

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0403.8
(117 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
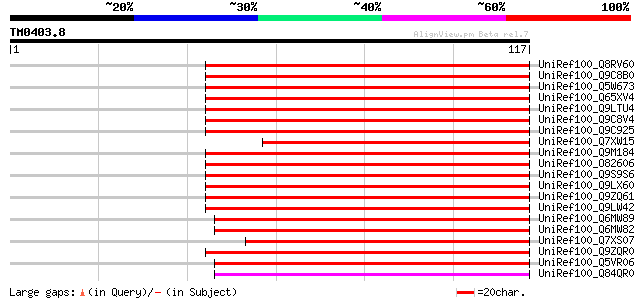
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis tha... 79 2e-14
UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thal... 75 3e-13
UniRef100_Q5W673 Putative helicase [Oryza sativa] 75 3e-13
UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa] 75 3e-13
UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana] 75 3e-13
UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis tha... 75 3e-13
UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis tha... 74 8e-13
UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa] 74 8e-13
UniRef100_Q9M184 Hypothetical protein T5C2_50 [Arabidopsis thali... 74 1e-12
UniRef100_O82606 T2L5.8 protein [Arabidopsis thaliana] 73 1e-12
UniRef100_Q9S9S6 F28J9.3 [Arabidopsis thaliana] 73 1e-12
UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thal... 71 5e-12
UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana] 71 5e-12
UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana] 71 5e-12
UniRef100_Q6MW89 B1248C03.15 protein [Oryza sativa] 71 7e-12
UniRef100_Q6MW82 B1340F09.1 protein [Oryza sativa] 71 7e-12
UniRef100_Q7XS07 OSJNBa0095H06.12 protein [Oryza sativa] 70 9e-12
UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana] 70 1e-11
UniRef100_Q5VR06 Helicase-like protein [Oryza sativa] 69 3e-11
UniRef100_Q84QR0 Helicase-like protein [Oryza sativa] 69 3e-11
>UniRef100_Q8RV60 Hypothetical protein At2g05640 [Arabidopsis thaliana]
Length = 1308
Score = 79.0 bits (193), Expect = 2e-14
Identities = 38/73 (52%), Positives = 49/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT L +I A V G+ G Y+PR+ LT +D +PF+F RRQF V CF MTIN
Sbjct: 1179 LQVTQLGDRVIEAKVLTGSNAGNKVYLPRLVLTPADFRIPFRFQRRQFPVVPCFGMTINK 1238
Query: 105 NQGQSLSHVSLYL 117
+QGQSLSHV +YL
Sbjct: 1239 SQGQSLSHVGIYL 1251
>UniRef100_Q9C8B0 Hypothetical protein F10O5.11 [Arabidopsis thaliana]
Length = 1678
Score = 75.1 bits (183), Expect = 3e-13
Identities = 34/73 (46%), Positives = 51/73 (69%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G IP ++LT +D+ LPFK RRQF +++ FAMTIN
Sbjct: 1555 LQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINK 1614
Query: 105 NQGQSLSHVSLYL 117
+QGQSL H+ LYL
Sbjct: 1615 SQGQSLEHIGLYL 1627
>UniRef100_Q5W673 Putative helicase [Oryza sativa]
Length = 1634
Score = 75.1 bits (183), Expect = 3e-13
Identities = 36/73 (49%), Positives = 50/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
M +T L + I A + G +G+ YIPRI +T ++SG PF RRQ+ +++CFAMTIN
Sbjct: 1515 MTITQLGKRFIEAQIITGTHVGEKVYIPRIIMTPTESGWPFLLKRRQYPLSVCFAMTINK 1574
Query: 105 NQGQSLSHVSLYL 117
+QGQSL+ V LYL
Sbjct: 1575 SQGQSLNMVGLYL 1587
>UniRef100_Q65XV4 Hypothetical protein P0016H04.14 [Oryza sativa]
Length = 1525
Score = 75.1 bits (183), Expect = 3e-13
Identities = 36/73 (49%), Positives = 50/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
M +T L + I A + G +G+ YIPRI +T ++SG PF RRQ+ +++CFAMTIN
Sbjct: 1406 MTITQLGKRFIEAQIITGTHVGEKVYIPRIIMTPTESGWPFLLKRRQYPLSVCFAMTINK 1465
Query: 105 NQGQSLSHVSLYL 117
+QGQSL+ V LYL
Sbjct: 1466 SQGQSLNMVGLYL 1478
>UniRef100_Q9LTU4 Helicase-like protein [Arabidopsis thaliana]
Length = 1428
Score = 75.1 bits (183), Expect = 3e-13
Identities = 35/73 (47%), Positives = 51/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G+ YIP I++T SD+ LPFK RRQF +++ F MTIN
Sbjct: 1306 LQITQLCSHIVEAKVITGDRIGQIVYIPLINITPSDTKLPFKMRRRQFPLSVAFVMTINK 1365
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V LYL
Sbjct: 1366 SQGQSLEQVGLYL 1378
>UniRef100_Q9C8V4 Hypothetical protein T22A15.15 [Arabidopsis thaliana]
Length = 729
Score = 75.1 bits (183), Expect = 3e-13
Identities = 34/73 (46%), Positives = 51/73 (69%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G IP ++LT +D+ LPFK RRQF +++ FAMTIN
Sbjct: 606 LQITQLCTQIVEAKVITGDRIGNIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINK 665
Query: 105 NQGQSLSHVSLYL 117
+QGQSL H+ LYL
Sbjct: 666 SQGQSLEHIGLYL 678
>UniRef100_Q9C925 Hypothetical protein F14G24.23 [Arabidopsis thaliana]
Length = 996
Score = 73.9 bits (180), Expect = 8e-13
Identities = 36/73 (49%), Positives = 50/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + +I A GN++GK IPR+ +T SD+ LPFK RRQF +++ FAMTIN
Sbjct: 898 LQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRLPFKMRRRQFPLSVAFAMTINK 957
Query: 105 NQGQSLSHVSLYL 117
+QGQ+L V LYL
Sbjct: 958 SQGQTLESVGLYL 970
>UniRef100_Q7XW15 OSJNBb0013O03.3 protein [Oryza sativa]
Length = 2052
Score = 73.9 bits (180), Expect = 8e-13
Identities = 32/60 (53%), Positives = 46/60 (76%)
Query: 58 TVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQGQSLSHVSLYL 117
T+ G+K+G+ ++PRISL +++S PF RRQF V +C+AMTIN +QGQ+LSHV +YL
Sbjct: 1309 TILTGSKIGEIVFVPRISLNTTNSKWPFTLQRRQFPVRVCYAMTINKSQGQTLSHVGVYL 1368
>UniRef100_Q9M184 Hypothetical protein T5C2_50 [Arabidopsis thaliana]
Length = 830
Score = 73.6 bits (179), Expect = 1e-12
Identities = 35/73 (47%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + +I A GN++GK IPR+ +T SD+ LPFK R+QF +++ FAMTIN
Sbjct: 616 LQVTQMADTVIQARFITGNRVGKIVLIPRMLITPSDTRLPFKMRRKQFALSVAFAMTINK 675
Query: 105 NQGQSLSHVSLYL 117
+QGQ+L V LYL
Sbjct: 676 SQGQTLESVGLYL 688
Score = 71.2 bits (173), Expect = 5e-12
Identities = 34/73 (46%), Positives = 49/73 (66%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
++VT + +I A GN++GK IPR+ +T D+ LPFK R+QF +++ FAMTIN
Sbjct: 732 LQVTQMADTVIQARFITGNRVGKIVLIPRMLITPLDTRLPFKMRRKQFALSVAFAMTINK 791
Query: 105 NQGQSLSHVSLYL 117
+QGQ+L V LYL
Sbjct: 792 SQGQTLESVGLYL 804
Score = 31.6 bits (70), Expect = 4.4
Identities = 14/20 (70%), Positives = 16/20 (80%)
Query: 98 FAMTINNNQGQSLSHVSLYL 117
FAMTIN +QGQ+L V LYL
Sbjct: 553 FAMTINKSQGQTLESVGLYL 572
>UniRef100_O82606 T2L5.8 protein [Arabidopsis thaliana]
Length = 1073
Score = 73.2 bits (178), Expect = 1e-12
Identities = 35/73 (47%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+ +G IP ++LT +D+ LPFK RRQF +++ FAMTIN
Sbjct: 950 LQITQLCTQIVEAKVITGDIIGHIVLIPTVNLTPTDTKLPFKMRRRQFPLSVAFAMTINT 1009
Query: 105 NQGQSLSHVSLYL 117
+QGQSL HV LYL
Sbjct: 1010 SQGQSLEHVGLYL 1022
>UniRef100_Q9S9S6 F28J9.3 [Arabidopsis thaliana]
Length = 436
Score = 73.2 bits (178), Expect = 1e-12
Identities = 36/73 (49%), Positives = 50/73 (68%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++THL ++ A V G+ +G+ I ++LT SD+ LPFK RRQFL+ + FAMTIN
Sbjct: 313 LQITHLCNQIVQARVIIGDTIGEIVLIRTMNLTPSDTKLPFKMRRRQFLLPVAFAMTINK 372
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V LYL
Sbjct: 373 SQGQSLQQVGLYL 385
>UniRef100_Q9LX60 Hypothetical protein F4M19_60 [Arabidopsis thaliana]
Length = 1752
Score = 71.2 bits (173), Expect = 5e-12
Identities = 35/73 (47%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L + ++ A V +++G IP I+LT SD+ LPFK RRQF +++ FAMTIN
Sbjct: 1629 LQITQLAKQVVQAKVITRDRIGDIVLIPLINLTPSDTKLPFKMRRRQFPLSVAFAMTINK 1688
Query: 105 NQGQSLSHVSLYL 117
+QGQSL V LYL
Sbjct: 1689 SQGQSLEQVGLYL 1701
>UniRef100_Q9ZQ61 Putative helicase [Arabidopsis thaliana]
Length = 1230
Score = 71.2 bits (173), Expect = 5e-12
Identities = 33/73 (45%), Positives = 48/73 (65%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++ L ++ + G ++GK IPR+SLT SD LPFK RR F +++ FAMTIN
Sbjct: 1150 LQIVRLGDKLVQGRILTGTRVGKLVIIPRMSLTPSDRRLPFKMKRRHFPLSVAFAMTINK 1209
Query: 105 NQGQSLSHVSLYL 117
+QGQSL +V +YL
Sbjct: 1210 SQGQSLGNVGMYL 1222
>UniRef100_Q9LW42 Helicase-like protein [Arabidopsis thaliana]
Length = 1669
Score = 71.2 bits (173), Expect = 5e-12
Identities = 33/73 (45%), Positives = 48/73 (65%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++ L ++ + G ++GK IPR+ LT SD LPFK RRQF +++ FAMTIN
Sbjct: 1547 LQIVRLGDKLVQGRILTGTRVGKLVIIPRMPLTPSDRRLPFKMKRRQFPLSVAFAMTINK 1606
Query: 105 NQGQSLSHVSLYL 117
+QGQSL +V +YL
Sbjct: 1607 SQGQSLGNVGIYL 1619
>UniRef100_Q6MW89 B1248C03.15 protein [Oryza sativa]
Length = 1550
Score = 70.9 bits (172), Expect = 7e-12
Identities = 38/71 (53%), Positives = 45/71 (62%)
Query: 47 VTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQ 106
VT LT +I + G G AYIPRI TS+ S PFK RRQF + L +AMTIN +Q
Sbjct: 1143 VTQLTNRVIEGEIITGKAKGTKAYIPRIITTSAQSKWPFKLRRRQFPIRLSYAMTINKSQ 1202
Query: 107 GQSLSHVSLYL 117
GQ+LS V LYL
Sbjct: 1203 GQTLSIVGLYL 1213
>UniRef100_Q6MW82 B1340F09.1 protein [Oryza sativa]
Length = 698
Score = 70.9 bits (172), Expect = 7e-12
Identities = 38/71 (53%), Positives = 45/71 (62%)
Query: 47 VTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQ 106
VT LT +I + G G AYIPRI TS+ S PFK RRQF + L +AMTIN +Q
Sbjct: 291 VTQLTNRVIEGEIITGKAKGTKAYIPRIITTSAQSKWPFKLRRRQFPIRLSYAMTINKSQ 350
Query: 107 GQSLSHVSLYL 117
GQ+LS V LYL
Sbjct: 351 GQTLSIVGLYL 361
>UniRef100_Q7XS07 OSJNBa0095H06.12 protein [Oryza sativa]
Length = 1724
Score = 70.5 bits (171), Expect = 9e-12
Identities = 32/64 (50%), Positives = 46/64 (71%)
Query: 54 MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQGQSLSHV 113
+I A + G+ +G+T YIPRI+L++++ PF RRQF V +C+AMTIN +QGQSL V
Sbjct: 1158 VIEARIITGSNIGQTVYIPRITLSANNKKWPFTLQRRQFPVRVCYAMTINKSQGQSLCSV 1217
Query: 114 SLYL 117
+YL
Sbjct: 1218 GIYL 1221
>UniRef100_Q9ZQR0 Putative helicase [Arabidopsis thaliana]
Length = 1265
Score = 70.1 bits (170), Expect = 1e-11
Identities = 32/73 (43%), Positives = 50/73 (67%)
Query: 45 MRVTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINN 104
+++T L ++ A V G+++G IP ++LT +++ LPFK RRQF +++ F MTIN
Sbjct: 1159 LQITQLCTQIVEAKVITGDRIGHIILIPTVNLTPTNTKLPFKMRRRQFPLSVAFVMTINK 1218
Query: 105 NQGQSLSHVSLYL 117
++GQSL HV LYL
Sbjct: 1219 SEGQSLEHVGLYL 1231
>UniRef100_Q5VR06 Helicase-like protein [Oryza sativa]
Length = 1427
Score = 68.9 bits (167), Expect = 3e-11
Identities = 32/71 (45%), Positives = 48/71 (67%)
Query: 47 VTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQ 106
+T+L +I + G +G+ AYIPRI+LT+ + PF RR F + +C++MTIN +Q
Sbjct: 1305 ITNLGDNIIEGIIITGTHIGEKAYIPRINLTTRGNQWPFTLCRRHFPIKVCYSMTINKSQ 1364
Query: 107 GQSLSHVSLYL 117
GQ+LS+V LYL
Sbjct: 1365 GQTLSNVGLYL 1375
>UniRef100_Q84QR0 Helicase-like protein [Oryza sativa]
Length = 1330
Score = 68.6 bits (166), Expect = 3e-11
Identities = 36/71 (50%), Positives = 43/71 (59%)
Query: 47 VTHLTQ*MIVATVFPGNKLGKTAYIPRISLTSSDSGLPFKFSRRQFLVTLCFAMTINNNQ 106
VT LT +I + G G AYIPRI TS+ S PFK RRQF + L +AMTIN +Q
Sbjct: 1205 VTQLTTRIIEGEIMTGKAKGSKAYIPRIITTSAQSKWPFKLKRRQFPIRLSYAMTINKSQ 1264
Query: 107 GQSLSHVSLYL 117
GQ+L V YL
Sbjct: 1265 GQTLQKVGAYL 1275
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.353 0.153 0.503
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 151,298,985
Number of Sequences: 2790947
Number of extensions: 4416167
Number of successful extensions: 17001
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 69
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 16924
Number of HSP's gapped (non-prelim): 76
length of query: 117
length of database: 848,049,833
effective HSP length: 93
effective length of query: 24
effective length of database: 588,491,762
effective search space: 14123802288
effective search space used: 14123802288
T: 11
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0403.8


