

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0404.10
(989 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
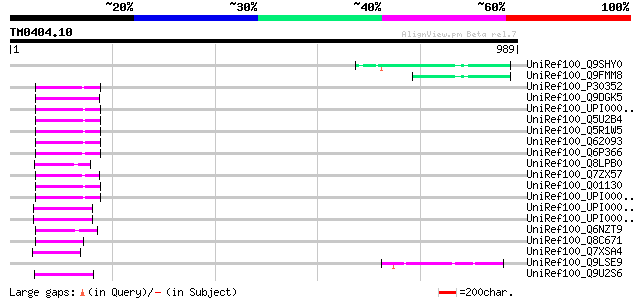
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana] 76 6e-12
UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like... 66 5e-09
UniRef100_P30352 Splicing factor, arginine/serine-rich 2 [Gallus... 66 6e-09
UniRef100_Q9DGK5 Splicing factor arginine/serine rich 2 [Oryzias... 65 1e-08
UniRef100_UPI000024E83B UPI000024E83B UniRef100 entry 64 2e-08
UniRef100_Q5U2B4 Hypothetical protein [Xenopus laevis] 64 2e-08
UniRef100_Q5R1W5 Arginine/serine-rich 2 splicing factor [Pan tro... 64 2e-08
UniRef100_Q62093 Splicing factor, arginine/serine-rich 2 [Mus mu... 64 2e-08
UniRef100_Q6P366 Hypothetical protein MGC75633 [Xenopus tropicalis] 63 5e-08
UniRef100_Q8LPB0 Glycine-rich RNA-binding protein [Physcomitrell... 63 5e-08
UniRef100_Q7ZX57 Sfrs2-prov protein [Xenopus laevis] 61 1e-07
UniRef100_Q01130 Splicing factor, arginine/serine-rich 2 [Homo s... 61 1e-07
UniRef100_UPI000036DCDF UPI000036DCDF UniRef100 entry 59 6e-07
UniRef100_UPI000029C32C UPI000029C32C UniRef100 entry 59 7e-07
UniRef100_UPI000001775C UPI000001775C UniRef100 entry 59 9e-07
UniRef100_Q6NZT9 Zgc:55876 protein [Brachydanio rerio] 58 2e-06
UniRef100_Q8C671 Mus musculus 0 day neonate head cDNA, RIKEN ful... 58 2e-06
UniRef100_Q7XSA4 OJ000126_13.13 protein [Oryza sativa] 58 2e-06
UniRef100_Q9LSE9 Reverse transcriptase-like protein [Arabidopsis... 58 2e-06
UniRef100_Q9U2S6 Hypothetical protein Y116A8C.34 [Caenorhabditis... 58 2e-06
>UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana]
Length = 1055
Score = 75.9 bits (185), Expect = 6e-12
Identities = 75/316 (23%), Positives = 120/316 (37%), Gaps = 36/316 (11%)
Query: 675 GAGWGMGKSTKRMVVKQKIRNSRAELVILQETKMDVHRNKVLLGWATSMD---------- 724
G GW K + N+R EL + + R++ L W S D
Sbjct: 329 GRGWDFAK-----IDPYTTNNTRLELRAVVLDLVTGARDR--LSWKFSQDGQFSVRSAYE 381
Query: 725 -MGLEIVPAVGSAGGLVTLWKTSLFQVSQQLMEC--NIAVKASLIRRGIPIENGGLCDIC 781
+ ++ VP A LWK + + + + N AV R + +C +C
Sbjct: 382 MLTVDEVPRPNMASFFNCLWKVRVPERVKTFLWLVGNQAVMTEEERHRRHLSASNVCQVC 441
Query: 782 GLIGESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWEII 841
ES H++ CP +W ++ + F S+ L + R+ + I W I
Sbjct: 442 KGGVESMLHVLRDCPAQLGIWVRVVPQRRQQGFFSKSLFEWLYDNLGDRSGCEDIPWSTI 501
Query: 842 PYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKI 901
+ W W+ R +F + D + ++K W E + H +
Sbjct: 502 FAVIIWWGWKWRCGNIFGENTKCRDRVK---------FVKEWAVEV------YRAHSGNV 546
Query: 902 RIKVAAPKVHSA-DWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVG 960
+ + P+V W P G +K N DG+SRGN G++ GGVL+D G FS +G
Sbjct: 547 LVGITQPRVERMIGWVSPCVGWVKVNTDGASRGNPGLASAGGVLRDCTGAWCGGFSLNIG 606
Query: 961 STWAYVAERKALLEGL 976
A AE + GL
Sbjct: 607 RCSAPQAELWGVYYGL 622
>UniRef100_Q9FMM8 Non-LTR retroelement reverse transcriptase-like protein
[Arabidopsis thaliana]
Length = 308
Score = 66.2 bits (160), Expect = 5e-09
Identities = 51/192 (26%), Positives = 75/192 (38%), Gaps = 16/192 (8%)
Query: 786 ESESHLMLHCPKIWLLWTTILAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWEIIPYAM 845
ES H+ CP +W + R F S+ L + R+ + I W I +
Sbjct: 25 ESVLHVFRDCPAQLGIWVRFVPRRRQQGFFSKSLFEWLYDNLCDRSSCEDIPWSTIFAVI 84
Query: 846 FWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFEKIRIKV 905
W W+ R +F + D + ++K W E + H +
Sbjct: 85 IWWGWKWRCSNIFGENTKCRDRVK---------FVKEWVVEV------YRAHLGNALVGS 129
Query: 906 AAPKVHSA-DWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAVGSTWA 964
P+V W P G +K N DG+SRGN G++ GGVL+D G FS +G A
Sbjct: 130 TQPRVERLIGWVLPCVGWVKVNTDGASRGNPGLASAGGVLRDCEGAWCGGFSLNIGRCSA 189
Query: 965 YVAERKALLEGL 976
AE + GL
Sbjct: 190 QHAELWGVYYGL 201
>UniRef100_P30352 Splicing factor, arginine/serine-rich 2 [Gallus gallus]
Length = 221
Score = 65.9 bits (159), Expect = 6e-09
Identities = 42/131 (32%), Positives = 67/131 (51%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G S +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRY-GSSGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>UniRef100_Q9DGK5 Splicing factor arginine/serine rich 2 [Oryzias latipes]
Length = 211
Score = 64.7 bits (156), Expect = 1e-08
Identities = 39/129 (30%), Positives = 65/129 (50%), Gaps = 3/129 (2%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 1 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYSKESRGFAFVRFFDKRDAEDAMDA 60
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P ++R P ++ G +G S+ G+ R R+
Sbjct: 61 MDGALLDGRELRVQMARYGRPPDSMYSRRGPPPRRYGGGGGYGRRSRSRSGSPRRRRRSR 120
Query: 167 IPLKRNSQS 175
+ S+S
Sbjct: 121 SRSRSRSRS 129
>UniRef100_UPI000024E83B UPI000024E83B UniRef100 entry
Length = 252
Score = 64.3 bits (155), Expect = 2e-08
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>UniRef100_Q5U2B4 Hypothetical protein [Xenopus laevis]
Length = 271
Score = 64.3 bits (155), Expect = 2e-08
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 65 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 124
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 125 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 183
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 184 SRSRSRSRSRY 194
>UniRef100_Q5R1W5 Arginine/serine-rich 2 splicing factor [Pan troglodytes]
Length = 221
Score = 64.3 bits (155), Expect = 2e-08
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 133
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 134 SRSRSRSRSRY 144
>UniRef100_Q62093 Splicing factor, arginine/serine-rich 2 [Mus musculus]
Length = 220
Score = 64.3 bits (155), Expect = 2e-08
Identities = 41/131 (31%), Positives = 66/131 (50%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 14 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 73
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 74 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 132
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 133 SRSRSRSRSRY 143
>UniRef100_Q6P366 Hypothetical protein MGC75633 [Xenopus tropicalis]
Length = 220
Score = 62.8 bits (151), Expect = 5e-08
Identities = 40/131 (30%), Positives = 65/131 (49%), Gaps = 6/131 (4%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H R P ++ ++G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHGRRGPPPRRY---GDYGRRSRSPRRRRRSRSRSK 131
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 132 SRSRSRSRSRY 142
>UniRef100_Q8LPB0 Glycine-rich RNA-binding protein [Physcomitrella patens]
Length = 155
Score = 62.8 bits (151), Expect = 5e-08
Identities = 43/113 (38%), Positives = 58/113 (51%), Gaps = 10/113 (8%)
Query: 48 GKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAM 107
G LFV GL TT +K AF++FG + V + + R FGFV F+ D DAE A+
Sbjct: 41 GSKLFVGGLAWGTTDDNIKEAFSAFGEVTEVKIICDRDTGRSRGFGFVTFATDQDAEAAL 100
Query: 108 RRMNGSILNGATINVSLARLPQDQHTRPRPTDNQWKGVSEWG--EHSKPNKGN 158
+ ++G L G TI V+ A T+ P D Q G +G +HS N GN
Sbjct: 101 QALDGRDLAGRTIRVNYA-------TKQSPQDRQ-SGGGMYGRRDHSGGNSGN 145
>UniRef100_Q7ZX57 Sfrs2-prov protein [Xenopus laevis]
Length = 215
Score = 61.2 bits (147), Expect = 1e-07
Identities = 39/129 (30%), Positives = 64/129 (49%), Gaps = 6/129 (4%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H R P ++ ++G S+ + RRS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHGRRGPAPRRY---GDYGRRSRSPRRRRRSRSRSK 131
Query: 167 IPLKRNSQS 175
+ S+S
Sbjct: 132 SRSRSRSRS 140
>UniRef100_Q01130 Splicing factor, arginine/serine-rich 2 [Homo sapiens]
Length = 220
Score = 61.2 bits (147), Expect = 1e-07
Identities = 40/131 (30%), Positives = 65/131 (49%), Gaps = 4/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F + R+ V++ + + F FVRF DAE AM
Sbjct: 14 SLKVDNLTYRTSPDTLRRVFEKYRRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 73
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR P H+R P ++ G +G S+ + RRS +
Sbjct: 74 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG-GGYGRRSRSPRRRRRSRSRSR 132
Query: 167 IPLKRNSQSDY 177
+ S+S Y
Sbjct: 133 SRSRSRSRSRY 143
>UniRef100_UPI000036DCDF UPI000036DCDF UniRef100 entry
Length = 224
Score = 59.3 bits (142), Expect = 6e-07
Identities = 38/131 (29%), Positives = 65/131 (49%), Gaps = 5/131 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
+L VD LT +T+ ++ F +GR+ V++ + F FVRF SDA+ A
Sbjct: 15 TLKVDNLTYRTSPDSLRRVFEKYGRVGDVYIPREPHTKAPRGFAFVRFHDRSDAQDAEAA 74
Query: 110 MNGSILNGATINVSLARLPQ---DQHTRPRPTDNQWKGVSEWGEHSKPNKGNRRSARKEW 166
M+G++L+G + V +AR + + ++ P+ W G +G S+ +G RS +
Sbjct: 75 MDGAVLDGRELRVQMARYGRRDLPRSSQEEPSGRSWGG--RYGRRSRSPRGRHRSQSRGP 132
Query: 167 IPLKRNSQSDY 177
K S+S Y
Sbjct: 133 SYSKSRSRSHY 143
>UniRef100_UPI000029C32C UPI000029C32C UniRef100 entry
Length = 391
Score = 58.9 bits (141), Expect = 7e-07
Identities = 39/118 (33%), Positives = 56/118 (47%), Gaps = 4/118 (3%)
Query: 46 RIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEV 105
R GK LF+ GL +TT ++ F+ +GRI V + + + F FV F SDA+
Sbjct: 6 RPGK-LFIGGLNTETTEKALEQFFSKYGRIAEVILMKDRETNKSRGFAFVTFESPSDAKD 64
Query: 106 AMRRMNGSILNGATINVSLARLPQDQ---HTRPRPTDNQWKGVSEWGEHSKPNKGNRR 160
A R MNG L+G I V A PQ + P P ++ +G+ S+ G R
Sbjct: 65 AAREMNGKSLDGKNIKVEQATKPQFESGGRRGPPPMHSRSRGLPRGMRGSRGGPGGMR 122
>UniRef100_UPI000001775C UPI000001775C UniRef100 entry
Length = 390
Score = 58.5 bits (140), Expect = 9e-07
Identities = 39/118 (33%), Positives = 55/118 (46%), Gaps = 4/118 (3%)
Query: 46 RIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEV 105
R GK LF+ GL +TT ++ F+ +GRI V + + + F FV F SDA+
Sbjct: 6 RPGK-LFIGGLNTETTEKALEQFFSKYGRIAEVLLMKDRETNKSRGFAFVTFESPSDAKD 64
Query: 106 AMRRMNGSILNGATINVSLARLPQDQ---HTRPRPTDNQWKGVSEWGEHSKPNKGNRR 160
A R MNG L+G I V A PQ + P P ++ +G S+ G R
Sbjct: 65 AAREMNGKSLDGKNIKVEQATKPQFESGGRRGPPPMHSRSRGAPRGMRGSRGGPGGMR 122
>UniRef100_Q6NZT9 Zgc:55876 protein [Brachydanio rerio]
Length = 220
Score = 57.8 bits (138), Expect = 2e-06
Identities = 41/123 (33%), Positives = 61/123 (49%), Gaps = 8/123 (6%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPETLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARLPQDQHTRPRPTDNQWKGVSEWGEHSKPNKGNRR-SARKEWIP 168
M+G+IL+G + V +AR RP D+ + G S G+RR AR+ P
Sbjct: 75 MDGAILDGRELRVQMARY-------GRPPDSYYGGGGGGRRGSGGGGGSRRHGARRSRSP 127
Query: 169 LKR 171
+R
Sbjct: 128 RRR 130
>UniRef100_Q8C671 Mus musculus 0 day neonate head cDNA, RIKEN full-length enriched
library, clone:4833401I21 product:splicing factor,
arginine/serine- rich 2 (SC-35), full insert sequence
[Mus musculus]
Length = 254
Score = 57.8 bits (138), Expect = 2e-06
Identities = 33/98 (33%), Positives = 52/98 (52%), Gaps = 3/98 (3%)
Query: 50 SLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMRR 109
SL VD LT +T+ ++ F +GR+ V++ + + F FVRF DAE AM
Sbjct: 15 SLKVDNLTYRTSPDTLRRVFEKYGRVGDVYIPRDRYTKESRGFAFVRFHDKRDAEDAMDA 74
Query: 110 MNGSILNGATINVSLARL---PQDQHTRPRPTDNQWKG 144
M+G++L+G + V +AR P H+R P ++ G
Sbjct: 75 MDGAVLDGRELRVQMARYGRPPDSHHSRRGPPPRRYGG 112
>UniRef100_Q7XSA4 OJ000126_13.13 protein [Oryza sativa]
Length = 137
Score = 57.8 bits (138), Expect = 2e-06
Identities = 31/93 (33%), Positives = 47/93 (50%)
Query: 45 RRIGKSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAE 104
R I LF+ GL+ T + AF+ +G++L + K R FGFV+F+ + A
Sbjct: 33 RGITYRLFIGGLSQFATEDSLAEAFSQYGQVLEATIVTDKMTNRPKGFGFVKFASEEAAN 92
Query: 105 VAMRRMNGSILNGATINVSLARLPQDQHTRPRP 137
A MNG +LNG I V +A+ ++ T P
Sbjct: 93 KAKEEMNGKVLNGRVIYVDIAKAKMNRTTDSSP 125
>UniRef100_Q9LSE9 Reverse transcriptase-like protein [Arabidopsis thaliana]
Length = 343
Score = 57.8 bits (138), Expect = 2e-06
Identities = 61/243 (25%), Positives = 99/243 (40%), Gaps = 16/243 (6%)
Query: 725 MGLEIVPAVGSAGGLVTLWKTS----LFQVSQQLMECNIAVKASLIRRGIPIENGGLCDI 780
M EI P G A +WK + +L+ +A +L RR I N C
Sbjct: 1 MDPEIQPPPGKAEIKAKIWKLKTAPKIKHFLWKLLSGALATGDNLKRRHI--RNHPQCHR 58
Query: 781 CGLIGESESHLMLHCPKIWLLWTTI-LAREEVYCCFPSSIQGLLLEWSSLRAISDPIMWE 839
C E+ HL C +W + +E+ + + L SS A P ++
Sbjct: 59 CCQEDETSQHLFFDCFYAQQVWRASGIPHQELRTTGITMETKMELLLSSCLANRQPQLFN 118
Query: 840 IIPYAMFWSIWRARNDLVFNQKDFSADYIWDLHLIRVMWWIKSWWKECPFTVLDFSQHFE 899
+ + + W +W++RN LVF QK S W L R ++ W++ V +Q
Sbjct: 119 LAIWIL-WRLWKSRNQLVFQQKSIS----WQNTLQRARNDVQE-WEDTNTYVQSLNQQVH 172
Query: 900 KIRIKVAAPKVHSADWCPPPPGCLKFNVDGSSRGNSGVSGIGGVLKDANSTMLGRFSCAV 959
R + P + W PP +K+N DG+ + + G +++D N +G A+
Sbjct: 173 SSRHQ--QPTMARTKWQRPPSTWIKYNYDGAFNHQTRNAKAGWLMRDENGVYMGS-GQAI 229
Query: 960 GST 962
GST
Sbjct: 230 GST 232
>UniRef100_Q9U2S6 Hypothetical protein Y116A8C.34 [Caenorhabditis elegans]
Length = 331
Score = 57.8 bits (138), Expect = 2e-06
Identities = 36/117 (30%), Positives = 57/117 (47%), Gaps = 3/117 (2%)
Query: 49 KSLFVDGLTDQTTHLQVKNAFASFGRILGVFVQNVKRKQRRSKFGFVRFSFDSDAEVAMR 108
++L+V G T+ T + AF FG ++ + + + FGFV F DA +A+
Sbjct: 11 RTLYVGGFTEDVTEKVLMAAFIPFGDVVAISIPMDYESGKHRGFGFVEFDMAEDAAMAID 70
Query: 109 RMNGSILNGATINVSLARLPQ--DQHTRPRPTDNQW-KGVSEWGEHSKPNKGNRRSA 162
MN S L G TI V+ AR P+ ++ +P D++W K GE + G+ A
Sbjct: 71 NMNESELFGKTIRVNFARPPKATERSQKPVWADDEWLKKYGRGGEAAAEEDGDAEKA 127
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.319 0.134 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,720,393,990
Number of Sequences: 2790947
Number of extensions: 74489972
Number of successful extensions: 198498
Number of sequences better than 10.0: 1913
Number of HSP's better than 10.0 without gapping: 814
Number of HSP's successfully gapped in prelim test: 1099
Number of HSP's that attempted gapping in prelim test: 195940
Number of HSP's gapped (non-prelim): 3331
length of query: 989
length of database: 848,049,833
effective HSP length: 137
effective length of query: 852
effective length of database: 465,690,094
effective search space: 396767960088
effective search space used: 396767960088
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 80 (35.4 bits)
Lotus: description of TM0404.10


