

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0391.11
(490 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
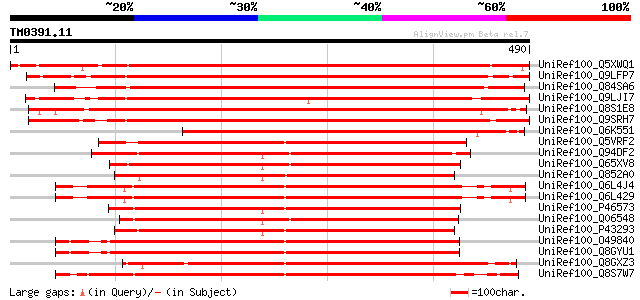
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q5XWQ1 Serine/threonine protein kinase-like [Solanum t... 682 0.0
UniRef100_Q9LFP7 Serine/threonine specific protein kinase-like [... 625 e-178
UniRef100_Q84SA6 Serine/threonine protein kinase [Aster tripolium] 620 e-176
UniRef100_Q9LJI7 Protein kinase [Arabidopsis thaliana] 610 e-173
UniRef100_Q8S1E8 Putative serine/threonine-specific protein kina... 608 e-173
UniRef100_Q9SRH7 Hypothetical protein T22N4.7 [Arabidopsis thali... 606 e-172
UniRef100_Q6K551 Putative serine/threonine protein kinase [Oryza... 490 e-137
UniRef100_Q5VRF2 Putative serine/threonine protein kinase [Oryza... 468 e-130
UniRef100_Q94DF2 Putative serine/threonine-specific protein kina... 454 e-126
UniRef100_Q65XV8 Hypothetical protein P0016H04.10 [Oryza sativa] 451 e-125
UniRef100_Q852A0 Hypothetical protein OSJNBb0081B07.25 [Oryza sa... 435 e-120
UniRef100_Q6L4J4 Hypothetical protein PGEC219.1 [Solanum demissum] 429 e-119
UniRef100_Q6L429 Hypothetical protein PGEC446O19.6 [Solanum demi... 427 e-118
UniRef100_P46573 Protein kinase APK1B, chloroplast precursor [Ar... 427 e-118
UniRef100_Q06548 Protein kinase APK1A, chloroplast precursor [Ar... 424 e-117
UniRef100_P43293 Probable serine/threonine-protein kinase NAK [A... 422 e-116
UniRef100_O49840 Protein kinase [Arabidopsis thaliana] 420 e-116
UniRef100_Q8GYU1 Hypothetical protein At2g02800/T20F6.6 [Arabido... 419 e-115
UniRef100_Q8GXZ3 Hypothetical protein At5g01020/F7J8_5 [Arabidop... 419 e-115
UniRef100_Q8S7W7 Hypothetical protein OSJNBa0091P11.5 [Oryza sat... 417 e-115
>UniRef100_Q5XWQ1 Serine/threonine protein kinase-like [Solanum tuberosum]
Length = 603
Score = 682 bits (1760), Expect = 0.0
Identities = 357/496 (71%), Positives = 403/496 (80%), Gaps = 13/496 (2%)
Query: 1 MGYGDNNAIQAGSLDVDKSKGSKKKVDDGAKKERGWWFRFSFFGSCIPSRPKVDSSISGT 60
MG G + + S DV+KSKG KKK + ++E G W + F GSCI SR KVDSSISG
Sbjct: 113 MGLGGDGE-KGESWDVEKSKGKKKK--EVTEEETGCWTKLWFIGSCISSRSKVDSSISGI 169
Query: 61 STHNGNS---IKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELK 117
STH S + S I +E + + ++ AP SSTTTSN ES +ST KL EELK
Sbjct: 170 STHCDKSTYVLTSCIALAESKSTNTSR--DQPVAPIISSTTTSNAESNSSTSKL-EEELK 226
Query: 118 VASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKIL 177
V+S LRKF FN LK+ATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVK L
Sbjct: 227 VSSRLRKFAFNDLKLATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKTL 286
Query: 178 NHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL 237
NH+G Q W AE+N+LGDL+HPNLVKLIG+CIEDDQRLLVYEFMPRGSLENHLFRR +
Sbjct: 287 NHDGLQVLSIWQAEVNFLGDLVHPNLVKLIGYCIEDDQRLLVYEFMPRGSLENHLFRRSM 346
Query: 238 PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPE 297
PLPWSIRMKIALGAAKGLAFLHE+++RP+IYRDFKTSNILLDA+YNAKLSDFGLAKDGPE
Sbjct: 347 PLPWSIRMKIALGAAKGLAFLHEEAERPVIYRDFKTSNILLDADYNAKLSDFGLAKDGPE 406
Query: 298 GEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHN 357
G+KTHVSTRVMGTYGYAAPEYVMTGHL+SKSDVYSFGVVLLEM+TGRRS+DK RPNGEHN
Sbjct: 407 GDKTHVSTRVMGTYGYAAPEYVMTGHLTSKSDVYSFGVVLLEMITGRRSMDKNRPNGEHN 466
Query: 358 LVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTL 417
LVEWARP LG RR F++++DPRLEGHFS+KGAQKAAQLAA+CLSRDPKARP+MS+VV L
Sbjct: 467 LVEWARPHLGERRRFYRLVDPRLEGHFSIKGAQKAAQLAARCLSRDPKARPMMSDVVEAL 526
Query: 418 KPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLPKKGQP-LRILSSPNCPNG 476
KPLPNLKDMA SSY+FQT + DR S P+ KNG+RTQ S + GQ R LS PN +
Sbjct: 527 KPLPNLKDMASSSYYFQTMQADRVGSSPSTKNGVRTQ-GSFSRNGQQHPRSLSIPNGSHA 585
Query: 477 SPYSRY--SKSPKPVG 490
SPY + SPKP G
Sbjct: 586 SPYHQQFPQNSPKPNG 601
>UniRef100_Q9LFP7 Serine/threonine specific protein kinase-like [Arabidopsis
thaliana]
Length = 493
Score = 625 bits (1612), Expect = e-178
Identities = 322/475 (67%), Positives = 376/475 (78%), Gaps = 16/475 (3%)
Query: 17 DKSKGSKKKVDDGAKKERGWWFRFSFFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSE 76
D +K ++ ++G + G W +F F CIPS+ +D+S S S + N + +
Sbjct: 31 DDNKSRNEEEEEG--EASGCWVKFRFMIGCIPSKSDLDASSS--SIYGSNCTVTTM---- 82
Query: 77 ENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRN 136
E+KS EK+ ++ S+TTTSN ES +STP + SEEL ++S LRKFTFN LK++TRN
Sbjct: 83 ESKSANEKSNDQPVGQVSSTTTTSNAESSSSTP-VISEELNISSHLRKFTFNDLKLSTRN 141
Query: 137 FRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLG 196
FRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVK LN +G QGHKEWLAE+N+LG
Sbjct: 142 FRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKTLNPDGLQGHKEWLAEINFLG 201
Query: 197 DLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLA 256
+LLHPNLVKL+G+CIEDDQRLLVYEFMPRGSLENHLFRR LPLPWSIRMKIALGAAKGL+
Sbjct: 202 NLLHPNLVKLVGYCIEDDQRLLVYEFMPRGSLENHLFRRSLPLPWSIRMKIALGAAKGLS 261
Query: 257 FLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAP 316
FLHE++ +P+IYRDFKTSNILLDA+YNAKLSDFGLAKD P+ KTHVSTRVMGTYGYAAP
Sbjct: 262 FLHEEALKPVIYRDFKTSNILLDADYNAKLSDFGLAKDAPDEGKTHVSTRVMGTYGYAAP 321
Query: 317 EYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQII 376
EYVMTGHL+SKSDVYSFGVVLLEMLTGRRS+DK RPNGEHNLVEWARP L +R F++++
Sbjct: 322 EYVMTGHLTSKSDVYSFGVVLLEMLTGRRSMDKNRPNGEHNLVEWARPHLLDKRRFYRLL 381
Query: 377 DPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTA 436
DPRLEGHFS+KGAQK QLAAQCLSRDPK RP MS+VV LKPLP+LKDMA SSY+FQT
Sbjct: 382 DPRLEGHFSIKGAQKVTQLAAQCLSRDPKIRPKMSDVVEALKPLPHLKDMASSSYYFQTM 441
Query: 437 RVDRTMSMPNHKNGIRTQLVSLPKKGQPL-RILSSPNCPNGSPYSRYSKSPKPVG 490
+ +R + G S + QP+ R LSSP+ SPY SPKP G
Sbjct: 442 QAERLKNGSGRSQGFG----SRNGQHQPVFRTLSSPH--GSSPYRHQIPSPKPKG 490
>UniRef100_Q84SA6 Serine/threonine protein kinase [Aster tripolium]
Length = 439
Score = 620 bits (1600), Expect = e-176
Identities = 321/446 (71%), Positives = 361/446 (79%), Gaps = 26/446 (5%)
Query: 43 FGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNE 102
FGSC+ SR KVDSS SG S+H FE + + + +T+ S
Sbjct: 14 FGSCVSSRSKVDSSTSGISSH------------------FEIKSTNNVSKDQPTTSNSEH 55
Query: 103 ESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTA 162
TP+ +ELKVAS LRKF FN LK+ATRNFRPESLLGEGGFGCVFKGWIEENGTA
Sbjct: 56 NLPTLTPE---DELKVASRLRKFGFNDLKMATRNFRPESLLGEGGFGCVFKGWIEENGTA 112
Query: 163 PVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEF 222
PVKPGTGLTVAVK LNH+G QGHKEWLAE+N+LGDL +PNLVKLIG+CIEDDQRLLVYEF
Sbjct: 113 PVKPGTGLTVAVKTLNHDGLQGHKEWLAEVNFLGDLGNPNLVKLIGYCIEDDQRLLVYEF 172
Query: 223 MPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEY 282
+PRGSLENHLFRR LPLPWSIRMKIALGAAKGLAFLHE+++RP+IYRDFKTSNILLDAEY
Sbjct: 173 LPRGSLENHLFRRSLPLPWSIRMKIALGAAKGLAFLHEEAKRPVIYRDFKTSNILLDAEY 232
Query: 283 NAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLT 342
NAKLSDFGLAKDGPEG+KTH+STRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLT
Sbjct: 233 NAKLSDFGLAKDGPEGDKTHISTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLT 292
Query: 343 GRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSR 402
GRRS+DKKRPNGEHNLVEWARP LG RR F+++IDPRLEGHFS+KGAQKAAQLA++CLSR
Sbjct: 293 GRRSMDKKRPNGEHNLVEWARPHLGERRRFYRLIDPRLEGHFSIKGAQKAAQLASRCLSR 352
Query: 403 DPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLPKKG 462
DPKARPLMSEVV LKPLP LKDMA SY+ QT + +R S P+ RT++ S + G
Sbjct: 353 DPKARPLMSEVVDCLKPLPALKDMAGPSYYLQTVQPERAGSSPDPN---RTRVGSFSRNG 409
Query: 463 -QPLRILSSPNC-PNGSPYSRYSKSP 486
Q R LS PN P + + + S +P
Sbjct: 410 SQHPRTLSIPNASPRHNQFLQDSPNP 435
>UniRef100_Q9LJI7 Protein kinase [Arabidopsis thaliana]
Length = 483
Score = 610 bits (1572), Expect = e-173
Identities = 319/483 (66%), Positives = 371/483 (76%), Gaps = 35/483 (7%)
Query: 16 VDKSKGSKKKVDDGAKKERGWWFRFSFFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQS 75
+ K K KK DG + E G+WFRF F SCI SR KVDSS++ T+ VI +
Sbjct: 25 MSKKKNVKK---DGDESESGFWFRFKFIFSCISSRSKVDSSMNATA---------VIAEP 72
Query: 76 EENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATR 135
+ K E+ E P + T ES +STP L S ELK +S LR F FN LK+ATR
Sbjct: 73 K-------KVIEKLEGHPAPTKDTGCAESGSSTP-LMSGELKYSSKLRIFMFNDLKLATR 124
Query: 136 NFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYL 195
NFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVK LN +G QGHKEWLAE+N+L
Sbjct: 125 NFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKTLNPDGLQGHKEWLAEINFL 184
Query: 196 GDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGL 255
G+L+HP+LVKL+G+C+E+DQRLLVYEFMPRGSLENHLFRR LPLPWS+RMKIALGAAKGL
Sbjct: 185 GNLVHPSLVKLVGYCMEEDQRLLVYEFMPRGSLENHLFRRTLPLPWSVRMKIALGAAKGL 244
Query: 256 AFLHEDSQRPIIYRDFKTSNILLDA-------EYNAKLSDFGLAKDGPEGEKTHVSTRVM 308
AFLHE++++P+IYRDFKTSNILLD EYNAKLSDFGLAKD P+ +K+HVSTRVM
Sbjct: 245 AFLHEEAEKPVIYRDFKTSNILLDGKKMLPLQEYNAKLSDFGLAKDAPDEKKSHVSTRVM 304
Query: 309 GTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGH 368
GTYGYAAPEYVMTGHL++KSDVYSFGVVLLE+LTGRRS+DK RPNGE NLVEW RP L
Sbjct: 305 GTYGYAAPEYVMTGHLTTKSDVYSFGVVLLEILTGRRSVDKSRPNGEQNLVEWVRPHLLD 364
Query: 369 RRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAI 428
++ F++++DPRLEGH+S+KGAQKA Q+AAQCL+RD KARP MSEVV LKPLPNLKD A
Sbjct: 365 KKRFYRLLDPRLEGHYSIKGAQKATQVAAQCLNRDSKARPKMSEVVEALKPLPNLKDFAS 424
Query: 429 SSYHFQTARVDRTMSMPNHKNGIRTQLVS-LPKKGQPLRILSSPNCPNGSPYSRYSKSPK 487
SS FQT + P KNG+RTQ + + G P+R LSS N P SPY +SPK
Sbjct: 425 SSSSFQTMQ-------PVAKNGVRTQGGGFVSRNGPPMRSLSSLNLPQASPYRYARQSPK 477
Query: 488 PVG 490
P G
Sbjct: 478 PKG 480
>UniRef100_Q8S1E8 Putative serine/threonine-specific protein kinase NAK [Oryza
sativa]
Length = 494
Score = 608 bits (1569), Expect = e-173
Identities = 313/479 (65%), Positives = 370/479 (76%), Gaps = 17/479 (3%)
Query: 18 KSKGSKKKV----DDGAKKERGWWFRFS--FFGSCIPSRPKVDSSISGTSTHNGNSIKSV 71
K KG+ + V ++ A G W R G C+ SR KVDSS + + G S +
Sbjct: 19 KGKGAGEMVLQQEEEDAAPAMGCWIRIPRRLGGGCMSSRSKVDSSTTTSGGGGGGSARV- 77
Query: 72 ITQSEENKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLK 131
E+KS + + + P S +TTS+ S + EELK+A LR+FTFN LK
Sbjct: 78 ---GGESKSANDGCRDHSVQPMASGSTTSSNTGSISPSSIVGEELKLAFQLRRFTFNELK 134
Query: 132 VATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAE 191
ATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVK LNH+G QGHKEW+AE
Sbjct: 135 CATRNFRPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKTLNHDGLQGHKEWVAE 194
Query: 192 LNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGA 251
+++LG+L HP+LVKL+G+CIEDDQRLLVYEFMPRGSLENHLFRR LPLPW+IRM+IALGA
Sbjct: 195 VDFLGNLQHPHLVKLVGYCIEDDQRLLVYEFMPRGSLENHLFRRSLPLPWAIRMRIALGA 254
Query: 252 AKGLAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTY 311
AKGLAFLHE+++RP+IYRDFKTSNILLDA+YNAKLSDFGLAKDGPEG+KTHVSTRVMGTY
Sbjct: 255 AKGLAFLHEEAERPVIYRDFKTSNILLDADYNAKLSDFGLAKDGPEGDKTHVSTRVMGTY 314
Query: 312 GYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRM 371
GYAAPEYVMTGHL+SKSDVYSFGVVLLEM++GRRS+DK RPNGEHNLVEWARP LG RR
Sbjct: 315 GYAAPEYVMTGHLTSKSDVYSFGVVLLEMMSGRRSMDKNRPNGEHNLVEWARPYLGERRR 374
Query: 372 FFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSY 431
F++++DPRLEG+FS++GAQK AQLA CL+RDPKARPLMS+VV LKPL NLKDMA SSY
Sbjct: 375 FYRLVDPRLEGNFSIRGAQKTAQLACACLNRDPKARPLMSQVVEVLKPLLNLKDMASSSY 434
Query: 432 HFQTARVDRTMSM--PNHKNGIRTQLVSLPKKGQPLRILSSPNCPNGSPYSRYSKSPKP 488
FQ+ + +R S+ P ++ Q QP+R LS P+ SP Y +SP+P
Sbjct: 435 FFQSMQQERAASLGNPIGSQSMKAQGTFARNGQQPMRSLSYG--PHASP---YRQSPRP 488
>UniRef100_Q9SRH7 Hypothetical protein T22N4.7 [Arabidopsis thaliana]
Length = 490
Score = 606 bits (1562), Expect = e-172
Identities = 315/475 (66%), Positives = 367/475 (76%), Gaps = 19/475 (4%)
Query: 18 KSKGSKKKVDDGAKKERGWWFRFSFFGSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEE 77
K + + + ++ G W +F + C S V++S++ +++ G+ +S I QS +
Sbjct: 30 KKNNNVRNSEHYEEEANGCWVKFRYIVCCASSTSDVETSLTLSTSTVGS--QSAIVQSND 87
Query: 78 NKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNF 137
G P S+TTTSN ES STP + SEEL + S L+KF+F LK+ATRNF
Sbjct: 88 QPVG----------PVSSTTTTSNAESSLSTP-IISEELNIYSHLKKFSFIDLKLATRNF 136
Query: 138 RPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGD 197
RPESLLGEGGFGCVFKGW+EENGTAPVKPGTGLTVAVK LN +G QGHKEWLAE+NYLG+
Sbjct: 137 RPESLLGEGGFGCVFKGWVEENGTAPVKPGTGLTVAVKTLNPDGLQGHKEWLAEINYLGN 196
Query: 198 LLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAF 257
LLHPNLVKL+G+CIEDDQRLLVYEFMPRGSLENHLFRR LPLPWSIRMKIALGAAKGL+F
Sbjct: 197 LLHPNLVKLVGYCIEDDQRLLVYEFMPRGSLENHLFRRSLPLPWSIRMKIALGAAKGLSF 256
Query: 258 LHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPE 317
LHE++ +P+IYRDFKTSNILLD EYNAKLSDFGLAKD P+ KTHVSTRVMGTYGYAAPE
Sbjct: 257 LHEEALKPVIYRDFKTSNILLDGEYNAKLSDFGLAKDAPDEGKTHVSTRVMGTYGYAAPE 316
Query: 318 YVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIID 377
YVMTGHL+SKSDVYSFGVVLLEMLTGRRS+DK RPNGEHNLVEWARP L +R F++++D
Sbjct: 317 YVMTGHLTSKSDVYSFGVVLLEMLTGRRSMDKNRPNGEHNLVEWARPHLLDKRRFYRLLD 376
Query: 378 PRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTAR 437
PRLEGHFSVKGAQK QLAAQCLSRD K RP MSEVV LKPLP+LKDMA +SY+FQT +
Sbjct: 377 PRLEGHFSVKGAQKVTQLAAQCLSRDSKIRPKMSEVVEVLKPLPHLKDMASASYYFQTMQ 436
Query: 438 VDRTMSMPNHKNGIRTQLVSLPKKGQPL-RILSSPNCPNG-SPYSRYSKSPKPVG 490
+R + +G + GQP+ R LSSP+ G SPY SPKP G
Sbjct: 437 AERLKAGSGSGSGRGFG----SRNGQPVFRTLSSPHGQAGSSPYRHQIPSPKPKG 487
>UniRef100_Q6K551 Putative serine/threonine protein kinase [Oryza sativa]
Length = 326
Score = 490 bits (1262), Expect = e-137
Identities = 242/326 (74%), Positives = 282/326 (86%), Gaps = 6/326 (1%)
Query: 164 VKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFM 223
+KPGTGLTVAVK LNH+G QGHKEW+AE+++LG+L HPNLV+LIG+C+EDDQRLLVYEFM
Sbjct: 1 MKPGTGLTVAVKTLNHDGLQGHKEWVAEVDFLGNLHHPNLVRLIGYCVEDDQRLLVYEFM 60
Query: 224 PRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEYN 283
PRGSL+NHLFRR LPLPWSIRMK+ALGAAKGLAFLHE+++RP+IYRDFKTSNILLDA+YN
Sbjct: 61 PRGSLDNHLFRRSLPLPWSIRMKVALGAAKGLAFLHEEAERPVIYRDFKTSNILLDADYN 120
Query: 284 AKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLTG 343
AKLSDFGLAKDGP G+KTHVSTRVMGTYGYAAPEYVMTGHL+SKSDVYSFGVVLLEM++G
Sbjct: 121 AKLSDFGLAKDGPVGDKTHVSTRVMGTYGYAAPEYVMTGHLTSKSDVYSFGVVLLEMMSG 180
Query: 344 RRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSRD 403
RRS+DK RPNGEHNLVEWARP+LG R+ F+++IDPRLEG+FSVKGAQKAAQLA CL+RD
Sbjct: 181 RRSMDKNRPNGEHNLVEWARPLLGERQRFYKLIDPRLEGNFSVKGAQKAAQLARACLNRD 240
Query: 404 PKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDR---TMSMPNHKNGIRTQLVSLPK 460
PKARPLMS+VV LKPL NLKDMA SSY +QT + +R + SM + ++ Q
Sbjct: 241 PKARPLMSQVVEVLKPLLNLKDMASSSYFYQTMQAERMAHSSSMNGRSHALKVQGSFARN 300
Query: 461 KGQPLRILSSPNCPNGSPYSRYSKSP 486
QP+R LS + P SP+ RYS P
Sbjct: 301 GQQPMRSLS--DGPRASPF-RYSPKP 323
>UniRef100_Q5VRF2 Putative serine/threonine protein kinase [Oryza sativa]
Length = 411
Score = 468 bits (1203), Expect = e-130
Identities = 238/348 (68%), Positives = 270/348 (77%), Gaps = 12/348 (3%)
Query: 85 NTEETEAPPESSTTTSNEESI-ASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLL 143
N E T +E + AST K L +FTF LK AT NFRP+S+L
Sbjct: 63 NASNRELGDHFQTNLDDENGVNASTEK----------KLLRFTFQELKSATVNFRPDSIL 112
Query: 144 GEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNL 203
GEGGFG VFKGWI+ N T+P KPGTGLTVAVK L + QGH+EW+AE+++LG L H +L
Sbjct: 113 GEGGFGYVFKGWIDPNSTSPAKPGTGLTVAVKSLKQDALQGHREWVAEVDFLGQLHHKHL 172
Query: 204 VKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPLPLPWSIRMKIALGAAKGLAFLHEDSQ 263
VKLIG+CIEDDQRLLVYEFM RGSLENHLFRR LPLPW RMKIALGAAKGLAFLH
Sbjct: 173 VKLIGYCIEDDQRLLVYEFMARGSLENHLFRRALPLPWPCRMKIALGAAKGLAFLH-GGP 231
Query: 264 RPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGH 323
+P+IYRDFKTSNILLDAEYNAKLSDFGLAK GP+G+KTHVSTRV+GTYGYAAPEYVMTGH
Sbjct: 232 KPVIYRDFKTSNILLDAEYNAKLSDFGLAKAGPQGDKTHVSTRVVGTYGYAAPEYVMTGH 291
Query: 324 LSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGH 383
L+SKSDVYSFGVVLLEMLTGRRS+DKKRP GE NLV WARP L RR +Q++DPRL +
Sbjct: 292 LTSKSDVYSFGVVLLEMLTGRRSMDKKRPTGEQNLVAWARPYLSDRRRLYQLVDPRLGLN 351
Query: 384 FSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSY 431
+SV+G QK AQ+ CLSRD K+RP M EVV L PL +L DMA +SY
Sbjct: 352 YSVRGVQKVAQICYHCLSRDTKSRPTMDEVVKHLTPLQDLNDMASASY 399
>UniRef100_Q94DF2 Putative serine/threonine-specific protein kinase [Oryza sativa]
Length = 467
Score = 454 bits (1167), Expect = e-126
Identities = 225/361 (62%), Positives = 277/361 (76%), Gaps = 8/361 (2%)
Query: 78 NKSGFEKNTEETEAPPESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNF 137
N SG + + ++ + A TP+ E L+ A+ +R F FN LK ATRNF
Sbjct: 76 NASGSRRRRSGSRRKGTGLSSRVSSSVAAVTPRSEGEILRCAN-VRSFAFNELKTATRNF 134
Query: 138 RPESLLGEGGFGCVFKGWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGD 197
RP+S+LGEGGFG VFKGW++EN P +PGTG+ +AVK LN +G QGH+EWLAE+NYLG
Sbjct: 135 RPDSVLGEGGFGSVFKGWVDENTFLPSRPGTGMVIAVKKLNQDGFQGHREWLAEVNYLGQ 194
Query: 198 LLHPNLVKLIGFCIEDDQRLLVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKG 254
L HPNLVKL+G+C++D+QRLLVYEFMPRGSLENHLFRR PL W++RMK+ALGAAKG
Sbjct: 195 LSHPNLVKLVGYCLQDEQRLLVYEFMPRGSLENHLFRRGSHFQPLSWNLRMKVALGAAKG 254
Query: 255 LAFLHEDSQRPIIYRDFKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYA 314
LAFLH D + +IYRDFKTSN+LLD+ YNAKLSDFGLAKDGP G+K+HVSTRVMGTYGYA
Sbjct: 255 LAFLHSDKAK-VIYRDFKTSNVLLDSNYNAKLSDFGLAKDGPTGDKSHVSTRVMGTYGYA 313
Query: 315 APEYVMTGHLSSKSDVYSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQ 374
APEY+ TGHLS+KSDVYSFGVV++EML+GRR++DK RP GEHNLVEWARP L RR F+
Sbjct: 314 APEYLATGHLSAKSDVYSFGVVMVEMLSGRRALDKNRPAGEHNLVEWARPYLSSRRRIFR 373
Query: 375 IIDPRLEGHFSVKGAQKAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQ 434
I+D RL G +S+ GA KAA LA QCLS D K RP M +VV L+ L++ +S+H +
Sbjct: 374 ILDARLAGQYSLAGAHKAAALALQCLSADAKNRPTMHQVVAALE---QLQETTTTSHHHR 430
Query: 435 T 435
+
Sbjct: 431 S 431
>UniRef100_Q65XV8 Hypothetical protein P0016H04.10 [Oryza sativa]
Length = 395
Score = 451 bits (1159), Expect = e-125
Identities = 220/334 (65%), Positives = 270/334 (79%), Gaps = 5/334 (1%)
Query: 95 SSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKG 154
+S ++ + S+ TP+ +E+ A++++ F FN L+ ATRNFRP+S+LGEGGFG VFKG
Sbjct: 29 ASNSSVSAASVPPTPRS-EDEILEAANVKAFAFNELRTATRNFRPDSVLGEGGFGSVFKG 87
Query: 155 WIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDD 214
WI+E AP KPGTG+ +AVK LN GHQGH+EWLAE+NYLG L HP LV+L+G+C+ED+
Sbjct: 88 WIDEKTLAPTKPGTGMVIAVKKLNQEGHQGHREWLAEVNYLGQLSHPYLVRLVGYCVEDE 147
Query: 215 QRLLVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDF 271
QRLLVYEFMPRGSLENHLFRR PL W++RMKIALGAAKGLAFLH D + +IYRDF
Sbjct: 148 QRLLVYEFMPRGSLENHLFRRSTHFQPLSWNLRMKIALGAAKGLAFLHSDKVK-VIYRDF 206
Query: 272 KTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVY 331
KTSN+LLDA Y+AKLSDFGLAKDGP G+K+HVSTRVMGTYGYAAPEY+ TGHL++KSDVY
Sbjct: 207 KTSNVLLDANYDAKLSDFGLAKDGPTGDKSHVSTRVMGTYGYAAPEYLATGHLTTKSDVY 266
Query: 332 SFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQK 391
SFGVVLLEML+GRR++DK RP GEHNLVEWARP L +R F+I+D RL G +S+ AQK
Sbjct: 267 SFGVVLLEMLSGRRALDKNRPTGEHNLVEWARPYLMSKRRIFRILDARLGGQYSLAKAQK 326
Query: 392 AAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKD 425
AA LA QC+S + K RP M +VV L+ L + K+
Sbjct: 327 AATLALQCISVEAKNRPNMEQVVAVLEQLQDSKE 360
>UniRef100_Q852A0 Hypothetical protein OSJNBb0081B07.25 [Oryza sativa]
Length = 416
Score = 435 bits (1119), Expect = e-120
Identities = 215/326 (65%), Positives = 258/326 (78%), Gaps = 6/326 (1%)
Query: 100 SNEESIASTPKLFSEELKVASS--LRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIE 157
S+ S AS P E ++ S ++ F+F+ L++ATRNFRP+S+LGEGGFG V+KGWI+
Sbjct: 39 SSRASSASMPPTAKTECEILQSANVKIFSFSDLRIATRNFRPDSVLGEGGFGSVYKGWID 98
Query: 158 ENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRL 217
EN + KPGTG+ VAVK LN QGH+EWLAE+NYLG HPNLVKL G+C+ED+ RL
Sbjct: 99 ENTLSACKPGTGIAVAVKRLNQESLQGHREWLAEVNYLGQFCHPNLVKLFGYCLEDEHRL 158
Query: 218 LVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTS 274
LVYEFMPRGSLENHLFRR PL W++RMK+ALGAAKGLA+LH S+ +IYRDFKTS
Sbjct: 159 LVYEFMPRGSLENHLFRRGSHFQPLSWNLRMKVALGAAKGLAYLHS-SEAKVIYRDFKTS 217
Query: 275 NILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFG 334
NILLD +Y+AKLSDFGLAKDGP GEK+HVSTRVMGTYGYAAPEY+ TGHL++KSDVYSFG
Sbjct: 218 NILLDTDYSAKLSDFGLAKDGPVGEKSHVSTRVMGTYGYAAPEYLSTGHLTAKSDVYSFG 277
Query: 335 VVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQ 394
VVLLEM++GRR+IDK RP GEHNLVEWARP L H+R F+++D RLEG +S GAQ A
Sbjct: 278 VVLLEMMSGRRAIDKNRPQGEHNLVEWARPYLTHKRKIFRVLDTRLEGQYSHVGAQTVAT 337
Query: 395 LAAQCLSRDPKARPLMSEVVHTLKPL 420
LA +CLS + K RP M VV L+ L
Sbjct: 338 LALECLSYEAKMRPSMEAVVTILEEL 363
>UniRef100_Q6L4J4 Hypothetical protein PGEC219.1 [Solanum demissum]
Length = 420
Score = 429 bits (1103), Expect = e-119
Identities = 232/450 (51%), Positives = 298/450 (65%), Gaps = 39/450 (8%)
Query: 44 GSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEE 103
G+C+ S +V++++S T T S S F + P S + +
Sbjct: 2 GNCVGSSARVEATLSST------------TPSAYEASRFPDRKSNSSVPSSLSIPSYGRK 49
Query: 104 SIAS---TPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENG 160
S + TP+ SE L + +++ F+FN LK ATRNFRP+SLLGEGGFGCVFKGWI+
Sbjct: 50 SSSESLPTPRSESEIL-FSPNVKSFSFNELKNATRNFRPDSLLGEGGFGCVFKGWIDAQT 108
Query: 161 TAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVY 220
KPG+G+ +AVK L G QGHKEWL E+NYLG L HPNLVKLIG+CI+ D LLVY
Sbjct: 109 LTASKPGSGIVIAVKKLKPEGFQGHKEWLTEVNYLGQLRHPNLVKLIGYCIDGDNHLLVY 168
Query: 221 EFMPRGSLENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLD 279
EFMP+GSLENHLFRR P PL W+ R+K+A+GAA+GLAFLH D++ +IYRDFK SNILLD
Sbjct: 169 EFMPKGSLENHLFRRGPQPLNWATRIKVAIGAARGLAFLH-DAKEQVIYRDFKASNILLD 227
Query: 280 AEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLE 339
AE+N+KLSDFGLAK GP G++THVST+VMGT GYAAPEYV TG L++KSDVYSFGVVLLE
Sbjct: 228 AEFNSKLSDFGLAKAGPTGDRTHVSTQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLE 287
Query: 340 MLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQC 399
+L+GRR++D + E NLV+WA+P LG +R F+I+D +LEG + KGA AA LA QC
Sbjct: 288 LLSGRRAVDNTKVGIEQNLVDWAKPYLGDKRKLFRIMDTKLEGQYPQKGAYTAANLAWQC 347
Query: 400 LSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLP 459
LS +PK RP MSEV+ L+ L + K ++ +S H+ + P
Sbjct: 348 LSNEPKLRPKMSEVLTALEELQSPKGVS-------------KLSHTEHR------AIPSP 388
Query: 460 KKGQPLRILSSP--NCPNGSPYSRYSKSPK 487
P+R SP P+ SP+ Y KSP+
Sbjct: 389 VAVSPMRHHRSPLHMTPSASPFQAYQKSPR 418
>UniRef100_Q6L429 Hypothetical protein PGEC446O19.6 [Solanum demissum]
Length = 420
Score = 427 bits (1099), Expect = e-118
Identities = 232/450 (51%), Positives = 297/450 (65%), Gaps = 39/450 (8%)
Query: 44 GSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEE 103
G+C+ S +V++++S T T S S F + P S + +
Sbjct: 2 GNCVGSSARVEATLSST------------TPSAYEASRFPDRKSNSSVPSSLSIPSYGRK 49
Query: 104 SIAS---TPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENG 160
S + TP+ SE L + +++ F+FN LK ATRNFRP+SLLGEGGFGCVFKGWI+
Sbjct: 50 SSSESLPTPRSESEIL-FSPNVKSFSFNELKNATRNFRPDSLLGEGGFGCVFKGWIDAQT 108
Query: 161 TAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVY 220
KPG+G+ +AVK L G QGHKEWL E+NYLG L HPNLVKLIG+CI+ D LLVY
Sbjct: 109 LTASKPGSGIVIAVKKLKPEGFQGHKEWLTEVNYLGQLRHPNLVKLIGYCIDGDNHLLVY 168
Query: 221 EFMPRGSLENHLFRR-PLPLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLD 279
EFMP+GSLENHLFRR P PL W+ R+K+A+GAA+GLAFLH D++ +IYRDFK SNILLD
Sbjct: 169 EFMPKGSLENHLFRRGPQPLNWATRIKVAIGAARGLAFLH-DAKEQVIYRDFKASNILLD 227
Query: 280 AEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLE 339
AE+N+KLSDFGLAK GP G++THVST+VMGT GYAAPEYV TG L++KSDVYSFGVVLLE
Sbjct: 228 AEFNSKLSDFGLAKAGPTGDRTHVSTQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLE 287
Query: 340 MLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQC 399
+L+GRR++D + E NLV+WA+P LG +R F+I+D +LEG + KGA AA LA QC
Sbjct: 288 LLSGRRAVDNAKVGIEQNLVDWAKPYLGDKRKLFRIMDTKLEGQYPQKGAYTAANLAWQC 347
Query: 400 LSRDPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLP 459
LS +PK RP MSEV+ L+ L + K ++ +S H+ + P
Sbjct: 348 LSNEPKLRPKMSEVLTALEELQSPKGVS-------------KLSHTEHR------AIPSP 388
Query: 460 KKGQPLRILSSP--NCPNGSPYSRYSKSPK 487
P+R SP P+ SP Y KSP+
Sbjct: 389 VAVSPMRHHRSPLHMTPSASPLQAYQKSPR 418
>UniRef100_P46573 Protein kinase APK1B, chloroplast precursor [Arabidopsis thaliana]
Length = 412
Score = 427 bits (1097), Expect = e-118
Identities = 211/335 (62%), Positives = 267/335 (78%), Gaps = 5/335 (1%)
Query: 94 ESSTTTSNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFK 153
+S + S+ SI + P+ E L+ + +L+ FTF LK ATRNFRP+S+LGEGGFG VFK
Sbjct: 27 DSLGSKSSSVSIRTNPRTEGEILQ-SPNLKSFTFAELKAATRNFRPDSVLGEGGFGSVFK 85
Query: 154 GWIEENGTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIED 213
GWI+E KPGTG+ +AVK LN +G QGH+EWLAE+NYLG HPNLVKLIG+C+ED
Sbjct: 86 GWIDEQTLTASKPGTGVVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLED 145
Query: 214 DQRLLVYEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRD 270
+ RLLVYEFMPRGSLENHLFRR PL W++R+K+ALGAAKGLAFLH +++ +IYRD
Sbjct: 146 EHRLLVYEFMPRGSLENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLH-NAETSVIYRD 204
Query: 271 FKTSNILLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDV 330
FKTSNILLD+EYNAKLSDFGLAKDGP G+K+HVSTR+MGTYGYAAPEY+ TGHL++KSDV
Sbjct: 205 FKTSNILLDSEYNAKLSDFGLAKDGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDV 264
Query: 331 YSFGVVLLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQ 390
YS+GVVLLE+L+GRR++DK RP GE LVEWARP+L ++R F++ID RL+ +S++ A
Sbjct: 265 YSYGVVLLEVLSGRRAVDKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEAC 324
Query: 391 KAAQLAAQCLSRDPKARPLMSEVVHTLKPLPNLKD 425
K A LA +CL+ + K RP M+EVV L+ + L +
Sbjct: 325 KVATLALRCLTFEIKLRPNMNEVVSHLEHIQTLNE 359
>UniRef100_Q06548 Protein kinase APK1A, chloroplast precursor [Arabidopsis thaliana]
Length = 410
Score = 424 bits (1089), Expect = e-117
Identities = 209/323 (64%), Positives = 261/323 (80%), Gaps = 5/323 (1%)
Query: 104 SIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAP 163
S+ +P+ E L+ + +L+ F+F LK ATRNFRP+S+LGEGGFGCVFKGWI+E
Sbjct: 36 SVRPSPRTEGEILQ-SPNLKSFSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTA 94
Query: 164 VKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFM 223
+PGTGL +AVK LN +G QGH+EWLAE+NYLG H +LVKLIG+C+ED+ RLLVYEFM
Sbjct: 95 SRPGTGLVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFM 154
Query: 224 PRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDA 280
PRGSLENHLFRR L PL W +R+K+ALGAAKGLAFLH R +IYRDFKTSNILLD+
Sbjct: 155 PRGSLENHLFRRGLYFQPLSWKLRLKVALGAAKGLAFLHSSETR-VIYRDFKTSNILLDS 213
Query: 281 EYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEM 340
EYNAKLSDFGLAKDGP G+K+HVSTRVMGT+GYAAPEY+ TGHL++KSDVYSFGVVLLE+
Sbjct: 214 EYNAKLSDFGLAKDGPIGDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLEL 273
Query: 341 LTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCL 400
L+GRR++DK RP+GE NLVEWA+P L ++R F++ID RL+ +S++ A K A L+ +CL
Sbjct: 274 LSGRRAVDKNRPSGERNLVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCL 333
Query: 401 SRDPKARPLMSEVVHTLKPLPNL 423
+ + K RP MSEVV L+ + +L
Sbjct: 334 TTEIKLRPNMSEVVSHLEHIQSL 356
>UniRef100_P43293 Probable serine/threonine-protein kinase NAK [Arabidopsis thaliana]
Length = 389
Score = 422 bits (1084), Expect = e-116
Identities = 207/324 (63%), Positives = 260/324 (79%), Gaps = 5/324 (1%)
Query: 100 SNEESIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEEN 159
S+ S + P+ E L+ A+ L+ F+ + LK ATRNFRP+S++GEGGFGCVFKGWI+E+
Sbjct: 32 SSTASFSYMPRTEGEILQNAN-LKNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDES 90
Query: 160 GTAPVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLV 219
AP KPGTG+ +AVK LN G QGH+EWLAE+NYLG L HPNLVKLIG+C+E++ RLLV
Sbjct: 91 SLAPSKPGTGIVIAVKRLNQEGFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLV 150
Query: 220 YEFMPRGSLENHLFRRPL---PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNI 276
YEFM RGSLENHLFRR PL W+ R+++ALGAA+GLAFLH ++Q +IYRDFK SNI
Sbjct: 151 YEFMTRGSLENHLFRRGTFYQPLSWNTRVRMALGAARGLAFLH-NAQPQVIYRDFKASNI 209
Query: 277 LLDAEYNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVV 336
LLD+ YNAKLSDFGLA+DGP G+ +HVSTRVMGT GYAAPEY+ TGHLS KSDVYSFGVV
Sbjct: 210 LLDSNYNAKLSDFGLARDGPMGDNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVV 269
Query: 337 LLEMLTGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLA 396
LLE+L+GRR+IDK +P GEHNLV+WARP L ++R +++DPRL+G +S+ A K A LA
Sbjct: 270 LLELLSGRRAIDKNQPVGEHNLVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLA 329
Query: 397 AQCLSRDPKARPLMSEVVHTLKPL 420
C+S D K+RP M+E+V T++ L
Sbjct: 330 LDCISIDAKSRPTMNEIVKTMEEL 353
>UniRef100_O49840 Protein kinase [Arabidopsis thaliana]
Length = 426
Score = 420 bits (1079), Expect = e-116
Identities = 222/383 (57%), Positives = 277/383 (71%), Gaps = 16/383 (4%)
Query: 44 GSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEE 103
G+C+ S KVDSS S + N S+ S ++ T + P SS + ++
Sbjct: 2 GNCLDSSAKVDSS-SHSPHANSASLSSRVSSK----------TSRSTVP--SSLSINSYS 48
Query: 104 SIASTPKLFSE-ELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTA 162
S+ S P +E E+ + +L+ FTFN LK ATRNFRP+SLLGEGGFG VFKGWI+
Sbjct: 49 SVESLPTPRTEGEILSSPNLKAFTFNELKNATRNFRPDSLLGEGGFGYVFKGWIDGTTLT 108
Query: 163 PVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEF 222
KPG+G+ VAVK L G+QGHKEWL E+NYLG L HPNLVKL+G+C+E + RLLVYEF
Sbjct: 109 ASKPGSGIVVAVKKLKTEGYQGHKEWLTEVNYLGQLSHPNLVKLVGYCVEGENRLLVYEF 168
Query: 223 MPRGSLENHLFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAE 281
MP+GSLENHLFRR PL W+IRMK+A+GAAKGL FLH D++ +IYRDFK +NILLDAE
Sbjct: 169 MPKGSLENHLFRRGAQPLTWAIRMKVAIGAAKGLTFLH-DAKSQVIYRDFKAANILLDAE 227
Query: 282 YNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEML 341
+N+KLSDFGLAK GP G+KTHVST+VMGT+GYAAPEYV TG L++KSDVYSFGVVLLE+L
Sbjct: 228 FNSKLSDFGLAKAGPTGDKTHVSTQVMGTHGYAAPEYVATGRLTAKSDVYSFGVVLLELL 287
Query: 342 TGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLS 401
+GRR++DK + E +LV+WA P LG +R F+I+D RL G + KGA AA LA QCL+
Sbjct: 288 SGRRAVDKSKVGMEQSLVDWATPYLGDKRKLFRIMDTRLGGQYPQKGAYTAASLALQCLN 347
Query: 402 RDPKARPLMSEVVHTLKPLPNLK 424
D K RP MSEV+ L L + K
Sbjct: 348 PDAKLRPKMSEVLAKLDQLESTK 370
>UniRef100_Q8GYU1 Hypothetical protein At2g02800/T20F6.6 [Arabidopsis thaliana]
Length = 426
Score = 419 bits (1076), Expect = e-115
Identities = 221/383 (57%), Positives = 277/383 (71%), Gaps = 16/383 (4%)
Query: 44 GSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEE 103
G+C+ S KVDSS S + N S+ S ++ T + P SS + ++
Sbjct: 2 GNCLDSSAKVDSS-SHSPHANSASLSSRVSSK----------TSRSTVP--SSLSINSYS 48
Query: 104 SIASTPKLFSE-ELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTA 162
S+ S P +E E+ + +L+ FTFN LK ATRNFRP+SLLGEGGFG VFKGWI+
Sbjct: 49 SVESLPTPRTEGEILSSPNLKAFTFNELKNATRNFRPDSLLGEGGFGYVFKGWIDGTTLT 108
Query: 163 PVKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEF 222
KPG+G+ VAVK L G+QGHKEWL E+NYLG L HPNLVKL+G+C+E + RLLVYEF
Sbjct: 109 ASKPGSGIVVAVKKLKTEGYQGHKEWLTEVNYLGQLSHPNLVKLVGYCVEGENRLLVYEF 168
Query: 223 MPRGSLENHLFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAE 281
MP+GSLENHLFRR PL W+IRMK+A+GAAKGL FLH D++ +IYRDFK +NILLDAE
Sbjct: 169 MPKGSLENHLFRRGAQPLTWAIRMKVAIGAAKGLTFLH-DAKSQVIYRDFKAANILLDAE 227
Query: 282 YNAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEML 341
+N+KLSDFGLAK GP G+KTHVST+VMGT+GYAAPEYV TG L++KSDVYSFGVVLLE+L
Sbjct: 228 FNSKLSDFGLAKAGPTGDKTHVSTQVMGTHGYAAPEYVATGRLTAKSDVYSFGVVLLELL 287
Query: 342 TGRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLS 401
+GRR++D+ + E +LV+WA P LG +R F+I+D RL G + KGA AA LA QCL+
Sbjct: 288 SGRRAVDRSKVGMEQSLVDWATPYLGDKRKLFRIMDTRLGGQYPQKGAYTAASLALQCLN 347
Query: 402 RDPKARPLMSEVVHTLKPLPNLK 424
D K RP MSEV+ L L + K
Sbjct: 348 PDAKLRPKMSEVLAKLDQLESTK 370
>UniRef100_Q8GXZ3 Hypothetical protein At5g01020/F7J8_5 [Arabidopsis thaliana]
Length = 410
Score = 419 bits (1076), Expect = e-115
Identities = 226/376 (60%), Positives = 274/376 (72%), Gaps = 15/376 (3%)
Query: 107 STPKLFSEELKVASSLRK---FTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAP 163
STP+ F ++ + S + FT L+ T++FRP+ +LGEGGFG V+KG+I++N
Sbjct: 37 STPR-FRDDSRTPISYAQVIPFTLFELETITKSFRPDYILGEGGFGTVYKGYIDDNLRVG 95
Query: 164 VKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFM 223
+K L VAVK+LN G QGH+EWL E+N+LG L HPNLVKLIG+C EDD RLLVYEFM
Sbjct: 96 LK---SLPVAVKVLNKEGLQGHREWLTEVNFLGQLRHPNLVKLIGYCCEDDHRLLVYEFM 152
Query: 224 PRGSLENHLFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEY 282
RGSLENHLFR+ PL WS RM IALGAAKGLAFLH +++RP+IYRDFKTSNILLD++Y
Sbjct: 153 LRGSLENHLFRKTTAPLSWSRRMMIALGAAKGLAFLH-NAERPVIYRDFKTSNILLDSDY 211
Query: 283 NAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLT 342
AKLSDFGLAK GP+G++THVSTRVMGTYGYAAPEYVMTGHL+++SDVYSFGVVLLEMLT
Sbjct: 212 TAKLSDFGLAKAGPQGDETHVSTRVMGTYGYAAPEYVMTGHLTARSDVYSFGVVLLEMLT 271
Query: 343 GRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSR 402
GR+S+DK RP+ E NLV+WARP L +R QIIDPRLE +SV+ AQKA LA CLS+
Sbjct: 272 GRKSVDKTRPSKEQNLVDWARPKLNDKRKLLQIIDPRLENQYSVRAAQKACSLAYYCLSQ 331
Query: 403 DPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLPKKG 462
+PKARPLMS+VV TL+PL D I +P+++ R K
Sbjct: 332 NPKARPLMSDVVETLEPLQCTGDALIPCATTTAGAAFAMGGVPDYRMHRR-----FAKNV 386
Query: 463 QPLRILSSPNCPNGSP 478
P I SPN PN SP
Sbjct: 387 GPGAICRSPN-PNYSP 401
>UniRef100_Q8S7W7 Hypothetical protein OSJNBa0091P11.5 [Oryza sativa]
Length = 422
Score = 417 bits (1073), Expect = e-115
Identities = 224/438 (51%), Positives = 298/438 (67%), Gaps = 29/438 (6%)
Query: 44 GSCIPSRPKVDSSISGTSTHNGNSIKSVITQSEENKSGFEKNTEETEAPPESSTTTSNEE 103
G+C+ + +VD S+ N + S +T S+ + S + + + + +
Sbjct: 2 GNCMDTTARVDHSM------NNGAYPSKVT-SKTSLSSVPSTFKSNSSRSTLTLPSMKDR 54
Query: 104 SIASTPKLFSEELKVASSLRKFTFNGLKVATRNFRPESLLGEGGFGCVFKGWIEENGTAP 163
S TP+ E L +S+L+ F+FN L+ AT+NFRP+SLLGEGGFG V+KGWI+E+ AP
Sbjct: 55 SELPTPRTEGEILS-SSNLKAFSFNDLRNATKNFRPDSLLGEGGFGHVYKGWIDEHTLAP 113
Query: 164 VKPGTGLTVAVKILNHNGHQGHKEWLAELNYLGDLLHPNLVKLIGFCIEDDQRLLVYEFM 223
KPG+G+ VAVK L G QGHKEWL E+NYLG L H NLVKLIG+C + D RLLVYEFM
Sbjct: 114 SKPGSGMVVAVKKLKPEGFQGHKEWLTEVNYLGQLHHKNLVKLIGYCSDGDNRLLVYEFM 173
Query: 224 PRGSLENHLFRRPL-PLPWSIRMKIALGAAKGLAFLHEDSQRPIIYRDFKTSNILLDAEY 282
P+GSLENHLFRR PL W+IR+K+A+GAA+GL+FLH D++ +IYRDFK SNILLD+E+
Sbjct: 174 PKGSLENHLFRRGADPLSWAIRLKVAIGAARGLSFLH-DAENQVIYRDFKASNILLDSEF 232
Query: 283 NAKLSDFGLAKDGPEGEKTHVSTRVMGTYGYAAPEYVMTGHLSSKSDVYSFGVVLLEMLT 342
N+KLSDFGLAK GP G+KTHVST+VMGT+GYAAPEY+ TG LS+K+DVYSFGVVLLE+LT
Sbjct: 233 NSKLSDFGLAKAGPTGDKTHVSTQVMGTHGYAAPEYIATGRLSAKADVYSFGVVLLELLT 292
Query: 343 GRRSIDKKRPNGEHNLVEWARPVLGHRRMFFQIIDPRLEGHFSVKGAQKAAQLAAQCLSR 402
GRR++DK +P E NLV+WA+P LG +R ++++D +L G + KGA A +A QC+
Sbjct: 293 GRRALDKSKPGIEQNLVDWAKPHLGDKRRLYRVMDTKLGGQYPKKGAHAIANIALQCICN 352
Query: 403 DPKARPLMSEVVHTLKPLPNLKDMAISSYHFQTARVDRTMSMPNHKNGIRTQLVSLPKKG 462
D K RP MSEV+ L+ L + S Y+ + +VD IR ++PK
Sbjct: 353 DAKMRPRMSEVLEELEQLQD------SKYNMASPQVD-----------IRRTSNAVPK-- 393
Query: 463 QPLRILSSPNCPNGSPYS 480
P+RI SP G+ S
Sbjct: 394 SPMRIQPSPRRSLGAAAS 411
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 863,780,372
Number of Sequences: 2790947
Number of extensions: 37598032
Number of successful extensions: 153375
Number of sequences better than 10.0: 17170
Number of HSP's better than 10.0 without gapping: 7486
Number of HSP's successfully gapped in prelim test: 9689
Number of HSP's that attempted gapping in prelim test: 116559
Number of HSP's gapped (non-prelim): 20966
length of query: 490
length of database: 848,049,833
effective HSP length: 131
effective length of query: 359
effective length of database: 482,435,776
effective search space: 173194443584
effective search space used: 173194443584
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0391.11


