQTSHR-------KA 753
Query: 668 KKPFSLSVSIDDPSISPLPHNPKQALS-----DPN------------------------W 698
KKP L +S PH ++ LS DP W
Sbjct: 754 KKPAYLQDYYCHSVVSSTPHEIRKFLSYDRINDPYLTFLACLDKTKEPSNYTEAEKLQVW 813
Query: 699 KSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQI 758
+ AM +EF+ L ++TWE+ P D I C WIF+ K S+G ERYKARLV G +Q
Sbjct: 814 RDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRWIFKIKYNSDGSVERYKARLVAQGYTQK 873
Query: 759 AGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFR- 817
G+D +ETFSPV K +++ +L +A + QLD+ NAFL+GDL E IYM P G+
Sbjct: 874 EGIDYNETFSPVAKLNSVKLLLGVAARFKLSLTQLDISNAFLNGDLDEEIYMRLPQGYAS 933
Query: 818 ---DPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIA 874
D P+ VCRLKKSLYGLKQA R WY +F+ + +GF S DH+ F+
Sbjct: 934 RQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFSSTLLGLGFIQSYCDHTCFLKISDGIFL 993
Query: 875 YILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTY 934
+L+Y+DDII+ +++ + + S F ++DLG L YFLG+ + R G+ +SQ Y
Sbjct: 994 CVLVYIDDIIIASNNDAAVDILKSQMKSFFKLRDLGELKYFLGLEIVRSDKGIHISQRKY 1053
Query: 935 ATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYA 994
A +++ G C PS+ P+D + SG + YR L G L YL TRPDI++A
Sbjct: 1054 ALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGDFVEVGPYRRLIGRLMYLNITRPDITFA 1113
Query: 995 VQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRR 1054
V ++ AP H+ A+ ++L+Y++GT+ GL + L Y +AD+ C D+RR
Sbjct: 1114 VNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQGLFYSATSELQLKVYANADYNSCRDSRR 1173
Query: 1055 STSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQA 1114
STSGYC+FLGD+LI W S++Q +S+SSAEAEYR ++ E W+ N L EL PLS+
Sbjct: 1174 STSGYCMFLGDSLICWKSRKQDVVSKSSAEAEYRSLSVATDELVWLTNFLKELQVPLSKP 1233
Query: 1115 TLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKG 1174
TL+ CDN +AI+++ N V H+RTKHIE D H VRE++++G + H+ + QIAD FTK
Sbjct: 1234 TLLFCDNEAAIHIANNHVFHERTKHIESDCHSVRERLLKGLFELYHINTELQIADPFTKP 1293
Query: 1175 L 1175
L
Sbjct: 1294 L 1294
>UniRef100_Q9FX79 Putative retroelement polyprotein [Arabidopsis thaliana]
Length = 1413
Score = 611 bits (1576), Expect = e-173
Identities = 350/1016 (34%), Positives = 534/1016 (52%), Gaps = 70/1016 (6%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W +D+GA+ H + + + + + + +G I + G G ++ + L L +
Sbjct: 447 WIVDSGATHHVCHVKDMFLNLD--TSVQHHVNLPTGTTIRVGGVGNIAV---NADLILKN 501
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRSSP 285
VL+ P NL+SV LTTD V FDP V D G
Sbjct: 502 VLYIPEFRLNLLSVSALTTDIGARVVFDPTCCVVHDLTKG-------------------- 541
Query: 286 FAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPFSSSAII 345
+ + S + + LG IF ++ CD C K +L + S I
Sbjct: 542 -STIGSDLLTDVLG---------------IFKTKNKGLLHCDICQRAKQKKLTYPSRHNI 585
Query: 346 TLRPFDILHSDLW---TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYETFTTLAT 402
L PFD+LH D+W + P + G+ Y++ +DDHT W + + KS V F T
Sbjct: 586 CLAPFDLLHIDVWGPFSEP--TQEGYHYFLTIVDDHTRVTWVYLMKYKSDVLTIFPDFIT 643
Query: 403 LIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERKIRTINN 462
+++TQ+ +K ++ DN E + +R G++ SCP T QN ERK + I N
Sbjct: 644 MVETQYDTKVKAVRSDNAPELKFEELYR---RKGIVAYHSCPETPEQNSVVERKHQHILN 700
Query: 463 MIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSHLRVFGCL 522
+ R LL S +P S+W + A +++N P + N + ++L + P Y+HL+ FGCL
Sbjct: 701 VARALLFQSQIPLSYWGDCILTAVFIINRTPSPVISNKTLFEMLTKKVPDYTHLKSFGCL 760
Query: 523 CFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETRFPFADL 582
C+ +K + R+ C FLGYP ++GYK DL I ISR+V+F E FPF
Sbjct: 761 CYASTSPKQRHKFEDRARTCAFLGYPSGYKGYKLLDLESHTIFISRNVVFYEDLFPF--- 817
Query: 583 SLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDSSPPLSALVSPTASPSPL 642
PA + E+ H + + P P+ + S+ P S + P
Sbjct: 818 KTKPAEN-----EESSVFFPHIYVDRNDSHPSQPLPVQETSASNVPAEKQNSRVSRP--- 869
Query: 643 PLPPVPPAPPVRTMTTRSMHGISKPKKPFSLS----VSIDDPSISPLPHNPKQALSDPNW 698
P ++T+ + H IS+ SLS + I+ + P PH QA W
Sbjct: 870 --PAYLKDYHCNSVTSSTDHPISEVLSYSSLSDPYMIFINAVNKIPEPHTYAQARQIKEW 927
Query: 699 KSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQI 758
AM E AL + TW + P + C W+++ K ++G ERYKARLV G +Q
Sbjct: 928 CDAMGMEITALEDNGTWVVCSLPVGKKAVGCKWVYKIKLNADGSLERYKARLVAKGYTQT 987
Query: 759 AGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFR- 817
G+D +TFSPV K T++ ++++A ++ W + QLD+ NAFL+G L E IYM P G+
Sbjct: 988 EGLDYVDTFSPVAKLTTVKLLIAVAAAKGWSLSQLDISNAFLNGSLDEEIYMTLPPGYSP 1047
Query: 818 ---DPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIA 874
D P+ VCRLKKSLYGLKQA R WY +F++ + ++GF S+ DH+LF + +
Sbjct: 1048 RQGDSFPPNAVCRLKKSLYGLKQASRQWYLKFSESLKALGFTQSSGDHTLFTRKSKNSYM 1107
Query: 875 YILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTY 934
+L+YVDDII+ +S + L ++DLG L YFLG+ + R+ G+ + Q Y
Sbjct: 1108 AVLVYVDDIIIASSCDRETELLRDALQRSSKLRDLGTLRYFLGLEIARNTDGISICQRKY 1167
Query: 935 ATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYA 994
E++A G+ C S+ P++ QKLS G +DA YR L G L YLTFTRPDI+YA
Sbjct: 1168 TLELLAETGLLGCKSSSVPMEPNQKLSQEDGELIDDAEHYRKLVGKLMYLTFTRPDITYA 1227
Query: 995 VQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRR 1054
V ++C AP H+ A+ +++ Y++GT+ GL + L + D+D+ C D+R+
Sbjct: 1228 VHRLCQFTSAPRVPHLKAVYKIIYYLKGTVGQGLFYSANVDLKLSGFADSDFSSCSDSRK 1287
Query: 1055 STSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQA 1114
T+GYC+FLG +L++W SK+Q +S SSAEAEY+ ++ V E W+R LL +L +S+A
Sbjct: 1288 LTTGYCMFLGTSLVAWKSKKQEVISMSSAEAEYKAMSMAVREMMWLRFLLEDLWIDVSEA 1347
Query: 1115 TLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADI 1170
++++CDN +AI+++ NPV H+RTKHIE D H +REK++ G R LHV + +Q+ADI
Sbjct: 1348 SVLYCDNTAAIHIANNPVFHERTKHIERDYHHIREKIILGLIRTLHVRTENQLADI 1403
>UniRef100_O23588 Retrotransposon like protein [Arabidopsis thaliana]
Length = 1433
Score = 602 bits (1551), Expect = e-170
Identities = 354/1031 (34%), Positives = 526/1031 (50%), Gaps = 51/1031 (4%)
Query: 156 TMSLTPPDTP--WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTS 213
++S P +P W +D+GA+ H ++ ++ +L + +L + + I G G+
Sbjct: 420 SVSQEPAVSPRGWVIDSGATHHVTHNRDLYLNFRSLENTFVRL--PNDCTVKIAGIGFIQ 477
Query: 214 IPTPHKPLALNHVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNS 273
+ ++L++VL+ P NLIS +LT + + V DF +
Sbjct: 478 LSDA---ISLHNVLYIPEFKFNLIS--ELTKELMIGRGSQVGNLYVLDFNENNHTVSLKG 532
Query: 274 LGDLYPVTRSSPFAGLASSVWHNRLGHPASSALNHLRNN-----KLIFCEPSRSSSVCDS 328
+ P + S WH RLGHPA S ++ L + K I E S VC
Sbjct: 533 TTSMCPEFSVCSSVVVDSVTWHKRLGHPAYSKIDLLSDVLNLKVKKINKEHSPVCHVCHV 592
Query: 329 CVLGKHVRLPFSSSAIITLRPFDILHSDLWTSPVLSTAGHRYYVLFLDDHTDFLWTFPIS 388
C L K L F S + FD++H D W + T D W + +
Sbjct: 593 CHLSKQKHLSFQSRQNMCSAAFDLVHIDTWGPFSVPT-------------NDATWIYLLK 639
Query: 389 KKSQVYETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSS 448
KS V F ++ TQ+ +K ++ DN E F A+G++ SCP T
Sbjct: 640 NKSDVLHVFPAFINMVHTQYQTKLKSVRSDNAHEL---KFTDLFAAHGIVAYHSCPETPE 696
Query: 449 QNGKAERKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYH 508
QN ERK + I N+ R LL S++P FW + A +L+N LP L N SP + L +
Sbjct: 697 QNSVVERKHQHILNVARALLFQSNIPLEFWGDCVLTAVFLINRLPTPVLNNKSPYEKLKN 756
Query: 509 RDPSYSHLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISR 568
P+Y L+ FGCLC+ +K +PR+ CVFLGYP+ ++GYK D+ + ISR
Sbjct: 757 IPPAYESLKTFGCLCYSSTSPKQRHKFEPRARACVFLGYPLGYKGYKLLDIETHAVSISR 816
Query: 569 HVIFDETRFPFADLSLTPAPSYECFTEDLPPSLIHHWQTVSSRPPDPPVQPSSPTDSSPP 628
HVIF E FPF ++ +D P L +R D P++ +S D+ P
Sbjct: 817 HVIFHEDIFPFISSTIKDD------IKDFFPLL-----QFPARTDDLPLEQTSIIDTHPH 865
Query: 629 LSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDDPSISPLPHN 688
S P PL PP + ++P F I++ + + +P
Sbjct: 866 QDVSSSKALVPFD-PLSKRQKKPPKHLQDFHCYNNTTEPFHAF-----INNITNAVIPQR 919
Query: 689 PKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKA 748
+A W AM+ E A++R+NTW +V P + I C W+F K ++G ERYKA
Sbjct: 920 YSEAKDFKAWCDAMKEEIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKA 979
Query: 749 RLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETI 808
RLV G +Q G+D +ETFSPV K ++R +L +A W +HQLD+ NAFL+GDL E I
Sbjct: 980 RLVAKGYTQEEGLDYEETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEI 1039
Query: 809 YMHHPLGFRD----PHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSL 864
YM P G+ D P +CRL KS+YGLKQA R WY + ++ + +GF+ S +DH+L
Sbjct: 1040 YMKIPPGYADLVGEALPPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTL 1099
Query: 865 FIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHA 924
FI + +L+YVDDI++V++S D F A L S F ++DLG YFLGI + R
Sbjct: 1100 FIKYANGVLMGVLVYVDDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSE 1159
Query: 925 GGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYL 984
G+ + Q Y E+++ G PS+ P+D KL+ G P D++ YR L G L YL
Sbjct: 1160 KGISICQRKYILELLSTTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYL 1219
Query: 985 TFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDA 1044
TRPDI+YAV +C HAP + H+ A+ +VLRY++GT+ GL L YTD+
Sbjct: 1220 QITRPDIAYAVNTLCQFSHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDS 1279
Query: 1045 DWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLL 1104
D+G C D+RR + YC+F+GD L+SW SK+Q T+S S+AEAE+R ++ E W+ L
Sbjct: 1280 DFGSCTDSRRCVAAYCMFIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLF 1339
Query: 1105 LELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSR 1164
+ P ++CDN +A+++ N V H+RTK +E+D + RE V G + + V +
Sbjct: 1340 DDFKVPFIPPAYLYCDNTAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETG 1399
Query: 1165 HQIADIFTKGL 1175
Q+AD TK +
Sbjct: 1400 EQVADPLTKAI 1410
>UniRef100_Q8W5M3 Putative gag-pol polyprotein [Oryza sativa]
Length = 725
Score = 600 bits (1546), Expect = e-169
Identities = 289/468 (61%), Positives = 357/468 (75%)
Query: 731 WIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPI 790
W+F+ K ++G ++YKAR V G +Q GVD ETFSPVVKPATIRTVL++ S+ WP+
Sbjct: 216 WVFKTKLHADGSLDKYKARWVVRGFNQCPGVDFGETFSPVVKPATIRTVLTLISSKQWPV 275
Query: 791 HQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYV 850
HQLDV NAFLHG L E + P GF D P VC L +SLYGL+QAPRAW++RFAD+V
Sbjct: 276 HQLDVSNAFLHGHLQERVLCQQPTGFEDAARPADVCLLSRSLYGLRQAPRAWFKRFADHV 335
Query: 851 SSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLG 910
+S+GF S +D SLF+ R+GSD AY+LLYVDD+IL ASS L + + L +EF +KD+G
Sbjct: 336 TSLGFVQSRADPSLFVLRRGSDTAYLLLYVDDMILSASSSSLLQRIIDRLQAEFKVKDMG 395
Query: 911 PLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCED 970
PL YFLGI V R A G LSQS YAT ++ RAGMA+C ATP D K KLSS G +D
Sbjct: 396 PLKYFLGIEVQRTADGFVLSQSKYATNVLERAGMANCKAVATPADAKPKLSSDEGPLFQD 455
Query: 971 ASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHL 1030
+S YRS+ GALQYLT TRPDI+YAVQQVCLHMHAP H+ LKR+LRY++GT +GLHL
Sbjct: 456 SSWYRSITGALQYLTLTRPDIAYAVQQVCLHMHAPREAHVTLLKRILRYIKGTAAFGLHL 515
Query: 1031 YPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLGDNLISWSSKRQPTLSRSSAEAEYRGV 1090
S TL +++DADW GCPDTRRSTSG+C+FLGD+LISWSSKRQ T+SRSSAEAEYRGV
Sbjct: 516 RASTSPTLTAFSDADWAGCPDTRRSTSGFCIFLGDSLISWSSKRQTTVSRSSAEAEYRGV 575
Query: 1091 ANVVSESCWIRNLLLELHFPLSQATLVHCDNVSAIYLSGNPVHHQRTKHIEMDIHFVREK 1150
AN V+E W+R LL ELH + QAT+ +CDN+S++Y+S NPVHH+R KHIE+DIHFVREK
Sbjct: 576 ANAVAECTWLRQLLGELHCRVPQATIAYCDNISSVYMSKNPVHHKRIKHIELDIHFVREK 635
Query: 1151 VVRGQARVLHVPSRHQIADIFTKGLPRLLFDDFRSSLSVGEPPASTAG 1198
V G+ RVL + S HQ AD+FTKGLP +F++FR+SL VG+ A+TAG
Sbjct: 636 VALGELRVLPISSAHQFADVFTKGLPSSMFNEFRASLCVGDTTAATAG 683
>UniRef100_Q5XWK9 Gag-pol polyprotein-like [Solanum tuberosum]
Length = 1212
Score = 598 bits (1541), Expect = e-169
Identities = 351/918 (38%), Positives = 513/918 (55%), Gaps = 51/918 (5%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLIVGSGQGIPIQGSGYTSIPTPHKPLALNH 225
W +D+GAS+H S L + +Q + + +G +PI G + PT +
Sbjct: 320 WIVDSGASNHMTNSTSILKNVRKYQGPSQ-IQIANGSNLPITKVGDIT-PT------FKN 371
Query: 226 VLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPVTRSSP 285
V +P++ +LISV QL DNN V+F G V D +G + + +G L+P+ S P
Sbjct: 372 VFVSPKLSTSLISVGQLV-DNNCDVNFSRNGCLVQDQVSGTIIAKGPKVGRLFPIHFSIP 430
Query: 286 ----FAGLASS----VWHNRLGHPASSALNHLRNNKLIFCEP--SRSSSVCDSCVLGKHV 335
FA +++ VWH RLGHP S L+H+ N+ L+ + S +S C +C LGK
Sbjct: 431 PVLSFACTSTASKTEVWHKRLGHPNSVVLSHISNSGLLGNKNKFSVASIDCSTCKLGKSK 490
Query: 336 RLPFSSSAIITLRPFDILHSDLW-TSPVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVY 394
LPF + + FD++HSD+W SP++S A +Y++ F+DD++ F W + + KS+V+
Sbjct: 491 TLPFPNFGSRATKCFDVIHSDVWGISPIISHAHFKYFMTFIDDYSRFTWVYFLRSKSEVF 550
Query: 395 ETFTTLATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAE 454
F T I+TQFS IK L+ D+G EY + F ++ G++ + SCP+T QNG AE
Sbjct: 551 SMFKTFLAYIETQFSTCIKLLRSDSGGEYMSYEFKKFLLDKGIVSQHSCPYTPQQNGVAE 610
Query: 455 RKIRTINNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYS 514
RK R + ++ RTLL SSVP +W AL A YL+N LP K L +SP LYH++P+YS
Sbjct: 611 RKNRHLLDVTRTLLIESSVPSKYWVEALSTAVYLINRLPSKVLNLESPYFRLYHQNPNYS 670
Query: 515 HLRVFGCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDE 574
FGC+CF P + NKL +ST C F+GY + +G+ CYD K ISR+V+F E
Sbjct: 671 DFHTFGCVCFVHLPPSQCNKLSVQSTKCAFMGYSTSQKGFICYDPCSHKFRISRNVVFFE 730
Query: 575 TRFPFADL----SLTPA-PSYECFTEDLP---PSLIHHWQTVSSRPPDPPVQPSSPTDSS 626
++ F + S++P P++E + P ++ + RP P P +++
Sbjct: 731 NQYFFPTIVDLSSVSPLLPTFEDLSSSFKRFKPGFVYERR----RPTLPYPNTDPPPETA 786
Query: 627 PPLSALVSPTASPSPLPLPPVPPAPPVRTMTTRSMHGISKPKKPFSLSVSIDDPSISPLP 686
P L + S + P L P TR +S+ + S ++ + S+ P
Sbjct: 787 PQLESENSSRSGP----LEP-----------TRRSTRVSRTPNWYGFSSTLSNISV---P 828
Query: 687 HNPKQALSDPNWKSAMQSEFNALIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERY 746
QA W+ AM+ E AL ++TW++V P +V I C W++ K S+G +RY
Sbjct: 829 SCYSQASKHECWQKAMEEELLALKENDTWDIVSCPSNVRPIGCKWVYSIKLHSDGTLDRY 888
Query: 747 KARLVGDGRSQIAGVDCDETFSPVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHE 806
KARLV G Q GVD +ETF+PV K T+RT+++IA S++W ++Q DV+NAFLHGDL E
Sbjct: 889 KARLVVLGNRQEYGVDYEETFAPVAKMTTVRTIIAIAASQNWSLYQKDVKNAFLHGDLKE 948
Query: 807 TIYMHHPLGFRDPHHPDYVCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFI 866
IYM P D VC+LK+SLYGLKQAPRAW+ +F + F S D SLF+
Sbjct: 949 DIYMKPPPDLFSSPTSD-VCKLKRSLYGLKQAPRAWFDKFRSTLLQFSFELSKYDSSLFL 1007
Query: 867 FRQGSDIAYILLYVDDIILVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGG 926
+ + +L+YVDDII+ + L L F MKDLG L+YFLG+ V A G
Sbjct: 1008 RKTSTSCVLLLVYVDDIIITGTDSSLITCLQQQLKDSFHMKDLGTLTYFLGLEVHNVASG 1067
Query: 927 LFLSQSTYATEIIARAGMASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTF 986
+FL+Q Y ++I+ AG+ + TP++ K G D +++R L G+L YLT
Sbjct: 1068 VFLNQHKYTQDLISLAGLQVSSSVDTPLEMNVKYRREEGDLLPDPTIFRQLVGSLNYLTI 1127
Query: 987 TRPDISYAVQQVCLHMHAPHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADW 1046
TRPDIS+AVQQV M AP H++A+ ++RY+ GT T GL L +++D+DW
Sbjct: 1128 TRPDISFAVQQVSQFMQAPRHLHLVAVCHIIRYLLGTSTRGLFFPSGSPIRLNAFSDSDW 1187
Query: 1047 GGCPDTRRSTSGYCVFLG 1064
GCPDTRRS SG+C+FLG
Sbjct: 1188 AGCPDTRRSVSGWCMFLG 1205
>UniRef100_O81617 F8M12.17 protein [Arabidopsis thaliana]
Length = 1633
Score = 596 bits (1537), Expect = e-168
Identities = 374/1071 (34%), Positives = 559/1071 (51%), Gaps = 112/1071 (10%)
Query: 166 WYMDTGASSHTAASQGTLTSYSNLSHLNQKLI-VGSGQGIPIQGSGYTSIPTPHKPLALN 224
W +D+GASSH + LT + L H++ + + +G + I +G I + L L+
Sbjct: 397 WIIDSGASSHVCSD---LTMFRELIHVSGVTVTLPNGTRVAITHTGTICITST---LILH 450
Query: 225 HVLHTPRIIKNLISVRQLTTDNNVSVSFDPFGFSVSDFKTGMPLLRCNSLGDLYPV---- 280
+VL P NLISV L + G+ + R + +LY +
Sbjct: 451 NVLLVPDFKFNLISVCCL------------------ELTRGLMIGRGKTYNNLYILETQR 492
Query: 281 TRSSPFAGLASSVWHNRLGHPASSALNHLRNNKLIFCEPSRSSSVCDSCVLGKHVRLPFS 340
T SP A+S HP+ AL L ++ S ++S C L K RL +
Sbjct: 493 TSFSPSLPAATS------RHPSLPALQKLVSSIPSLKSVSSTASHCRISPLAKQKRLAYV 546
Query: 341 SSAIITLRPFDILHSDLWTS-PVLSTAGHRYYVLFLDDHTDFLWTFPISKKSQVYETFTT 399
S + PFD++H D+W + S G RY++ +DD T W + + KS+V F
Sbjct: 547 SHNNLASSPFDLIHLDIWGPFSIESVDGFRYFLTLVDDCTRTTWVYMMKNKSEVSNIFPV 606
Query: 400 LATLIKTQFSANIKCLQCDNGREYDNDSFHRYCDANGLIFRFSCPHTSSQNGKAERKIRT 459
LI TQ++A IK ++ DN +E +F ++ G+I +FSC +T QN ERK +
Sbjct: 607 FVKLIFTQYNAKIKAIRSDNVKEL---AFTKFVKEQGMIHQFSCAYTPQQNSVVERKHQH 663
Query: 460 INNMIRTLLAHSSVPPSFWHHALQMATYLLNILPRKTLQNDSPTQLLYHRDPSYSHLRVF 519
+ N+ R+LL S+VP +W + A YL+N LP L N +P +LL + P Y+ L+
Sbjct: 664 LLNIARSLLFQSNVPLQYWSDCVLTAAYLINRLPSPLLDNKTPFELLLKKIPDYTLLK-- 721
Query: 520 GCLCFPLFPSATINKLQPRSTPCVFLGYPMNHRGYKCYDLSHRKILISRHVIFDETRFPF 579
CLC+ NK PR+ PCVFLGYP ++GYK DL I I+R+V+F ET+FPF
Sbjct: 722 SCLCYASTNVHDRNKFSPRARPCVFLGYPSGYKGYKVLDLESHSISITRNVVFHETKFPF 781
Query: 580 ---------ADL---SLTPAPSYECFTEDLP-----------PSLIHHWQTVSSRPPDPP 616
D+ S+ P P+ F E +P S + + SS PP P
Sbjct: 782 KTSKFLKESVDMFPNSILPLPAPLHFVESMPLDDDLRADDNNASTSNSASSASSIPPLPS 841
Query: 617 VQPSSPTDS------SPPLSALVSPTASPSPLP------LPPVPPAPPVRTMTTRSMHGI 664
+ TD+ S P++ +P+ L +P + P + + +
Sbjct: 842 TVNTQNTDALDIDTNSVPIARPKRNAKAPAYLSEYHCNSVPFLSSLSPTTSTSIETPSSS 901
Query: 665 SKPKK---PFSLSVSIDDPSISPLPHN----------PK---QALSDPNWKSAMQSEFNA 708
PKK P+ +S +I ++PL H+ PK QA+ W A E +A
Sbjct: 902 IPPKKITTPYPMSTAISYDKLTPLFHSYICAYNVETEPKAFTQAMKSEKWTRAANEELHA 961
Query: 709 LIRSNTWELVPRPCDVNVIRCMWIFRHKKQSNGLFERYKARLVGDGRSQIAGVDCDETFS 768
L ++ TW + NV+ C W+F K +G ERYKARLV G +Q G+D ETFS
Sbjct: 962 LEQNKTWIVESLTEGKNVVGCKWVFTIKYNPDGSIERYKARLVAQGFTQQEGIDYMETFS 1021
Query: 769 PVVKPATIRTVLSIALSRSWPIHQLDVQNAFLHGDLHETIYMHHPLGFRDPHHPDY---- 824
PV K +++ +L +A + W + Q+DV NAFLHG+L E IYM P G+ P
Sbjct: 1022 PVAKFGSVKLLLGLAAATGWSLTQMDVSNAFLHGELDEEIYMSLPQGYTPPTGISLPSKP 1081
Query: 825 VCRLKKSLYGLKQAPRAWYQRFADYVSSIGFRHSTSDHSLFIFRQGSDIAYILLYVDDII 884
VCRL KSLYGLKQA R WY+R + F S +D+++F+ + I +L+YVDD++
Sbjct: 1082 VCRLLKSLYGLKQASRQWYKRLSSVFLGANFIQSPADNTMFVKVSCTSIIVVLVYVDDLM 1141
Query: 885 LVASSHDLRKSFMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYATEIIARAGM 944
+ ++ ++ LL SEF +KDLGP +FLG+ + R + G+ + Q YA ++ G+
Sbjct: 1142 IASNDSSAVENLKELLRSEFKIKDLGPARFFLGLEIARSSEGISVCQRKYAQNLLEDVGL 1201
Query: 945 ASCNPSATPVDTKQKLSSSSGTPCEDASLYRSLDGALQYLTFTRPDISYAVQQVCLHMHA 1004
+ C PS+ P+D L+ GT +A+ YR L G L YL TRPDI++AV + + A
Sbjct: 1202 SGCKPSSIPMDPNLHLTKEMGTLLPNATSYRELVGRLLYLCITRPDITFAVHTLSQFLSA 1261
Query: 1005 PHTKHMLALKRVLRYVRGTLTYGLHLYPSPVETLVSYTDADWGGCPDTRRSTSGYCVFLG 1064
P HM A +VLRY++G +P + DADWG C D+RRS +G+C++LG
Sbjct: 1262 PTDIHMQAAHKVLRYLKG----------NPGQ------DADWGTCKDSRRSVTGFCIYLG 1305
Query: 1065 DNLISWSSKRQPTLSRSSAEAEYRGVANVVSESCWIRNLLLELHFPLSQATLVHCDNVSA 1124
+LI+W SK+Q +SRSS E+EYR +A E W++ LL +LH ++ + CDN SA
Sbjct: 1306 TSLITWKSKKQSVVSRSSTESEYRSLAQATCEIIWLQQLLKDLHVTMTCPAKLFCDNKSA 1365
Query: 1125 IYLSGNPVHHQRTKHIEMDIHFVREKVVRGQARVLHVPSRHQIADIFTKGL 1175
++L+ NPV H+RTKHIE+D H VR+++ G+ + LHVP+ +Q+ADI TK L
Sbjct: 1366 LHLATNPVFHERTKHIEIDCHTVRDQIKAGKLKTLHVPTGNQLADILTKPL 1416
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.134 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,166,808,571
Number of Sequences: 2790947
Number of extensions: 102912889
Number of successful extensions: 699286
Number of sequences better than 10.0: 8266
Number of HSP's better than 10.0 without gapping: 2466
Number of HSP's successfully gapped in prelim test: 6252
Number of HSP's that attempted gapping in prelim test: 513019
Number of HSP's gapped (non-prelim): 79146
length of query: 1199
length of database: 848,049,833
effective HSP length: 139
effective length of query: 1060
effective length of database: 460,108,200
effective search space: 487714692000
effective search space used: 487714692000
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0490.3


