

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388b.3
(149 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
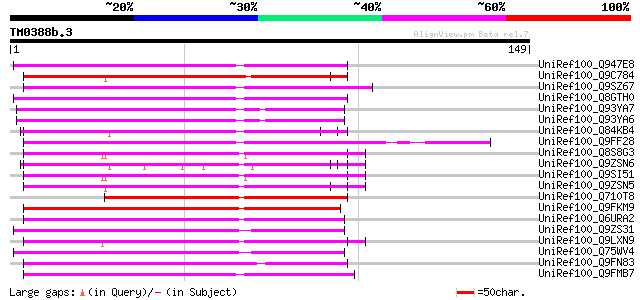
Sequences producing significant alignments: (bits) Value
UniRef100_Q947E8 Resistance gene analog PU3 [Helianthus annuus] 72 3e-12
UniRef100_Q9C784 Disease resistance protein, putative [Arabidops... 70 1e-11
UniRef100_Q9SZ67 Probable WRKY transcription factor 19 [Arabidop... 69 2e-11
UniRef100_Q8GTH0 NBS-LRR resistance protein RS6-8 [Helianthus an... 68 4e-11
UniRef100_Q93YA7 Resistance gene-like [Solanum tuberosum subsp. ... 68 4e-11
UniRef100_Q93YA6 Resistance gene-like [Solanum tuberosum subsp. ... 68 4e-11
UniRef100_Q84KB4 MRGH5 [Cucumis melo] 67 6e-11
UniRef100_Q9FF28 Disease resistance protein-like [Arabidopsis th... 66 1e-10
UniRef100_Q8S8G3 Disease resistance protein (TIR-NBS-LRR class),... 65 3e-10
UniRef100_Q9ZSN6 Disease resistance protein RPP1-WsA [Arabidopsi... 65 3e-10
UniRef100_Q9SI51 Disease resistance protein (TIR-NBS-LRR class),... 65 3e-10
UniRef100_Q9ZSN5 Disease resistance protein RPP1-WsB [Arabidopsi... 63 2e-09
UniRef100_Q710T8 TIR/NBS/LRR protein [Populus deltoides] 63 2e-09
UniRef100_Q9FKM9 Disease resistance protein-like [Arabidopsis th... 62 2e-09
UniRef100_Q6URA2 TIR-NBS-LRR type R protein 7 [Malus baccata] 62 2e-09
UniRef100_Q9ZS31 NL27 [Solanum tuberosum] 62 3e-09
UniRef100_Q9LXN9 Disease resistance protein-like [Arabidopsis th... 62 3e-09
UniRef100_Q75WV4 N protein [Nicotiana tabacum] 61 6e-09
UniRef100_Q9FN83 Disease resistance protein-like [Arabidopsis th... 61 6e-09
UniRef100_Q9FMB7 Disease resistance protein-like [Arabidopsis th... 61 6e-09
>UniRef100_Q947E8 Resistance gene analog PU3 [Helianthus annuus]
Length = 770
Score = 72.0 bits (175), Expect = 3e-12
Identities = 39/96 (40%), Positives = 54/96 (55%), Gaps = 2/96 (2%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
+P LK LDL+ S L+ TPNFDG LERLD GC +L ++HPSIG L ++ + CS
Sbjct: 676 LPNLKILDLAMSSNLITTPNFDGLPCLERLDLEGCESLEEIHPSIGYHKSLVYVDMRRCS 735
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
+L + ++ L L LS C L+ P+ N
Sbjct: 736 TLKR--FSPIIQMQMLETLILSECRELQQFPDIQSN 769
>UniRef100_Q9C784 Disease resistance protein, putative [Arabidopsis thaliana]
Length = 1398
Score = 70.1 bits (170), Expect = 1e-11
Identities = 41/93 (44%), Positives = 60/93 (64%), Gaps = 1/93 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK L+LS L+E P+ G+ L++LD +GC++L+++ SIG L L L L GCSSLV
Sbjct: 1076 LKTLNLSGCSSLVELPSSIGNLNLKKLDLSGCSSLVELPSSIGNLINLKKLDLSGCSSLV 1135
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 1136 ELPL-SIGNLINLQELYLSECSSLVELPSSIGN 1167
Score = 62.4 bits (150), Expect = 2e-09
Identities = 40/94 (42%), Positives = 59/94 (62%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+LDLS L+E P+ G+ L++LD +GC++L+++ SIG L L L L CSSL
Sbjct: 1099 LKKLDLSGCSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSL 1158
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 1159 VELP-SSIGNLINLQELYLSECSSLVELPSSIGN 1191
Score = 62.0 bits (149), Expect = 3e-09
Identities = 41/94 (43%), Positives = 59/94 (62%), Gaps = 3/94 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L+ L LS L+E P+ G+ L++LD +GC++L+++ SIG L L L+L GCSSL
Sbjct: 1028 LQELYLSECSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSGCSSL 1087
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ LN L+ L LSGC+ L P+ IGN
Sbjct: 1088 VEL-PSSIGNLN-LKKLDLSGCSSLVELPSSIGN 1119
Score = 59.7 bits (143), Expect = 1e-08
Identities = 39/94 (41%), Positives = 56/94 (59%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L RLDL L+E P+ G+ LE F GC++LL++ SIG L L L L+ SSL
Sbjct: 788 LPRLDLMGCSSLVELPSSIGNLINLEAFYFHGCSSLLELPSSIGNLISLKILYLKRISSL 847
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V + ++ L +L++L+LSGC+ L P+ IGN
Sbjct: 848 VEI-PSSIGNLINLKLLNLSGCSSLVELPSSIGN 880
Score = 58.2 bits (139), Expect = 4e-08
Identities = 34/93 (36%), Positives = 53/93 (56%), Gaps = 1/93 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK +DL S +L E PN + L + + C++L+++ SIG T + L ++GCSSL+
Sbjct: 693 LKVMDLRYSSHLKELPNLSTAINLLEMVLSDCSSLIELPSSIGNATNIKSLDIQGCSSLL 752
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L ++ L +L L L GC+ L P+ IGN
Sbjct: 753 KL-PSSIGNLITLPRLDLMGCSSLVELPSSIGN 784
Score = 58.2 bits (139), Expect = 4e-08
Identities = 38/94 (40%), Positives = 58/94 (61%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L+ L LS L+E P+ G+ L++LD +GC++L+++ SIG L L L+L CSSL
Sbjct: 956 LQELYLSECSSLVELPSSIGNLINLKKLDLSGCSSLVELPLSIGNLINLKTLNLSECSSL 1015
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 1016 VEL-PSSIGNLINLQELYLSECSSLVELPSSIGN 1048
Score = 58.2 bits (139), Expect = 4e-08
Identities = 36/94 (38%), Positives = 53/94 (56%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
+K LD+ L++ P+ G+ L RLD GC++L+++ SIG L L L L GCSSL
Sbjct: 740 IKSLDIQGCSSLLKLPSSIGNLITLPRLDLMGCSSLVELPSSIGNLINLPRLDLMGCSSL 799
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L + GC+ L P+ IGN
Sbjct: 800 VEL-PSSIGNLINLEAFYFHGCSSLLELPSSIGN 832
Score = 57.8 bits (138), Expect = 5e-08
Identities = 39/94 (41%), Positives = 57/94 (60%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK L L L+E P+ G+ L+ L+ +GC++L+++ SIG L L L L GCSSL
Sbjct: 836 LKILYLKRISSLVEIPSSIGNLINLKLLNLSGCSSLVELPSSIGNLINLKKLDLSGCSSL 895
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 896 VELPL-SIGNLINLQELYLSECSSLVELPSSIGN 928
Score = 56.2 bits (134), Expect = 1e-07
Identities = 38/94 (40%), Positives = 56/94 (59%), Gaps = 2/94 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+LDLS L+E P G+ L+ L + C++L+++ SIG L L L+L CSSL
Sbjct: 884 LKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLKTLNLSECSSL 943
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++ L +L+ L+LS C+ L P+ IGN
Sbjct: 944 VEL-PSSIGNLINLQELYLSECSSLVELPSSIGN 976
Score = 51.2 bits (121), Expect = 5e-06
Identities = 36/89 (40%), Positives = 52/89 (57%), Gaps = 2/89 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+LDLS L+E P G+ L+ L + C++L+++ SIG L L L L CSSL
Sbjct: 1123 LKKLDLSGCSSLVELPLSIGNLINLQELYLSECSSLVELPSSIGNLINLQELYLSECSSL 1182
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTP 92
V L ++ L +L+ L L+ CT+L S P
Sbjct: 1183 VEL-PSSIGNLINLKKLDLNKCTKLVSLP 1210
>UniRef100_Q9SZ67 Probable WRKY transcription factor 19 [Arabidopsis thaliana]
Length = 1895
Score = 69.3 bits (168), Expect = 2e-11
Identities = 37/100 (37%), Positives = 57/100 (57%), Gaps = 2/100 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++ LS S L + P + LE +D GC +LL + SI L +L FL+L+GCS L
Sbjct: 1260 LKKMRLSYSDQLTKIPRLSSATNLEHIDLEGCNSLLSLSQSISYLKKLVFLNLKGCSKLE 1319
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCG 104
+ + ++ +L SL +L+LSGC++L + P N G
Sbjct: 1320 N--IPSMVDLESLEVLNLSGCSKLGNFPEISPNVKELYMG 1357
>UniRef100_Q8GTH0 NBS-LRR resistance protein RS6-8 [Helianthus annuus]
Length = 577
Score = 68.2 bits (165), Expect = 4e-11
Identities = 36/96 (37%), Positives = 56/96 (57%), Gaps = 2/96 (2%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
+P LK LDLS S L++TP+F+G LERL C L ++HPSIG L +++++GC+
Sbjct: 381 LPNLKILDLSGSSNLIKTPDFEGLPCLERLILKYCERLEEIHPSIGYHKRLVYVNMKGCA 440
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L + + L L+LS C++L+ P+ N
Sbjct: 441 RLKR--FPPIIHMKKLETLNLSDCSKLQQFPDIQSN 474
>UniRef100_Q93YA7 Resistance gene-like [Solanum tuberosum subsp. andigena]
Length = 1126
Score = 68.2 bits (165), Expect = 4e-11
Identities = 41/94 (43%), Positives = 51/94 (53%), Gaps = 3/94 (3%)
Query: 3 PFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSS 62
PFL+RLDLS+ LM TP+F LE L C+NL +VH S+ +L L+L C +
Sbjct: 628 PFLRRLDLSSCANLMRTPDFTDMPNLEYLGLEECSNLKEVHHSLRCSKKLIKLNLRDCKN 687
Query: 63 LVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L S VC SL LHL GC+ LE P G
Sbjct: 688 LES--FSYVC-WESLECLHLQGCSNLEKFPRIRG 718
Score = 32.7 bits (73), Expect = 1.7
Identities = 25/68 (36%), Positives = 34/68 (49%), Gaps = 2/68 (2%)
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTR 87
L LD +G NL + SIG L L L + CS L SL + +L +L IL +G T
Sbjct: 747 LTELDLSGMKNLATLSCSIGELKSLVMLKVSYCSKLKSL-PEEIGDLENLEILK-AGYTL 804
Query: 88 LESTPNFI 95
+ P+ I
Sbjct: 805 ISQPPSSI 812
>UniRef100_Q93YA6 Resistance gene-like [Solanum tuberosum subsp. andigena]
Length = 1101
Score = 68.2 bits (165), Expect = 4e-11
Identities = 41/94 (43%), Positives = 51/94 (53%), Gaps = 3/94 (3%)
Query: 3 PFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSS 62
PFL+RLDLS+ LM TP+F LE L C+NL +VH S+ +L L+L C +
Sbjct: 603 PFLRRLDLSSCANLMRTPDFTDMPNLEYLGLEECSNLKEVHHSLRCSKKLIKLNLRDCKN 662
Query: 63 LVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L S VC SL LHL GC+ LE P G
Sbjct: 663 LES--FSYVC-WESLECLHLQGCSNLEKFPRIRG 693
Score = 32.7 bits (73), Expect = 1.7
Identities = 25/68 (36%), Positives = 34/68 (49%), Gaps = 2/68 (2%)
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTR 87
L LD +G NL + SIG L L L + CS L SL + +L +L IL +G T
Sbjct: 722 LTELDLSGMKNLATLSCSIGELKSLVMLKVSYCSKLKSL-PEEIGDLENLEILK-AGYTL 779
Query: 88 LESTPNFI 95
+ P+ I
Sbjct: 780 ISQPPSSI 787
>UniRef100_Q84KB4 MRGH5 [Cucumis melo]
Length = 1092
Score = 67.4 bits (163), Expect = 6e-11
Identities = 38/90 (42%), Positives = 52/90 (57%), Gaps = 2/90 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ LDLS+ K L + P+ + L L F CTNL+ +H SIG LT+L L L+ CS+L
Sbjct: 701 LEDLDLSHCKKLEKIPDISSASNLRSLSFEQCTNLVMIHDSIGSLTKLVTLKLQNCSNLK 760
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNF 94
L N L+ L+LS C +LE P+F
Sbjct: 761 K--LPRYISWNFLQDLNLSWCKKLEEIPDF 788
Score = 67.0 bits (162), Expect = 8e-11
Identities = 41/94 (43%), Positives = 56/94 (58%), Gaps = 2/94 (2%)
Query: 4 FLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
FL+ L+LS K L E P+F + L+ L CT+L VH SIG L++L L+LE CS+L
Sbjct: 770 FLQDLNLSWCKKLEEIPDFSSTSNLKHLSLEQCTSLRVVHDSIGSLSKLVSLNLEKCSNL 829
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L + +L SL+ L LSGC +LE+ P N
Sbjct: 830 EK--LPSYLKLKSLQNLTLSGCCKLETFPEIDEN 861
Score = 45.8 bits (107), Expect = 2e-04
Identities = 35/88 (39%), Positives = 47/88 (52%), Gaps = 7/88 (7%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQR---LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
L+ L LS L P D + + + RLD T + ++ PSIG LT L L+GC+
Sbjct: 841 LQNLTLSGCCKLETFPEIDENMKSLYILRLDSTA---IRELPPSIGYLTHLYMFDLKGCT 897
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLE 89
+L+SL T L SL LHLSG +R E
Sbjct: 898 NLISLPC-TTHLLKSLGELHLSGSSRFE 924
Score = 40.4 bits (93), Expect = 0.008
Identities = 31/98 (31%), Positives = 46/98 (46%), Gaps = 2/98 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK LDL +S L + + LE L + C+NL + S L +L L L C +L
Sbjct: 630 LKLLDLRHSVILKKISESSAAPNLEELYLSNCSNLKTIPKSFLSLRKLVTLDLHHCVNLK 689
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
+ + +L L LS C +LE P+ I + S+ R
Sbjct: 690 KIPRSYI-SWEALEDLDLSHCKKLEKIPD-ISSASNLR 725
Score = 31.2 bits (69), Expect = 4.9
Identities = 15/35 (42%), Positives = 22/35 (62%), Gaps = 2/35 (5%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNL 39
L+ L+L N K+L E PN ++R+D TGC +L
Sbjct: 1016 LRNLELRNCKFLQEIPNL--PLCIQRVDATGCVSL 1048
>UniRef100_Q9FF28 Disease resistance protein-like [Arabidopsis thaliana]
Length = 833
Score = 66.2 bits (160), Expect = 1e-10
Identities = 45/134 (33%), Positives = 69/134 (50%), Gaps = 9/134 (6%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR+DLS SK+L E P+ + LE L +GC +L+++ SIG L +L LSL GCS L
Sbjct: 480 LKRMDLSESKHLKELPDLSTATNLEYLIMSGCISLVELPSSIGKLRKLLMLSLRGCSKLE 539
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFRCGFDIVVPGNKIPYWFDSQFKG 124
+ L T L SL L L+ C ++ P N + + ++P S K
Sbjct: 540 A--LPTNINLESLDYLDLTDCLLIKKFPEISTNIKDLKLTKTAI---KEVP----STIKS 590
Query: 125 GSRIREVDHVYEQD 138
S +R+++ Y ++
Sbjct: 591 WSHLRKLEMSYSEN 604
>UniRef100_Q8S8G3 Disease resistance protein (TIR-NBS-LRR class), putative
[Arabidopsis thaliana]
Length = 972
Score = 65.1 bits (157), Expect = 3e-10
Identities = 37/98 (37%), Positives = 58/98 (58%), Gaps = 1/98 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR+DL +SK L E P+ + LE L+ GC++L+++ SIG T+L L L GCSSL+
Sbjct: 676 LKRMDLFSSKNLKELPDLSSATNLEVLNLNGCSSLVELPFSIGNATKLLKLELSGCSSLL 735
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L ++ +L+ + S C L P+ IGN ++ +
Sbjct: 736 EL-PSSIGNAINLQTIDFSHCENLVELPSSIGNATNLK 772
Score = 57.8 bits (138), Expect = 5e-08
Identities = 36/99 (36%), Positives = 57/99 (57%), Gaps = 2/99 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L +L+LS L+E P+ G+ L+ +DF+ C NL+++ SIG T L L L CSSL
Sbjct: 723 LLKLELSGCSSLLELPSSIGNAINLQTIDFSHCENLVELPSSIGNATNLKELDLSCCSSL 782
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L ++ +L+ LHL C+ L+ P+ IGN ++ +
Sbjct: 783 KEL-PSSIGNCTNLKKLHLICCSSLKELPSSIGNCTNLK 820
Score = 51.2 bits (121), Expect = 5e-06
Identities = 37/99 (37%), Positives = 52/99 (52%), Gaps = 2/99 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK LDLS L E P+ G+ L++L C++L ++ SIG T L L L CSSL
Sbjct: 771 LKELDLSCCSSLKELPSSIGNCTNLKKLHLICCSSLKELPSSIGNCTNLKELHLTCCSSL 830
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
+ L +N L L L+GC L P+FIG ++ +
Sbjct: 831 IKLPSSIGNAIN-LEKLILAGCESLVELPSFIGKATNLK 868
Score = 48.5 bits (114), Expect = 3e-05
Identities = 36/96 (37%), Positives = 53/96 (54%), Gaps = 6/96 (6%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+L L L E P+ G+ L+ L T C++L+++ SIG L L L GC SL
Sbjct: 795 LKKLHLICCSSLKELPSSIGNCTNLKELHLTCCSSLIKLPSSIGNAINLEKLILAGCESL 854
Query: 64 VSL--YLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++G + +L+IL+L + L P+FIGN
Sbjct: 855 VELPSFIG---KATNLKILNLGYLSCLVELPSFIGN 887
>UniRef100_Q9ZSN6 Disease resistance protein RPP1-WsA [Arabidopsis thaliana]
Length = 1189
Score = 65.1 bits (157), Expect = 3e-10
Identities = 38/88 (43%), Positives = 49/88 (55%), Gaps = 2/88 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK +DLS S YL E PN + LE L+ C++L+++ SI LT L L L+GCSSLV
Sbjct: 669 LKWMDLSYSSYLKELPNLSTATNLEELNLRNCSSLVELPSSIEKLTSLQILDLQGCSSLV 728
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTP 92
L + L IL+L C LE P
Sbjct: 729 E--LPSFGNATKLEILYLDYCRSLEKLP 754
Score = 53.5 bits (127), Expect = 9e-07
Identities = 31/95 (32%), Positives = 53/95 (55%), Gaps = 3/95 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAF--LSLEGCSS 62
L++L L N ++E P + + L L+ C++L+++ SIG L L++ GCSS
Sbjct: 762 LQKLSLRNCSRIVELPAIENATNLWELNLLNCSSLIELPLSIGTARNLFLKELNISGCSS 821
Query: 63 LVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
LV L ++ ++ +L+ LS C+ L P+ IGN
Sbjct: 822 LVKL-PSSIGDMTNLKEFDLSNCSNLVELPSSIGN 855
Score = 50.8 bits (120), Expect = 6e-06
Identities = 33/91 (36%), Positives = 53/91 (57%), Gaps = 4/91 (4%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQR---LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
L L+L N L+E P G+ R L+ L+ +GC++L+++ SIG +T L L CS
Sbjct: 785 LWELNLLNCSSLIELPLSIGTARNLFLKELNISGCSSLVKLPSSIGDMTNLKEFDLSNCS 844
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTP 92
+LV L ++ L +L L + GC++LE+ P
Sbjct: 845 NLVEL-PSSIGNLQNLCKLIMRGCSKLEALP 874
Score = 45.4 bits (106), Expect = 3e-04
Identities = 33/93 (35%), Positives = 52/93 (55%), Gaps = 6/93 (6%)
Query: 4 FLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCT--NLLQVHPSIGLLTELAFLSLEGCS 61
FL LD+S SK ++G+++L L + + + L+ P++ T L L+L CS
Sbjct: 645 FLVELDMSFSKL---QKLWEGTKQLRNLKWMDLSYSSYLKELPNLSTATNLEELNLRNCS 701
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNF 94
SLV L ++ +L SL+IL L GC+ L P+F
Sbjct: 702 SLVELP-SSIEKLTSLQILDLQGCSSLVELPSF 733
Score = 43.9 bits (102), Expect = 7e-04
Identities = 38/122 (31%), Positives = 57/122 (46%), Gaps = 25/122 (20%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGL---------------- 48
L+ LDL L+E P+F + +LE L C +L ++ PSI
Sbjct: 716 LQILDLQGCSSLVELPSFGNATKLEILYLDYCRSLEKLPPSINANNLQKLSLRNCSRIVE 775
Query: 49 ------LTELAFLSLEGCSSLVSLYL--GTVCELNSLRILHLSGCTRLESTPNFIGNPSH 100
T L L+L CSSL+ L L GT L L+ L++SGC+ L P+ IG+ ++
Sbjct: 776 LPAIENATNLWELNLLNCSSLIELPLSIGTARNL-FLKELNISGCSSLVKLPSSIGDMTN 834
Query: 101 FR 102
+
Sbjct: 835 LK 836
Score = 37.7 bits (86), Expect = 0.052
Identities = 31/102 (30%), Positives = 44/102 (42%), Gaps = 25/102 (24%)
Query: 1 DVPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGC 60
D+ LK DLSN L+E P+ SIG L L L + GC
Sbjct: 831 DMTNLKEFDLSNCSNLVELPS-----------------------SIGNLQNLCKLIMRGC 867
Query: 61 SSLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
S L +L + L SL L+L+ C++L+S P + + R
Sbjct: 868 SKLEALPIN--INLKSLDTLNLTDCSQLKSFPEISTHIKYLR 907
>UniRef100_Q9SI51 Disease resistance protein (TIR-NBS-LRR class), putative
[Arabidopsis thaliana]
Length = 554
Score = 65.1 bits (157), Expect = 3e-10
Identities = 37/98 (37%), Positives = 58/98 (58%), Gaps = 1/98 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LKR+DL +SK L E P+ + LE L+ GC++L+++ SIG T+L L L GCSSL+
Sbjct: 15 LKRMDLFSSKNLKELPDLSSATNLEVLNLNGCSSLVELPFSIGNATKLLKLELSGCSSLL 74
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L ++ +L+ + S C L P+ IGN ++ +
Sbjct: 75 EL-PSSIGNAINLQTIDFSHCENLVELPSSIGNATNLK 111
Score = 57.8 bits (138), Expect = 5e-08
Identities = 36/99 (36%), Positives = 57/99 (57%), Gaps = 2/99 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
L +L+LS L+E P+ G+ L+ +DF+ C NL+++ SIG T L L L CSSL
Sbjct: 62 LLKLELSGCSSLLELPSSIGNAINLQTIDFSHCENLVELPSSIGNATNLKELDLSCCSSL 121
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L ++ +L+ LHL C+ L+ P+ IGN ++ +
Sbjct: 122 KEL-PSSIGNCTNLKKLHLICCSSLKELPSSIGNCTNLK 159
Score = 51.2 bits (121), Expect = 5e-06
Identities = 37/99 (37%), Positives = 52/99 (52%), Gaps = 2/99 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK LDLS L E P+ G+ L++L C++L ++ SIG T L L L CSSL
Sbjct: 110 LKELDLSCCSSLKELPSSIGNCTNLKKLHLICCSSLKELPSSIGNCTNLKELHLTCCSSL 169
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
+ L +N L L L+GC L P+FIG ++ +
Sbjct: 170 IKLPSSIGNAIN-LEKLILAGCESLVELPSFIGKATNLK 207
Score = 48.5 bits (114), Expect = 3e-05
Identities = 36/96 (37%), Positives = 53/96 (54%), Gaps = 6/96 (6%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+L L L E P+ G+ L+ L T C++L+++ SIG L L L GC SL
Sbjct: 134 LKKLHLICCSSLKELPSSIGNCTNLKELHLTCCSSLIKLPSSIGNAINLEKLILAGCESL 193
Query: 64 VSL--YLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
V L ++G + +L+IL+L + L P+FIGN
Sbjct: 194 VELPSFIG---KATNLKILNLGYLSCLVELPSFIGN 226
Score = 30.4 bits (67), Expect = 8.4
Identities = 31/99 (31%), Positives = 44/99 (44%), Gaps = 11/99 (11%)
Query: 3 PFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCT--NLLQVHPSIGLLTELAFLSLEGC 60
P L+ L + S+ L E S LER+ + N+ ++ P + +T L L L GC
Sbjct: 295 PRLEDLQMLYSENLSEF-----SHVLERITVLELSDINIREMTPWLNRITRLRRLKLSGC 349
Query: 61 SSLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPS 99
LVSL + +SL IL C LE NP+
Sbjct: 350 GKLVSLPQLS----DSLIILDAENCGSLERLGCSFNNPN 384
>UniRef100_Q9ZSN5 Disease resistance protein RPP1-WsB [Arabidopsis thaliana]
Length = 1221
Score = 62.8 bits (151), Expect = 2e-09
Identities = 32/93 (34%), Positives = 54/93 (57%), Gaps = 1/93 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ L L+N ++E P + + L +L+ C++L+++ SIG T L L GCSSLV
Sbjct: 794 LQELSLTNCSRVVELPAIENATNLWKLNLLNCSSLIELPLSIGTATNLKHLDFRGCSSLV 853
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L ++ ++ +L + +LS C+ L P+ IGN
Sbjct: 854 KL-PSSIGDMTNLEVFYLSNCSNLVELPSSIGN 885
Score = 59.3 bits (142), Expect = 2e-08
Identities = 36/88 (40%), Positives = 46/88 (51%), Gaps = 2/88 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK +DLS S YL E PN + LE L C++L+++ SI LT L L L CSSLV
Sbjct: 701 LKWMDLSYSSYLKELPNLSTATNLEELKLRNCSSLVELPSSIEKLTSLQILDLHRCSSLV 760
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTP 92
L + L IL+L C+ L P
Sbjct: 761 E--LPSFGNATKLEILNLENCSSLVKLP 786
Score = 52.4 bits (124), Expect = 2e-06
Identities = 35/99 (35%), Positives = 51/99 (51%), Gaps = 3/99 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQ-RLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK LD L++ P+ G LE + C+NL+++ SIG L +L L + GCS L
Sbjct: 841 LKHLDFRGCSSLVKLPSSIGDMTNLEVFYLSNCSNLVELPSSIGNLRKLTLLLMRGCSKL 900
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
+ L T L SL L+L C+RL+S P + + R
Sbjct: 901 ET--LPTNINLKSLHTLNLIDCSRLKSFPEISTHIKYLR 937
>UniRef100_Q710T8 TIR/NBS/LRR protein [Populus deltoides]
Length = 1147
Score = 62.8 bits (151), Expect = 2e-09
Identities = 32/70 (45%), Positives = 49/70 (69%), Gaps = 1/70 (1%)
Query: 28 LERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTR 87
LE+L+ GC++L++VH SIG LT L FL+LEGC L +L ++ + SL L++SGC++
Sbjct: 666 LEKLNLKGCSSLVEVHQSIGNLTSLDFLNLEGCWRLKNL-PESIGNVKSLETLNISGCSQ 724
Query: 88 LESTPNFIGN 97
LE P +G+
Sbjct: 725 LEKLPESMGD 734
Score = 34.3 bits (77), Expect = 0.58
Identities = 34/90 (37%), Positives = 46/90 (50%), Gaps = 12/90 (13%)
Query: 5 LKRLDLSN---SKYLMETPNFDGSQRLERLDFTGCTNLLQVHPS-IGLLTELAFLSLEGC 60
+KRL+L + S + +F G LE LD G N PS IG L++L FLS++ C
Sbjct: 800 VKRLELPHGGLSDRAAKCVDFSGLSALEVLDLIG--NKFSSLPSGIGFLSKLKFLSVKAC 857
Query: 61 SSLVSLYLGTVCEL-NSLRILHLSGCTRLE 89
LVS + +L +SL L S C LE
Sbjct: 858 KYLVS-----IPDLPSSLDCLDASYCKSLE 882
>UniRef100_Q9FKM9 Disease resistance protein-like [Arabidopsis thaliana]
Length = 1059
Score = 62.4 bits (150), Expect = 2e-09
Identities = 37/91 (40%), Positives = 57/91 (61%), Gaps = 1/91 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK ++LSNS+ L E P+ + +L+ L+ T C++L+++ SIG T L L+L C+SLV
Sbjct: 681 LKWMNLSNSRNLKELPDLSTATKLQDLNLTRCSSLVEIPFSIGNTTNLEKLNLVMCTSLV 740
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFI 95
L ++ L+ LR L L GC++LE P I
Sbjct: 741 EL-PSSIGSLHKLRELRLRGCSKLEVLPTNI 770
>UniRef100_Q6URA2 TIR-NBS-LRR type R protein 7 [Malus baccata]
Length = 1095
Score = 62.4 bits (150), Expect = 2e-09
Identities = 37/92 (40%), Positives = 50/92 (54%), Gaps = 1/92 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK LDLS S+ L ++P+F LE L C L ++HPSIG L L+ ++LE C L+
Sbjct: 644 LKTLDLSESRSLQKSPDFSQVPNLEELILYNCKELSEIHPSIGHLKRLSLVNLEWCDKLI 703
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
SL G + S+ L L+GC L IG
Sbjct: 704 SL-PGDFYKSKSVEALLLNGCLILRELHEDIG 734
Score = 32.0 bits (71), Expect = 2.9
Identities = 37/141 (26%), Positives = 57/141 (40%), Gaps = 31/141 (21%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ L+LS+ + + D + D N PS+ L++L L L C L
Sbjct: 785 LRELNLSSFELADDEIPKDLGSLISLQDLNLQRNDFHTLPSLSGLSKLETLRLHHCEQL- 843
Query: 65 SLYLGTVCEL-NSLRILHLSGCTRLESTPNF--IGN-------------PSHFR------ 102
T+ +L +L+ L +GC LE+ PNF + N +H R
Sbjct: 844 ----RTITDLPTNLKFLLANGCPALETMPNFSEMSNIRELKVSDSPNNLSTHLRKNILQG 899
Query: 103 ---CGF-DIVVPGNKIPYWFD 119
CGF I + N +P WF+
Sbjct: 900 WTSCGFGGIFLHANYVPDWFE 920
Score = 31.2 bits (69), Expect = 4.9
Identities = 29/88 (32%), Positives = 42/88 (46%), Gaps = 10/88 (11%)
Query: 5 LKRLDLSNSKY---LMETP-NFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGC 60
LKRL L N ++ L+ P +F S+ +E L GC L ++H IG + L L E
Sbjct: 688 LKRLSLVNLEWCDKLISLPGDFYKSKSVEALLLNGCLILRELHEDIGEMISLRTLEAEYT 747
Query: 61 S------SLVSLYLGTVCELNSLRILHL 82
S+V L T L+S+ +HL
Sbjct: 748 DIREVPPSIVRLKNLTRLSLSSVESIHL 775
>UniRef100_Q9ZS31 NL27 [Solanum tuberosum]
Length = 821
Score = 62.0 bits (149), Expect = 3e-09
Identities = 36/95 (37%), Positives = 51/95 (52%), Gaps = 3/95 (3%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
+PFL++LDL +S+ LM+TP+F L+ LD + C NL +VH S+G EL L+L C
Sbjct: 621 LPFLQKLDLRDSRSLMQTPDFTWMPNLKYLDLSYCRNLSEVHHSLGYSRELIELNLYNCG 680
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
L + SL + L C+ LE P G
Sbjct: 681 RLKRF---PCVNVESLDYMDLEFCSSLEKFPIIFG 712
>UniRef100_Q9LXN9 Disease resistance protein-like [Arabidopsis thaliana]
Length = 1220
Score = 61.6 bits (148), Expect = 3e-09
Identities = 38/98 (38%), Positives = 50/98 (50%), Gaps = 2/98 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK +DLS S YL E PN + LE L C++L+++ SI LT L L LE CSSL
Sbjct: 716 LKWMDLSYSSYLKELPNLSTATNLEELKLRNCSSLVELPSSIEKLTSLQILDLENCSSLE 775
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
L + LR L L C+ L P IG ++ +
Sbjct: 776 K--LPAIENATKLRELKLQNCSSLIELPLSIGTATNLK 811
Score = 61.2 bits (147), Expect = 4e-09
Identities = 33/93 (35%), Positives = 53/93 (56%), Gaps = 1/93 (1%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L+ LDL N L + P + + +L L C++L+++ SIG T L L++ GCSSLV
Sbjct: 763 LQILDLENCSSLEKLPAIENATKLRELKLQNCSSLIELPLSIGTATNLKQLNISGCSSLV 822
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
L ++ ++ L + LS C+ L + P+ IGN
Sbjct: 823 KL-PSSIGDITDLEVFDLSNCSSLVTLPSSIGN 854
Score = 50.4 bits (119), Expect = 8e-06
Identities = 34/99 (34%), Positives = 54/99 (54%), Gaps = 3/99 (3%)
Query: 5 LKRLDLSNSKYLMETPNFDGS-QRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSL 63
LK+L++S L++ P+ G LE D + C++L+ + SIG L L L + GCS L
Sbjct: 810 LKQLNISGCSSLVKLPSSIGDITDLEVFDLSNCSSLVTLPSSIGNLQNLCKLIMRGCSKL 869
Query: 64 VSLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPSHFR 102
+L + L SL L+L+ C++L+S P + S R
Sbjct: 870 EALPIN--INLKSLDTLNLTDCSQLKSFPEISTHISELR 906
>UniRef100_Q75WV4 N protein [Nicotiana tabacum]
Length = 1128
Score = 60.8 bits (146), Expect = 6e-09
Identities = 36/95 (37%), Positives = 49/95 (50%), Gaps = 3/95 (3%)
Query: 2 VPFLKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCS 61
+P L+R+DLS SK L TP+F G LE ++ C+NL +VH S+G +++ L L C
Sbjct: 610 LPSLRRIDLSWSKRLTRTPDFTGMPNLEYVNLYQCSNLEEVHHSLGCCSKVIGLYLNDCK 669
Query: 62 SLVSLYLGTVCELNSLRILHLSGCTRLESTPNFIG 96
SL + SL L L C LE P G
Sbjct: 670 SLKRF---PCVNVESLEYLGLRSCDSLEKLPEIYG 701
>UniRef100_Q9FN83 Disease resistance protein-like [Arabidopsis thaliana]
Length = 1295
Score = 60.8 bits (146), Expect = 6e-09
Identities = 34/93 (36%), Positives = 52/93 (55%), Gaps = 2/93 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++DLS KYL+E P+ + LE L+ + C +L++V PSI L L+ L C L
Sbjct: 628 LKKMDLSRCKYLVEVPDLSKATNLEELNLSYCQSLVEVTPSIKNLKGLSCFYLTNCIQLK 687
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGN 97
+ +G + L SL + +SGC+ L+ P N
Sbjct: 688 DIPIGII--LKSLETVGMSGCSSLKHFPEISWN 718
Score = 40.4 bits (93), Expect = 0.008
Identities = 28/87 (32%), Positives = 42/87 (48%), Gaps = 1/87 (1%)
Query: 6 KRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLVS 65
+RL LS++K + L +LD + C L + +G L L L+L+GC L +
Sbjct: 720 RRLYLSSTKIEELPSSISRLSCLVKLDMSDCQRLRTLPSYLGHLVSLKSLNLDGCRRLEN 779
Query: 66 LYLGTVCELNSLRILHLSGCTRLESTP 92
L T+ L SL L +SGC + P
Sbjct: 780 L-PDTLQNLTSLETLEVSGCLNVNEFP 805
Score = 36.6 bits (83), Expect = 0.12
Identities = 41/153 (26%), Positives = 58/153 (37%), Gaps = 23/153 (15%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
L LDLS + + + RL RL+ C L Q P L L ++ + C+SLV
Sbjct: 981 LLELDLSGNNFEFIPASIKRLTRLNRLNLNNCQRL-QALPD-ELPRGLLYIYIHSCTSLV 1038
Query: 65 SLY--LGTVCELNSLRILHLSGCTRLESTPNFI---------GNPSHFRCGFDIVVPGNK 113
S+ C LR L S C +L+ + P H PG+
Sbjct: 1039 SISGCFNQYC----LRKLVASNCYKLDQAAQILIHRNLKLESAKPEHS------YFPGSD 1088
Query: 114 IPYWFDSQFKGGSRIREVDHVYEQDHWLGFAFC 146
IP F+ Q G S ++ LGF+ C
Sbjct: 1089 IPTCFNHQVMGPSLNIQLPQSESSSDILGFSAC 1121
Score = 33.9 bits (76), Expect = 0.76
Identities = 32/109 (29%), Positives = 48/109 (43%), Gaps = 22/109 (20%)
Query: 5 LKRLDLSNSKYLMETPN-FDGSQRLERLDFTGC--------------------TNLLQVH 43
LK L+L + L P+ LE L+ +GC T++ ++
Sbjct: 766 LKSLNLDGCRRLENLPDTLQNLTSLETLEVSGCLNVNEFPRVSTSIEVLRISETSIEEIP 825
Query: 44 PSIGLLTELAFLSLEGCSSLVSLYLGTVCELNSLRILHLSGCTRLESTP 92
I L++L L + L SL + ++ EL SL L LSGC+ LES P
Sbjct: 826 ARICNLSQLRSLDISENKRLASLPV-SISELRSLEKLKLSGCSVLESFP 873
>UniRef100_Q9FMB7 Disease resistance protein-like [Arabidopsis thaliana]
Length = 986
Score = 60.8 bits (146), Expect = 6e-09
Identities = 33/95 (34%), Positives = 54/95 (56%), Gaps = 2/95 (2%)
Query: 5 LKRLDLSNSKYLMETPNFDGSQRLERLDFTGCTNLLQVHPSIGLLTELAFLSLEGCSSLV 64
LK++ LS+S YL + P+ + LE LD C NL+++ S L +L +L++ GC L
Sbjct: 616 LKKMSLSSSWYLKKLPDLSNATNLEELDLRACQNLVELPSSFSYLHKLKYLNMMGCRRLK 675
Query: 65 SLYLGTVCELNSLRILHLSGCTRLESTPNFIGNPS 99
+ L SL ++++ GC+RL+S P+ N S
Sbjct: 676 E--VPPHINLKSLELVNMYGCSRLKSFPDISTNIS 708
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.143 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 263,736,980
Number of Sequences: 2790947
Number of extensions: 10231319
Number of successful extensions: 38151
Number of sequences better than 10.0: 527
Number of HSP's better than 10.0 without gapping: 222
Number of HSP's successfully gapped in prelim test: 314
Number of HSP's that attempted gapping in prelim test: 36483
Number of HSP's gapped (non-prelim): 1479
length of query: 149
length of database: 848,049,833
effective HSP length: 125
effective length of query: 24
effective length of database: 499,181,458
effective search space: 11980354992
effective search space used: 11980354992
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0388b.3


