

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0388a.1
(728 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
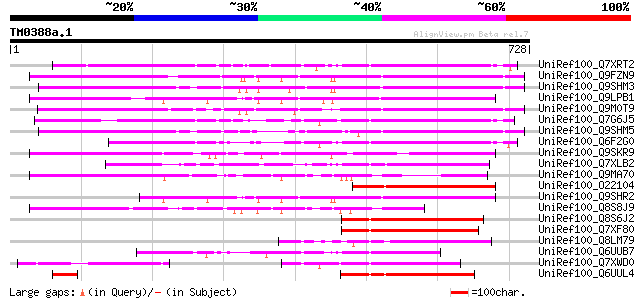
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XRT2 OSJNBa0042F21.11 protein [Oryza sativa] 321 4e-86
UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis ... 297 7e-79
UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana] 291 5e-77
UniRef100_Q9LPB1 T32E20.9 [Arabidopsis thaliana] 289 2e-76
UniRef100_Q9M0T9 Putative athila transposon protein [Arabidopsis... 288 4e-76
UniRef100_Q7G6J5 Putative polyprotein [Oryza sativa] 282 3e-74
UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana] 272 3e-71
UniRef100_Q6F2G0 Hypothetical protein OSJNBa0083K01.26 [Oryza sa... 229 2e-58
UniRef100_Q9SKR9 Putative Athila retroelement ORF1 protein [Arab... 228 4e-58
UniRef100_Q7XLB2 OSJNBa0011K22.17 protein [Oryza sativa] 219 3e-55
UniRef100_Q9MA70 F2J6.11 protein [Arabidopsis thaliana] 198 4e-49
UniRef100_O22104 Vicia faba mRNA expressed from retrotransposon-... 184 6e-45
UniRef100_Q9SHR2 F28L22.6 protein [Arabidopsis thaliana] 176 3e-42
UniRef100_Q8S8J9 Putative Athila retroelement ORF1 protein [Arab... 175 5e-42
UniRef100_Q8S6J2 Hypothetical protein OSJNBa0019N10.34 [Oryza sa... 171 9e-41
UniRef100_Q7XF80 Hypothetical protein [Oryza sativa] 167 8e-40
UniRef100_Q8LM79 Hypothetical protein OSJNAa0082N11.11 [Oryza sa... 167 1e-39
UniRef100_Q6UUB7 Hypothetical protein [Oryza sativa] 160 2e-37
UniRef100_Q7XWD0 OSJNBb0069N01.18 protein [Oryza sativa] 158 5e-37
UniRef100_Q6UUL4 Hypothetical protein [Oryza sativa] 155 3e-36
>UniRef100_Q7XRT2 OSJNBa0042F21.11 protein [Oryza sativa]
Length = 920
Score = 321 bits (823), Expect = 4e-86
Identities = 224/676 (33%), Positives = 348/676 (51%), Gaps = 56/676 (8%)
Query: 61 EDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAE 120
ED +AH+ +F+ C+ Y ++ VS D +RL LFPFSL A++W + +I +W+ +
Sbjct: 274 EDANAHLLQFLEICSMYTIKGVSPDAVRLRLFPFSLLRRAKQWFYANC-AAINTWDKCST 332
Query: 121 KFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYD 180
F ++F P L+ I F Q+ DE++ EAWER ++ + CP H + + FY+
Sbjct: 333 SFLSKFFPIGKTNALRGRISRFQQTRDESIPEAWERLQEYVAACPHHGMDDWLILQNFYN 392
Query: 181 GLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMNSSNDRQAR-RGVLEVEAYDQL 239
GL SR LDAA+ G F + Q ELIEKM S Q R RG+ V+ + L
Sbjct: 393 GLTLMSRDHLDAAAGGAFFSKTVQGAVELIEKMVSNTGWSKERLQTRQRGMHTVKETELL 452
Query: 240 MASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGGP-HDSEECTETRPEEEVKAMGQA 298
A L K +++ + + + T + + + V CGG H +C ET E AM
Sbjct: 453 AAKLDLLMKCLDDHEKRPQGT-VKALDSHVTCEVCGGTGHSGNDCLETHEE----AMYMG 507
Query: 299 RNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQPQERGEGEGSGS 358
+N+ + GW N P + G + GN F Q PS + + + + + +
Sbjct: 508 KNNEYRPQGGEGW-NQPR-PYYPGGNNNGN-FSNQ-PSLMDLVFAQAKTTDALRKKLAAN 563
Query: 359 KKSLEELVETFINRTENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQEN 418
K LE + ++ + ++NQ + NK +E Q QLA + G P P+ EN
Sbjct: 564 DKILEN-INVKLDGFASAFQNQLSFNKMIETQLAQLASLVPANESGRIPGQ--PDSSIEN 620
Query: 419 ASVVATRSGRV--------------MSELKKKTEGEKREEIVGGEVVVPIKVREEVDLTP 464
+ TR G+ MS+ T+ + +E + + VP +E D
Sbjct: 621 VKAITTRGGKSTRDPPYPNPAGTNGMSKETPSTDSDDKE--IQPDKTVP---QEYCD--- 672
Query: 465 ETSKIPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRL 524
T +PFP+ K S+DK+F+ FV+V +K+HI++P DA+ Q+P YA+++KDIL+ +R L
Sbjct: 673 -TRLLPFPQQSRKPSVDKQFACFVEVIQKIHINVPLLDAM-QVPTYARYLKDILNNKRPL 730
Query: 525 KGVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTM 584
E V +TE CS ++ K+P+K++DPG TI I +ALCDLGAS+++MP +
Sbjct: 731 P-TTEVVKLTEHCSNLILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKDV 789
Query: 585 FERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLI 644
F++LN + PT + LQ+AD S+ P GI EDV + + PVDFVVLDM+ ++ PLI
Sbjct: 790 FDKLNFTVLAPTPMRLQLADSSVHYPVGIAEDVPAKIRDFFIPVDFVVLDMDTGKETPLI 849
Query: 645 LGRPFLATGRAKIDVDKGHLILPVGKEKVRFSVFNPMIETNHDNDFVCDVIRSRSK---- 700
LGRPFL+T A IDV G + ++ +F F P +E C ++R + +
Sbjct: 850 LGRPFLSTAGANIDVGTGSIRFHTNGKEEKFE-FQPRMEQ-------CTMVRIKYRPNPQ 901
Query: 701 ----VSEETPKVNSQI 712
V E PK +S +
Sbjct: 902 NIQVVDVEPPKTDSLV 917
>UniRef100_Q9FZN9 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1864
Score = 297 bits (761), Expect = 7e-79
Identities = 226/781 (28%), Positives = 362/781 (45%), Gaps = 110/781 (14%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G + P NFE++ LI +V N++ G +EDP H++ F R C+ ++ VS D +
Sbjct: 60 GIVPPPVQNNNFEIKSGLIAMVQSNKFHGLPMEDPLDHLDEFDRLCSLTKINGVSEDGFK 119
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A +W S QGSITSW D + F +F + +L+NDI FTQ+ +E
Sbjct: 120 LRLFPFSLGDKAHQWEKSLLQGSITSWNDCKKAFLAKFFSNSRTARLRNDISGFTQTNNE 179
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYE 208
EAWERFK +CP H ++A ++ Y G+ R LD AS+G F + G+E
Sbjct: 180 TFCEAWERFKGYQTQCPHHGFSKASLLSTLYRGVLPKIRMLLDTASNGNFLNKDVEDGWE 239
Query: 209 LIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAK 268
L+E +A N + D YD+ + ++ ++ M + +
Sbjct: 240 LVENLAQSDGNYNED------------YDRSVRTSSDSDEKHRREMKAMNDKLDKLLLVQ 287
Query: 269 VEFVTCGGPHDSEECT--ETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWRQGS--- 323
+ + G ++ + ET EEV + ++ +N +NHPN S+R +
Sbjct: 288 QKHIHFLGDDETFQVQDGETMQSEEVSYV--QNQGGYNKGFNNFKQNHPNLSYRSTNVAN 345
Query: 324 ---------------------SGQG--------NGFQRQFPSQGFQGQSLRQP------- 347
GQG +Q Q P GF Q +QP
Sbjct: 346 PQDQVYPSQQQNQPRPFVPYNQGQGYVPKQQYQGNYQPQLPPPGFTQQQ-QQPASTTPDS 404
Query: 348 ------QERGEGEGSGSKKSLEELVETFINRTENNYKN----QEAANKNLENQFGQLAKQ 397
Q+ +G+ +G+ + + E N+ + +Y + EA + GQ
Sbjct: 405 DLKNMLQQILQGQATGAMDLSKRMAEIH-NKVDCSYNDINIKVEALTSKIRYIEGQTGST 463
Query: 398 IAERPQGIFPSDCIPNPKQENASVVATRSGRVMSELKKKTEGEKREEIVGGE-------- 449
A + G PS + +E A + RSG+ + + + + GE
Sbjct: 464 AAPKFTG--PSRKSMSNSEEYAHAITLRSGKELPTKESPNQNTEDSVDQDGEDFCQNGNS 521
Query: 450 ----VVVPI----------------------KVREEVDLTPETSK-IPFPKALAKKSLDK 482
+ PI K ++ V + P +PFP K + K
Sbjct: 522 AEKAIEEPILDQPTRLLAPAASPLVEKPAATKTKDNVFVPPPYKPPLPFPGRFKKVMIQK 581
Query: 483 KFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQ 542
+ K L +++P D L +P K++KD++ +R +K V V+++ ECSAI+Q
Sbjct: 582 YKALLEKQLKNLEVTMPLVDCLALIPDSNKYVKDMITER--IKEVQGMVVLSHECSAIIQ 639
Query: 543 RKM-PKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQ 601
+K+ PKK DPGSFT+P + +A + LCDLGAS++LMPL++ ++L + P +SL
Sbjct: 640 QKIIPKKLGDPGSFTLPCALGPLAFNKCLCDLGASVSLMPLSVAKKLGFNKYKPCNISLI 699
Query: 602 MADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDK 661
+ADRS++ P+G++ED+ V + P DFVVL+M+E+ K PLILGRPFL T A IDV K
Sbjct: 700 LADRSVRIPHGLLEDLPVMIGMVEVPTDFVVLEMDEEPKDPLILGRPFLVTVGAIIDVKK 759
Query: 662 GHLILPVGKE-KVRFSVFNPMIETNHDNDFVCDVIRSRSKVSEETPKVNSQIDEFHPALI 720
G + L +G++ K+ F + N M + + + I +++E + + D AL
Sbjct: 760 GKIDLNLGRDLKMTFDITNTMKKPTIERNIFW--IEEMDMLADEMLEELGETDHLQSALT 817
Query: 721 K 721
K
Sbjct: 818 K 818
>UniRef100_Q9SHM3 F7F22.17 [Arabidopsis thaliana]
Length = 1799
Score = 291 bits (745), Expect = 5e-77
Identities = 231/782 (29%), Positives = 368/782 (46%), Gaps = 120/782 (15%)
Query: 41 ELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTA 100
E+R + N++ G +EDP H++ F R CN ++ VS D +L LFPFSL D A
Sbjct: 23 EIRKRRTTVNPGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKA 82
Query: 101 EEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKL 160
W + P SIT+W+D + F ++F A +L+N+I F+Q T E+ EAWERFK
Sbjct: 83 HIWEKNLPHDSITTWDDCKKVFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFKGY 142
Query: 161 LRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMNS 220
+CP H T+A ++ Y G+ R LD AS+G F + G+EL+E +A N
Sbjct: 143 TNQCPHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSDGNY 202
Query: 221 SND-RQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGGPHD 279
+ D + RG + + D+ K L+ +++ I + S+ V F+ ++
Sbjct: 203 NEDCDRTVRGTADSD--DKHRKEIKALNDKLDRI--------LLSQHKHVHFLVDDEQYE 252
Query: 280 SEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWR--------------QGSSG 325
++ E EEV + N YN N+PN S+R Q
Sbjct: 253 VQD-GEGNQLEEVSYIN--NNQGGYKGYNYFKTNNPNLSYRSTNVANPQDQVYPPQQQQS 309
Query: 326 QGNGF----QRQFPSQGFQGQSLRQP----------------------QERGEGEGSGSK 359
Q F Q P Q FQG +QP Q+ G+ S S
Sbjct: 310 QNKPFVPYNQGFVPKQQFQGNYQQQPPGFAPPQHQGPAAPDADMKQMLQQLLHGQASCSM 369
Query: 360 KSLEELVETFINRTENNYKNQEAANKNLENQF----GQLAKQIAERPQGIFPSDCIPNPK 415
+ +++ E N+ + +Y + + L+ + G A + P + NPK
Sbjct: 370 EMAKKISELH-NKLDCSYNDLNVKMETLDTKVRYLEGHSTSSSATKQTSQLPGKAVQNPK 428
Query: 416 QENASVVATRSGRVM-SELKKKTEGEKREEIVGGEV------------------------ 450
E A + RSG+ + + + KT E E+ G ++
Sbjct: 429 -EYAHAITLRSGKALPTREEPKTVTEDSEDQDGEDLSLEKDQADKPLDLSLEQPLDLSLQ 487
Query: 451 -----------------VVP-----------IKVREEVDLTPE-TSKIPFPKALAKKSLD 481
V+P +K +E+V + P S++PFP K D
Sbjct: 488 QSLDPPLDSFTRPTTRPVIPAASPTAPKPVAVKNKEKVFVPPPYKSQLPFPGRHKKALAD 547
Query: 482 KKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAIL 541
K + F K++ + IP DAL +P KF+KD++ + R++ V V+++ ECSAI+
Sbjct: 548 KYRAMFAKNIKEVELRIPLVDALALIPDSHKFLKDLIVE--RIQEVQGMVVLSHECSAII 605
Query: 542 QRK-MPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSL 600
Q+K +PKK DPGSFT+P + +A LCDLGAS++LMPL++ +RL + +SL
Sbjct: 606 QKKIIPKKLSDPGSFTLPCSLGPLAFNRCLCDLGASVSLMPLSVAKRLGFTQYKSCNISL 665
Query: 601 QMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVD 660
+ADRS++ P+G++E++ + + P DFVVL+M+E+ K PLILGRPFLAT A IDV
Sbjct: 666 ILADRSVRIPHGLLENLPIRIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVK 725
Query: 661 KGHLILPVGKE-KVRFSVFNPMIETNHDNDFVCDVIRSRSKVSEETPKVNSQIDEFHPAL 719
KG + L +GK+ ++ F V + M + + I ++++E + ++ D + AL
Sbjct: 726 KGKIDLNLGKDFRMTFDVKDAMKKPTIEGQLFW--IEEMDQLADELLEELAEEDHLNSAL 783
Query: 720 IK 721
K
Sbjct: 784 TK 785
>UniRef100_Q9LPB1 T32E20.9 [Arabidopsis thaliana]
Length = 1586
Score = 289 bits (739), Expect = 2e-76
Identities = 217/734 (29%), Positives = 342/734 (46%), Gaps = 113/734 (15%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G + P NFE++ LI +V N++ G +EDP H++ F R C+ ++ VS D +
Sbjct: 60 GIVPPPVQNNNFEIKSGLIAMVQSNKFHGLPMEDPLDHLDEFDRLCSLTKINRVSEDGFK 119
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A +W S PQGSITSW D + F +F + +L+NDI FTQ+ +E
Sbjct: 120 LRLFPFSLGDKAHQWEKSLPQGSITSWNDCKKAFLAKFFSNSRTARLRNDISGFTQTNNE 179
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYE 208
YEAWERFK +CP H + LD AS+G F + G+E
Sbjct: 180 TFYEAWERFKGYQTQCPHHEML-------------------LDTASNGNFLNKDVEDGWE 220
Query: 209 LIEKMA-------------IRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQN 255
++E +A IR + S+++ R + D+L+ ++ + + +
Sbjct: 221 VVENLAQSDGNYNEDYDRSIRTSSDSDEKHRREMKAMNDKLDKLLLMQQKHIHFLGDDET 280
Query: 256 QMKTTKIGSRVAKVEFVTCGG-------------PHDSEECTETRPEEEVKAMGQARNDP 302
++ +V +V G P+ S T ++ Q +N P
Sbjct: 281 LQVQDGETLQLEEVSYVQNQGGYNKGFNNFKQNHPNLSYRSTNVANRQDQVYPSQQQNQP 340
Query: 303 FSNT-YNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQP-------------Q 348
YN G P + QGN +Q+Q P GF Q +QP Q
Sbjct: 341 KPFVPYNQGQGYVPKQQY------QGN-YQQQLPPPGFT-QQQQQPALTTPDSDLKNMLQ 392
Query: 349 ERGEGEGSGSKKSLEELVETFINRTENNYKN----QEAANKNLENQFGQLAKQIAERPQG 404
+ +G+ +G+ + + E N+ + +Y + EA + GQ A + G
Sbjct: 393 QILQGQATGAMDLSKRMAEIH-NKVDCSYNDINIKVEALTSKIRYIEGQTGSTAAPKFTG 451
Query: 405 IFPSDCIPNPKQENASVVATRSGRVMSELKKKTE----------------GEKREEIVGG 448
PS + +E A + RSG+ + + + G E+ +
Sbjct: 452 --PSGKSMSNSKEYAHAITLRSGKELPTKESPNQNTEDSLDQDGEDFCQNGNSAEKAIEE 509
Query: 449 EVV------------------VPIKVREEVDLTPE-TSKIPFPKALAKKSLDKKFSKFVD 489
++ K +E V + P +PFP K + K +
Sbjct: 510 PILHQPTRPLAPAASPLVEKPAAAKTKENVFIPPPYKPPLPFPGRFKKVMIQKYKALLEK 569
Query: 490 VFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRK-MPKK 548
K L +++P D L +P K++KD++ R+K V V+++ ECSAI+Q+K +PKK
Sbjct: 570 QLKNLEVTMPLVDCLALIPDSNKYVKDMI--TERIKEVQGMVVLSHECSAIIQQKIIPKK 627
Query: 549 RRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLK 608
DPGSFT+P + +A + LCDLGAS++LMPL + ++L + P +SL +ADRS++
Sbjct: 628 LGDPGSFTLPCALGPLAFSKCLCDLGASVSLMPLPVAKKLGFNKYKPCNISLILADRSVR 687
Query: 609 TPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPV 668
+G++ED+ V + P DFVVL+M+E+ K PLILGRPFLA RA IDV KG + L +
Sbjct: 688 ISHGLLEDLPVMIGVVEVPTDFVVLEMDEEPKDPLILGRPFLARARAIIDVKKGKIDLNL 747
Query: 669 GKE-KVRFSVFNPM 681
G++ K+ F + N M
Sbjct: 748 GRDLKMTFDITNTM 761
>UniRef100_Q9M0T9 Putative athila transposon protein [Arabidopsis thaliana]
Length = 866
Score = 288 bits (737), Expect = 4e-76
Identities = 221/710 (31%), Positives = 350/710 (49%), Gaps = 68/710 (9%)
Query: 39 NFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRD 98
NFE++ LI+++ N++ G +EDP H++ F R CN ++ VS D +L LFPFSL D
Sbjct: 38 NFEIKSGLISMIQGNKFYGLPMEDPLDHLDEFDRLCNLTKINGVSADGFKLRLFPFSLGD 97
Query: 99 TAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFK 158
A W + P SI +W+D + F ++F A +L+N+I F+Q T E+ EAWERFK
Sbjct: 98 KAHIWEKNLPHDSIITWDDCKKAFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFK 157
Query: 159 KLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAM 218
+CP H T+A ++ Y G+ R LD AS+G F + G+EL++ +A
Sbjct: 158 GYTNQCPHHGFTKASMLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVDNLAQSDG 217
Query: 219 NSSNDRQARRGVLEV-EAYDQLMASNKQLSKQMNEIQ-NQMKTTKIGSRVAKVEFVTCGG 276
N + D R V + ++ D+ K L+ +++ I NQ K + V +F G
Sbjct: 218 NYNED--CDRTVRDTADSDDKHRKEIKALNDKLDRILLNQQK--HVHFLVDDEQFQVQDG 273
Query: 277 PHDSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWR--------------QG 322
E EEV + N YN N+PN S+R Q
Sbjct: 274 --------EGNQLEEVSYINN--NQSGYKGYNNFKTNNPNLSYRSTNIANPQDQVYPPQQ 323
Query: 323 SSGQGNGF----QRQFPSQGFQGQSLRQPQERGEGEGSGSKKSLEELVETFINRT-ENNY 377
Q F Q P Q FQG QP G+ +++ + + E ++ E+
Sbjct: 324 QQVQNKPFVPYNQGFVPKQQFQGNY--QPPPPPGGKALPTREEPKTVTEDSEDQDGEDLS 381
Query: 378 KNQEAANKNLENQFGQLAKQ----IAERPQGIFPSDCIPNPKQENASVVATRSGRVMSEL 433
++ A+K E Q +Q E+P + P D + P A+ + +
Sbjct: 382 LEKDQADKPHEQPLDQSLEQPLDLSLEQPLDL-PLDNVTRPTTRPIFPAASATAPKPITV 440
Query: 434 KKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSKFVDVFKK 493
K K + V VP P ++PFP K DK + F K+
Sbjct: 441 KNKEK-----------VFVP---------PPYKPELPFPGRHKKALADKYRAMFAKNIKE 480
Query: 494 LHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKM-PKKRRDP 552
+ + IP DAL +P KF+KD++ +R ++ V V+++ CSAI+Q+K+ PKK DP
Sbjct: 481 VELRIPLVDALALIPDSHKFLKDLIVER--IQEVQGMVVLSHGCSAIIQKKIIPKKLSDP 538
Query: 553 GSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYG 612
GSFT+P + +A LCDLGAS++LM L++ +RL + + +SL +ADRS++ P+G
Sbjct: 539 GSFTLPCSLGPLAFNRCLCDLGASVSLMALSVAKRLGFTQYKSSNISLILADRSVRIPHG 598
Query: 613 IVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVGKE- 671
++++ + + P DFVVL+M+E+ K PLILGRPFLAT A IDV KG + L +GK+
Sbjct: 599 LLKNFGITIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLSLGKDF 658
Query: 672 KVRFSVFNPMIETNHDNDFVCDVIRSRSKVSEETPKVNSQIDEFHPALIK 721
++ F V + M + + I+ ++++E + ++ D + AL K
Sbjct: 659 RMTFDVKDAMKKPTIEGQLFW--IKEMDQLADELLEELAEEDHLNSALTK 706
>UniRef100_Q7G6J5 Putative polyprotein [Oryza sativa]
Length = 689
Score = 282 bits (721), Expect = 3e-74
Identities = 205/683 (30%), Positives = 333/683 (48%), Gaps = 90/683 (13%)
Query: 36 GTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFS 95
G +F+L+ + I + + + G ED +AH+++F+ C+TY ++ VS D ++L LFPFS
Sbjct: 26 GDADFDLKSSHITMAQASPFCGKPNEDANAHLQQFLEICSTYTIKGVSPDVVKLRLFPFS 85
Query: 96 LRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWE 155
L A++W + + ++ +W+ + F ++F P EAWE
Sbjct: 86 LLGRAKQWFYAN-RTAVNTWDKCSTAFLSKFFP----------------------MEAWE 122
Query: 156 RFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAI 215
R ++ + CP + + FY+GL SR LDAA+ G F + Q ELIEKM
Sbjct: 123 RLQEYVAACPHLGMDDWLILQNFYNGLTPMSRDHLDAAARGAFFSKTVQGAVELIEKMVS 182
Query: 216 RAMNSSNDRQAR-RGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTC 274
S Q R RG+ V+ + L A L K++++ + + T + + + V C
Sbjct: 183 NMGWSEERLQTRQRGMHTVKETELLAAKLDLLMKRLDDHEKGPQRT-VKALDSHVMCEVC 241
Query: 275 GGP-HDSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQ 333
G H +C ET E AM N+ + GW N P + QG + N
Sbjct: 242 GNTGHSGNDCLETCEE----AMYMGNNNGYCPQGGQGW-NQPR-PYYQGGNNNSN----- 290
Query: 334 FPSQGFQGQSLRQPQERGEGEGSGSKKSLEELVETFINRTENNYKNQEAANKNLENQFGQ 393
FP+Q SL++LV T+ K A +K
Sbjct: 291 FPNQ----------------------PSLKDLVFAQAKTTDALSKKLAANDK-------- 320
Query: 394 LAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVMSE--------LKKKTEGEKREEI 445
LA + G P P+P EN + R G+ + + ++G +
Sbjct: 321 LASLVPANETGRIPGQ--PDPSIENVKAITMRRGKSTRDPPYPNPVGTNEISKGAPSNDS 378
Query: 446 VGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALE 505
EV V +E +T + FP+ + K S+D++F++FV+V +K+HI++P DA+
Sbjct: 379 ADKEVQPENTVPQEY---CDTRLLSFPQRMRKPSVDEQFARFVEVIQKIHINVPLLDAM- 434
Query: 506 QMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMA 565
Q+P YA+++KDIL+ +R L E V +TE+CS ++ K+P+K++DPG TI I
Sbjct: 435 QVPTYARYLKDILNNKRPLP-TTEVVKLTEQCSNLILHKLPEKKKDPGCPTITCSIGAQQ 493
Query: 566 DVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYV 625
+ C+LGAS++++ +F++LN + PT + LQ+AD S++ P GI EDV V + +
Sbjct: 494 FDQVSCNLGASVSVVLKDVFDKLNFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFF 553
Query: 626 FPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVGKEKVRFSVFNPMIETN 685
PVDFVVLDM+ ++ PLILGRPFL+T A IDV G + + +K +F F P E
Sbjct: 554 IPVDFVVLDMDTGKETPLILGRPFLSTAGANIDVGTGSIRFHINGKKEKFE-FQPRTEQ- 611
Query: 686 HDNDFVCDVIRSRSKVSEETPKV 708
C ++R + ++ + +V
Sbjct: 612 ------CSMVRIKHGLNPQNTQV 628
>UniRef100_Q9SHM5 F7F22.15 [Arabidopsis thaliana]
Length = 1862
Score = 272 bits (695), Expect = 3e-71
Identities = 216/707 (30%), Positives = 346/707 (48%), Gaps = 97/707 (13%)
Query: 41 ELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFSLRDTA 100
E+R + N++ G +EDP H++ F R CN ++ VS D +L LFPFSL D A
Sbjct: 158 EIRKRRTTVNPGNKFHGLPMEDPLDHLDEFDRLCNLTKINGVSEDGFKLRLFPFSLGDKA 217
Query: 101 EEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDENLYEAWERFKKL 160
W + P SIT+W+D + F ++F A +L+N+I F+Q T E+ EAWERFK
Sbjct: 218 HIWEKNLPHDSITTWDDCKKAFLSKFFSNARTARLRNEISGFSQKTGESFCEAWERFKGY 277
Query: 161 LRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMN- 219
+C H T+A ++ Y G+ R LD AS+G F + G+EL+E +A N
Sbjct: 278 TNQCSHHGFTKASLLSTLYRGVLPRIRMLLDTASNGNFQNKDVEEGWELVENLAQSNGNY 337
Query: 220 SSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGGPHD 279
+ N + RG + + D+ K L+ +++ I + S+ V F+ ++
Sbjct: 338 NENCDRTVRGTADSD--DKHRKEIKALNDKLDRI--------LLSQHKHVHFLVDDEQYE 387
Query: 280 SEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGF 339
++ E EEV + N YN N+PN S+R S+ N + +P Q
Sbjct: 388 VQD-GEGNQLEEVSYINN--NQGGYKGYNNFKTNNPNLSYR--STNVANPQDQVYPPQ-- 440
Query: 340 QGQSLRQPQERGEGEGSGSKKSLEELVETFINRTENNYKNQEAANKNLENQFGQLAKQIA 399
Q QS +P F+ + Q KQ +
Sbjct: 441 QQQSQNKP---------------------FV-------------------PYNQATKQTS 460
Query: 400 ERPQGIFPSDCIPNPKQENASVVATRSGRVMSELKK-KTEGEKREEIVGGEVVVPIKVRE 458
+ P + NPK E A + SG+ + ++ KT E E+ G ++ + +
Sbjct: 461 Q-----LPGKAVQNPK-EYAHAITLHSGKALPTREEPKTVTEDSEDQDGEDLSLKKDQAD 514
Query: 459 E-VDLTPETSKIPFPKALAKKSLDKKFSKF---------------------VDVFKKLHI 496
+ +DL+ E P +L ++SLD F V +K+ +
Sbjct: 515 KPLDLSLEQ---PLDLSL-QQSLDPPLDSFTRPTTRPVIPAASPTAPKPVAVKNKEKVEL 570
Query: 497 SIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKM-PKKRRDPGSF 555
IP DAL +P KF+KD++ +R ++ V V+++ ECSAI+Q+K+ PKK DPGSF
Sbjct: 571 RIPLVDALALIPDSHKFLKDLIVER--IQEVQGMVVLSHECSAIIQKKIIPKKLSDPGSF 628
Query: 556 TIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTPYGIVE 615
T+P + +A LCDLGAS++LMPL++ +RL + +SL +ADRS++ P+G++E
Sbjct: 629 TLPCSLGPLAFNRCLCDLGASVSLMPLSVAKRLGFTQYKSCNISLILADRSVRIPHGLLE 688
Query: 616 DVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVGKE-KVR 674
++ + + P DFVVL+M+E+ K PLILGRPFLAT A IDV KG + L +GK+ ++
Sbjct: 689 NLPIRIGAVEIPTDFVVLEMDEEPKDPLILGRPFLATAGAMIDVKKGKIDLNLGKDFRMT 748
Query: 675 FSVFNPMIETNHDNDFVCDVIRSRSKVSEETPKVNSQIDEFHPALIK 721
F V + M + + I ++++E + ++ D + AL K
Sbjct: 749 FDVKDAMKKPTIEGQLFW--IEEMDQLADELLEELAEEDHLNSALTK 793
>UniRef100_Q6F2G0 Hypothetical protein OSJNBa0083K01.26 [Oryza sativa]
Length = 585
Score = 229 bits (584), Expect = 2e-58
Identities = 183/596 (30%), Positives = 278/596 (45%), Gaps = 108/596 (18%)
Query: 139 IMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEF 198
I +F Q+ DE++ EAWE ++ + CP H + + FY+ L SR LDAA+ G F
Sbjct: 11 ISSFQQTRDESIPEAWELLQEYVAACPHHGMDDWLILQNFYNRLTPMSRDHLDAAAGGAF 70
Query: 199 DALLPQVGYELIEKMAIRAMNSSNDRQAR-RGVLEVEAYDQLMASNKQLSKQMNEIQNQM 257
+ Q ELIEKM S Q R RG+ V+ + L A L K+++ + +
Sbjct: 71 FSKTVQGTVELIEKMVSNMGWSKERLQTRQRGMHTVKETELLAAKLDLLMKRLDNHEKRP 130
Query: 258 KTTKIGSRVAKVEFVTCGGP-HDSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHPN 316
+ T + + + V CGG H +C ETR E AM N+ N Y P
Sbjct: 131 QGT-VKALDSHVTCEVCGGTDHSGNDCPETREE----AMYMGNNN---NGYRP------- 175
Query: 317 FSWRQGSSGQGNGFQRQFPSQGFQGQSLRQPQERGEGEGSGSKKSLEELVETFINRTENN 376
QG QG + QP+ +G
Sbjct: 176 --------------------QGVQGWN--QPRSYYQG----------------------- 190
Query: 377 YKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVMSE---- 432
N E Q QLA + G P P EN + TR G+ +
Sbjct: 191 -------GNNNETQLAQLAYLVPANETGRIPGQ--PYSSIENVKAITTRGGKSTRDPPYP 241
Query: 433 --------LKKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKF 484
K+ + ++ + E VP +E D T +PFP+ K S+D+ F
Sbjct: 242 NPAGTNGMSKETPSNDSADKEIQPEKTVP---QEYCD----TRLLPFPQWSRKTSVDELF 294
Query: 485 SKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRK 544
++FV+V +++HI++ DA+ Q+PIYA+++KDIL+ +R L E V +TE CS ++ K
Sbjct: 295 ARFVEVIQRIHINVLLLDAM-QVPIYARYLKDILNNKRPLP-TTEVVKLTEHCSNVILHK 352
Query: 545 MPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMAD 604
+P+K++ TI I +ALCDLGAS+++MP +F+RLN + PT + LQMAD
Sbjct: 353 LPEKKKYSECPTITCSIGAQQFDQALCDLGASVSVMPKDVFDRLNFTVLAPTPMRLQMAD 412
Query: 605 RSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHL 664
S++ GI EDV + + + PVDFVVLDM+ ++ PLILGRPFL+T A IDV+ +
Sbjct: 413 SSVRYQAGIAEDVPIKIWDFFIPVDFVVLDMDTRKETPLILGRPFLSTAGANIDVETSSI 472
Query: 665 ILPVGKEKVRFSVFNPMIETNHDNDFVCDVIRSR--------SKVSEETPKVNSQI 712
+ +++ +F F P E C ++R + +V E PK +S +
Sbjct: 473 CFHINEKEEKFE-FQPRTEQ-------CSMVRIKYGPNPQNIQEVEVEPPKTDSLV 520
>UniRef100_Q9SKR9 Putative Athila retroelement ORF1 protein [Arabidopsis thaliana]
Length = 929
Score = 228 bits (582), Expect = 4e-58
Identities = 197/687 (28%), Positives = 310/687 (44%), Gaps = 76/687 (11%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G ++ P NF+++ LI +V N++ G +EDP H++ R C ++ VS D +
Sbjct: 60 GIVLRPVQNNNFKIKSGLIAMVQGNKFHGLPMEDPLDHLDELERLCGLTKINGVSEDGFK 119
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A W + P+ SIT+W+D F +F + +L+N+I FT +E
Sbjct: 120 LRLFPFSLGDKAHLWEKTLPKNSITTWDDCKMAFLAKFFSNSRTARLRNEISGFTLKQNE 179
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYE 208
+ EAWERFK +CP H +QA + Y G+ R LD AS+G F + G+E
Sbjct: 180 SFCEAWERFKGYQTKCPHHGFSQASLLNTLYRGVLPKIRMLLDTASNGNFLKKDIEEGWE 239
Query: 209 LIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNE-IQNQMKTTKIGSRVA 267
L+E +A ++ + N+ R D+ K L+ ++++ IQ Q K
Sbjct: 240 LVENLA-QSGGNYNEDYGRSIYTSSNTDDKHHREMKALNDKLDKIIQMQQK--------- 289
Query: 268 KVEFVTCGGPH-----DSEECTETR------PEEEVKAMGQARNDPFSNTYNPGWRNHPN 316
V F++ P ++++C E R P A PF YN P
Sbjct: 290 HVHFISEDEPFQVQEGENDQCAEIRYVHNQGPSLPSAAAAAEPTKPFV-PYNQSLGFVPK 348
Query: 317 FSWRQGSSGQGNGFQRQFPSQGF--QGQSLRQPQERG---------EGEGSGSKKSLEEL 365
++ G+Q+Q P GF Q PQ +G+ +G+ + ++L
Sbjct: 349 QQFQ-------GGYQQQQPPPGFTPHQQQAHAPQNSDIMTVLQQLIQGQATGAMEIAKKL 401
Query: 366 --VETFINR--TENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQ-ENAS 420
V ++R E N K E+ + + G LA P I NPKQ A
Sbjct: 402 SKVNNKVDRQGVELNSK-FESMSTRMRYVEGILASPSVNNNPCQLPGKAIQNPKQYSTAH 460
Query: 421 VVATRSGRVMSELKKKTEGEKREEIVGGE-----VVVPIKVREEVDLTPETSKIPFPKAL 475
+ R + T + I+ GE V+ EE LT + ++ P A
Sbjct: 461 AITITHDRELPTRYVPTSNTEDSVILEGEDFYQDDVLADNPIEEPILTSQPTRPQAPPA- 519
Query: 476 AKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTE 535
F+K + K I +P +F K ++ K R L +
Sbjct: 520 ------TPFAKKTEAAKTKDIDFVPPPYKPLLPFPGRFKKVLVAKYRALLEKHIKDIPLV 573
Query: 536 ECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTP 595
+C A++ E + + LCDLGAS++LMPL++ ++L P
Sbjct: 574 DCLALIHD----------------EHKQLTFNNCLCDLGASVSLMPLSVVKKLGFVHYKP 617
Query: 596 TMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRA 655
L+L +ADR+ KTP+G++EDV V ++ P DFVVL+M+ + K PLILGRPFLA+ A
Sbjct: 618 CDLTLILADRTSKTPFGLLEDVPVMINGVEVPTDFVVLEMDGESKDPLILGRPFLASVGA 677
Query: 656 KIDVDKGHLILPVGKE-KVRFSVFNPM 681
IDV +G + L +G++ K++F + + M
Sbjct: 678 VIDVKQGKINLNLGEDVKMKFDIRDAM 704
>UniRef100_Q7XLB2 OSJNBa0011K22.17 protein [Oryza sativa]
Length = 551
Score = 219 bits (557), Expect = 3e-55
Identities = 167/543 (30%), Positives = 265/543 (48%), Gaps = 78/543 (14%)
Query: 135 LKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAAS 194
L+ I +F Q+ DE++ EAWER ++ + CP H + + FY+GL S LDAA+
Sbjct: 7 LRGRISSFQQTRDESIPEAWERLQEYMAACPHHGMDDWLILQNFYNGLTPMSHDHLDAAA 66
Query: 195 SGEFDALLPQVGYELIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQ 254
G F + Q +LIEKM D LM K+L + Q
Sbjct: 67 RGAFFSKTVQGAVDLIEKML----------------------DLLM---KRLDDHDKKPQ 101
Query: 255 NQMKTTKIGSRVAKVEFVTCGGP-HDSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRN 313
+K + V CG H +C +TR +AM N+ + GW
Sbjct: 102 GTVKVLD-----SHVTCEVCGNTGHSGNDCLDTR-----EAMYMGNNNGYCPQRGQGWNQ 151
Query: 314 HPNFSWRQGSSGQG--NGFQRQFPSQGF-QGQSLRQPQERGEGEGSGSKKSLEELVETFI 370
+ R+ + + Q F Q +++ ++ + + E + +
Sbjct: 152 PCPYVDRKHEAWEICLTPVQPSLKDLVFAQAKTIDALSKK-----LATNDKILENINVKL 206
Query: 371 NRTENNYKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVM 430
+ + ++NQ NK LE Q Q A + P E T + +
Sbjct: 207 DGFASAFQNQLRFNKMLETQLAQFASLV---------------PANE------TGTNGIA 245
Query: 431 SELKKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSKFVDV 490
E+ +K ++ E +VP +E D T + F + K S+D++F++F +V
Sbjct: 246 KEVPSSDSADKEVQL---EKIVP---QEYCD----TWLLSFHQRSRKPSVDEQFARFAEV 295
Query: 491 FKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKRR 550
+K+HI++P DA+ Q+P YA+++KD+L+ +R L + E V +TE+CS+++ K+P+K++
Sbjct: 296 IQKIHINVPLLDAM-QVPTYARYLKDMLNNKRLLPTM-EVVKLTEQCSSVILHKLPEKKK 353
Query: 551 DPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTP 610
DPG TI I +A CDLGASI++MP + ++LN +TPT++ LQ+AD S+ P
Sbjct: 354 DPGCPTITCSIGAQQFDQAFCDLGASISVMPKDVPDKLNFTVLTPTLMCLQLADLSVHYP 413
Query: 611 YGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPV-G 669
GI EDV V + + PVDFVVLDM+ ++ PLILGRPFL+T A IDV G + + G
Sbjct: 414 TGIAEDVPVKIRDFFIPVDFVVLDMDMGKETPLILGRPFLSTAGANIDVGMGSIRFDING 473
Query: 670 KEK 672
KEK
Sbjct: 474 KEK 476
>UniRef100_Q9MA70 F2J6.11 protein [Arabidopsis thaliana]
Length = 823
Score = 198 bits (504), Expect = 4e-49
Identities = 180/684 (26%), Positives = 282/684 (40%), Gaps = 140/684 (20%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G + P NFE++ LI+++ N++ G +EDP H++ F R C+ ++ VS D+ +
Sbjct: 64 GIVPPPIQNNNFEIKSGLISMIQSNKFHGLPMEDPLDHLDNFDRLCSLTKINGVSEDSFK 123
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A W + P S+ + +D + F +F + +L+N+I F Q E
Sbjct: 124 LRLFPFSLGDKAHLWEKTLPVDSVDTLDDCKKAFLAKFFSNSRTARLRNEISGFNQKNSE 183
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGLDAASSGEFDALLPQVGYE 208
+ EAWERFK +CP H +A ++ Y G R LD S+G F G+E
Sbjct: 184 SFAEAWERFKGYSTQCPHHGFKKASLLSTLYRGALPKIRMLLDTTSNGNFLNKDVAEGWE 243
Query: 209 LIEKMAI-------------RAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQN 255
L+E +A R + S DR RR L + N+++S + +
Sbjct: 244 LVENLAQSDGNYNENYDRTNRGSSDSEDRHKRRTKLLMIRLTNWCLLNREMSTTLQK--K 301
Query: 256 QMKTTKIGSRVAKVEFVTCGGPHDSEECTETRPEEEVKAMGQARNDPFSNTYNPGWRNHP 315
+ +K G + P D + +P + PF YN G+
Sbjct: 302 SLHNSKKGRILLLRRSNNVANPQDQVYPPQNQPPQA---------KPFI-PYNQGYNQKQ 351
Query: 316 NFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQP-QERGEGEGSGSKKSLEELVETFINRTE 374
NF GF +Q Q ++ + +G+ S S ++L E R +
Sbjct: 352 NFG--------PPGFTQQPQQTSAQDSEMKTLLHQLVQGQASCSMTMDKKLAE-LTTRID 402
Query: 375 NNYKNQ----EAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVM 430
+Y + +A N ++ G +A A + G P + + NPK E A + T S
Sbjct: 403 CSYNDLNIKIDALNTRVKTMEGHIASTSAPKHPGQLPRNSVQNPK-EYAHAITTVS---- 457
Query: 431 SELKKKTEGEKREEIVGGEVVVPIKVREEVDLT------------PETSKIP-------- 470
T I GEV+ P + R+E++L +IP
Sbjct: 458 ------TSATADSGIQEGEVLRP-RSRQEIELDFFAWLVERAYDPSNPIRIPPAYEPKPY 510
Query: 471 FPKALAK---KSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGV 527
FP+ +A+ + K F+ K+L IP D +++ + R +
Sbjct: 511 FPERIAQVNARIFQKHKMMFIKCIKELEEKIPLVDTPKEVIM------------ERPQEA 558
Query: 528 DETVLMTEECSAILQRK-MPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFE 586
+ V ++ ECSAI+QRK +PKK D
Sbjct: 559 QQIVELSFECSAIIQRKVIPKKLAD----------------------------------- 583
Query: 587 RLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILG 646
RS++ P+G++ED+ V + P DFVVL+M+E+ K PLILG
Sbjct: 584 ------------------RSVRVPHGMLEDLPVKIGSIEIPTDFVVLEMDEEPKDPLILG 625
Query: 647 RPFLATGRAKIDVDKGHLILPVGK 670
RPFLAT A IDV G + L +GK
Sbjct: 626 RPFLATAGALIDVQMGKIDLNLGK 649
>UniRef100_O22104 Vicia faba mRNA expressed from retrotransposon-like gene [Vicia
faba]
Length = 238
Score = 184 bits (468), Expect = 6e-45
Identities = 96/202 (47%), Positives = 141/202 (69%), Gaps = 3/202 (1%)
Query: 481 DKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAI 540
+K F F+++FKKL ++I +ALE+MP YAKFMKDI+ K+R + + ++ TE CS+I
Sbjct: 11 EKNFEIFLEMFKKLELNILVLEALEKMPTYAKFMKDIISKKRTID--TDPIIHTETCSSI 68
Query: 541 LQ-RKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLS 599
LQ K+P K++D G T P I + + +LGAS++L+P ++++L + V T ++
Sbjct: 69 LQGMKIPVKKKDRGFVTTPCTIGDRSFKKYFINLGASVSLLPFYIYKKLGISNVQNTRMT 128
Query: 600 LQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDV 659
L+ AD S+K PYGIVEDV+V +DK+VFPVDFVVL+M EDE+I LILGRPFL T R ID+
Sbjct: 129 LRFADHSVKIPYGIVEDVLVKIDKFVFPVDFVVLEMPEDEEITLILGRPFLETRRCLIDI 188
Query: 660 DKGHLILPVGKEKVRFSVFNPM 681
++G + L V E+++ V N M
Sbjct: 189 EEGTVTLKVYDEELKIDVQNTM 210
>UniRef100_Q9SHR2 F28L22.6 protein [Arabidopsis thaliana]
Length = 719
Score = 176 bits (445), Expect = 3e-42
Identities = 160/581 (27%), Positives = 262/581 (44%), Gaps = 94/581 (16%)
Query: 182 LQYSSRFGLDAASSGEFDALLPQVGYELIEKMA-------------IRAMNSSNDRQARR 228
L S+ LD AS+G F + G+E++E +A IR + S+++ R
Sbjct: 91 LALSTEMLLDTASNGNFLNKDVEDGWEVVENLAQSDGNYNEDYDRSIRTSSDSDEKHRRE 150
Query: 229 GVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTCGG------------ 276
+ D+L+ ++ + + + ++ +V +V G
Sbjct: 151 MKAMNDKLDKLLLMQQKHIHFLGDDETLQVQDGETLQLEEVSYVQNQGGYNKGFNNFKQN 210
Query: 277 -PHDSEECTETRPEEEVKAMGQARNDPFSNT-YNPGWRNHPNFSWRQGSSGQGNGFQRQF 334
P+ S T ++ Q +N P YN G P + QGN +Q+Q
Sbjct: 211 HPNLSYRSTNVANRQDQVYPSQQQNQPKPFVPYNQGQGYVPKQQY------QGN-YQQQL 263
Query: 335 PSQGFQGQSLRQP-------------QERGEGEGSGSKKSLEELVETFINRTENNYKN-- 379
P GF Q +QP Q+ +G+ +G+ + + E N+ + +Y +
Sbjct: 264 PPPGFTQQQ-QQPALTTPDSDLKNMLQQILQGQATGAMDLSKRMAEIH-NKVDCSYNDIN 321
Query: 380 --QEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVMSELKKKT 437
EA + GQ A + G PS + +E A + RSG+ + +
Sbjct: 322 IKVEALTSKIRYIEGQTGSTAAPKFTG--PSGKSMSNSKEYAHAITLRSGKELPTKESPN 379
Query: 438 EGEKREEIVGGE------------VVVPI----------------------KVREEVDLT 463
+ + GE + PI K +E V +
Sbjct: 380 QNTEDSLDQDGEDFCQNGNSAEKAIEEPILHQPTRPLAPAASPLVEKPAAAKTKENVFIP 439
Query: 464 PETSK-IPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRR 522
P +PFP K + K + K L +++P D L +P K++KD++ +R
Sbjct: 440 PPYKPPLPFPGRFKKVMIQKYKALLEKQLKNLEVTMPLVDCLALIPDSNKYVKDMITER- 498
Query: 523 RLKGVDETVLMTEECSAILQRKM-PKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMP 581
+K V V+++ ECSAI+Q+K+ PKK DPGSFT+P + +A + LCDLGAS++LMP
Sbjct: 499 -IKEVQGMVVLSHECSAIIQQKIIPKKLGDPGSFTLPCALGPLAFSKCLCDLGASVSLMP 557
Query: 582 LTMFERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKI 641
L + ++L + P +SL +ADRS++ +G++ED+ V + P DFVVL+M+E+ K
Sbjct: 558 LPVAKKLGFNKYKPCNISLILADRSVRISHGLLEDLPVMIGVVEVPTDFVVLEMDEEPKD 617
Query: 642 PLILGRPFLATGRAKIDVDKGHLILPVGKE-KVRFSVFNPM 681
PLILGRPFLA RA IDV KG + L +G++ K+ F + N M
Sbjct: 618 PLILGRPFLARARAIIDVKKGKIDLNLGRDLKMTFDITNTM 658
>UniRef100_Q8S8J9 Putative Athila retroelement ORF1 protein [Arabidopsis thaliana]
Length = 1012
Score = 175 bits (443), Expect = 5e-42
Identities = 163/624 (26%), Positives = 269/624 (42%), Gaps = 121/624 (19%)
Query: 29 GSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERFIRNCNTYRVQNVSTDTIR 88
G + P NF+++ LI+++ N++ G +EDP H++ F R C+ ++ VS ++ +
Sbjct: 23 GIVPPPIQNNNFKIKSDLISMIQGNKFHGLPMEDPLDHLDNFDRLCSLTKINGVSEESFK 82
Query: 89 LSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRALLRKLKNDIMTFTQSTDE 148
L LFPFSL D A W + P SI +W+D + F +F + +L+++I F Q E
Sbjct: 83 LRLFPFSLGDKAHLWEKTLPVKSIDTWDDCKKAFLAKFFSNSRKARLRSEISGFNQKNSE 142
Query: 149 NLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQY--SSRFGLDAASSGEFDALLPQVG 206
+ EAWERFK +CP H E + Y L+ R LD AS+G G
Sbjct: 143 SFSEAWERFKGYTTQCPHH-----ESLPPQYSILRCLPKIRMLLDTASNGNILNKDVAEG 197
Query: 207 YELIEKMAIRAMNSSNDRQARRGVLEVEAYDQLMASNKQLSKQMNEIQNQMKTTKIGSRV 266
+EL+E +A N + D YD+ +N+ S ++ + ++KT + R+
Sbjct: 198 WELVENLAQSHRNYNED------------YDR---TNRGSSDSEDKHKKEIKT--LNDRI 240
Query: 267 AKVEFVTCGGPH--DSEECTETRPEEE--VKAMGQARNDPFSNTYNPGWRN----HPNFS 318
K+ + EE T+ + E ++ + +N YN G+ N HPN S
Sbjct: 241 DKLVLAQQRNVYYITEEELTQLQDGENLTIEEVSYLQN---QGGYNKGFNNYKPPHPNLS 297
Query: 319 WRQGS-----------------------SGQGNGFQRQFPSQGFQGQSLRQ--------- 346
+R + QG ++ F GF Q +
Sbjct: 298 YRSNNVANPQDQVYPPQNQPTQAKPFVPYNQGYNQKQNFGPPGFTQQPQQPSAQDSEMKT 357
Query: 347 -PQERGEGEGSGSKKSLEELVETFINRTENNYKNQ----EAANKNLENQFGQLAKQIAER 401
PQ+ +G S S ++L E R + +Y + +A N +++ G +A A +
Sbjct: 358 LPQQLVQGHASCSMTMDKKLAE-LTTRIDCSYNDLNIKIDALNTRVKSMEGHIASTSAPK 416
Query: 402 PQGIFPSDCIPNPKQENASVVATRSGRVMSELKKKTEGEKREEIVGGEVVVPIKVREEVD 461
G P + NPK E A ++T T + I GEV+ P + R+E++
Sbjct: 417 HLGQLPGKAVQNPK-EYAHAIST----------VNTSATEDSGIQEGEVLRP-RSRQEIE 464
Query: 462 L--------------------TPETSKIPFPKALA---KKSLDKKFSKFVDVFKKLHISI 498
L +P K FP+ +A + K F+ K+L +
Sbjct: 465 LDFFARLVERAHDPSNPIHIPSPYVPKPAFPERIAQIDEMIFQKHKMMFIKCIKELEEKV 524
Query: 499 PFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRK-MPKKRRDPGSFTI 557
P D ++M + R + + V ++ ECSAI+Q+K +PKK DPGSFT+
Sbjct: 525 PLVDTPKEMIM------------ERPQEAQQIVELSFECSAIIQKKVIPKKLGDPGSFTL 572
Query: 558 PVEIEGMADVEALCDLGASINLMP 581
P + + +LCDLGAS++ P
Sbjct: 573 PCSLAPLVFNNSLCDLGASMDEEP 596
>UniRef100_Q8S6J2 Hypothetical protein OSJNBa0019N10.34 [Oryza sativa]
Length = 237
Score = 171 bits (432), Expect = 9e-41
Identities = 88/199 (44%), Positives = 132/199 (66%), Gaps = 2/199 (1%)
Query: 466 TSKIPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLK 525
T +PFP+ K S+D++F+ FV+V +K+HI +P DA+ Q+P YA ++KDIL+ +R L
Sbjct: 30 TRLLPFPQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAM-QVPTYACYLKDILNNKRPLP 88
Query: 526 GVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMF 585
E V +T +CS ++ K+P+K++DPG TI I +ALCDLGAS+++MP +F
Sbjct: 89 -TTEVVKLTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVF 147
Query: 586 ERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLIL 645
++LN + PT + LQ+AD S++ P GI EDV V + + VDFVVLDM+ +++ +IL
Sbjct: 148 DKLNFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIIL 207
Query: 646 GRPFLATGRAKIDVDKGHL 664
GRPFL T A I V G +
Sbjct: 208 GRPFLGTAGANIHVGTGSI 226
>UniRef100_Q7XF80 Hypothetical protein [Oryza sativa]
Length = 725
Score = 167 bits (424), Expect = 8e-40
Identities = 86/192 (44%), Positives = 129/192 (66%), Gaps = 2/192 (1%)
Query: 466 TSKIPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLK 525
T +PFP+ K S+D++F+ FV+V +K+HI +P DA+ Q+P YA ++KDIL+ +R L
Sbjct: 61 TRLLPFPQRSRKPSVDEQFAHFVEVIQKIHIDVPLLDAM-QVPTYACYLKDILNNKRPLP 119
Query: 526 GVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMF 585
E V +T +CS ++ K+P+K++DPG TI I +ALCDLGAS+++MP +F
Sbjct: 120 -TTEVVKLTLQCSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKYVF 178
Query: 586 ERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLIL 645
++LN + PT + LQ+AD S++ P GI EDV V + + VDFVVLDM+ +++ +IL
Sbjct: 179 DKLNFTVLAPTPMRLQLADSSVRYPAGIAEDVPVKIRDFFILVDFVVLDMDTGKEMSIIL 238
Query: 646 GRPFLATGRAKI 657
GRPFL T A I
Sbjct: 239 GRPFLGTAGANI 250
>UniRef100_Q8LM79 Hypothetical protein OSJNAa0082N11.11 [Oryza sativa]
Length = 322
Score = 167 bits (422), Expect = 1e-39
Identities = 105/306 (34%), Positives = 163/306 (52%), Gaps = 41/306 (13%)
Query: 377 YKNQEAANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVMSELKKK 436
++NQ + NK +E Q QLA + P P+ EN + TR G
Sbjct: 37 FQNQLSFNKMIETQLAQLASLVPANESERIPGQ--PDSSIENIKAITTRGG--------- 85
Query: 437 TEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLD-------KKFSKFVD 489
T G +E P + ++ PE K + ++ D ++F++ V+
Sbjct: 86 TNGMSKE--------TPSTDLADKEIQPE-------KTVPQEYCDTRGVGNRQQFARSVE 130
Query: 490 VFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKR 549
V +K+HI++P DA+ Q+P YA+++KDIL+ +R L TE CS ++ K+P+K+
Sbjct: 131 VIQKIHINVPLLDAM-QVPTYARYLKDILNNKRPLP-------TTEHCSNVILHKLPEKK 182
Query: 550 RDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKT 609
+DP TI I +ALCDLGAS+++MP +F++LN + PT + LQ+AD S+
Sbjct: 183 KDPRCPTITCSIGAQQFDQALCDLGASVSVMPKDVFDKLNFTVLAPTPMRLQLADSSVHC 242
Query: 610 PYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLILGRPFLATGRAKIDVDKGHLILPVG 669
P GI EDV V + + PVDFVVLDM+ ++ LILG PFL+T A IDV G + +
Sbjct: 243 PVGIEEDVPVKIWDFFIPVDFVVLDMDTGKETSLILGHPFLSTAGANIDVGMGSIRFHIN 302
Query: 670 KEKVRF 675
++ +F
Sbjct: 303 GKEEKF 308
>UniRef100_Q6UUB7 Hypothetical protein [Oryza sativa]
Length = 450
Score = 160 bits (404), Expect = 2e-37
Identities = 136/436 (31%), Positives = 208/436 (47%), Gaps = 69/436 (15%)
Query: 178 FYDGLQYSSRFGLDAASSGEFDALLPQVGYELIEKMAIRAMNSSNDRQA-RRGVLEVEAY 236
FY+GL SR LDAA+ G F + + Q +L+EKM S Q +RG+ V+
Sbjct: 74 FYNGLTPMSRDHLDAAAGGAFFSKMVQGAVDLVEKMVSNMGWSEEQLQTCQRGMHTVKET 133
Query: 237 DQLMASNKQLSKQMNEIQNQMKTTKIGSRVAKVEFVTC----GGPHDSEECTETRPEEEV 292
+ L A L K+++ + + + G+ A VTC H +C ETR EE
Sbjct: 134 ELLAAKLDLLMKRLDNHEKRPQ----GTVKALDSHVTCEVCSSTGHSGNDCPETR--EEA 187
Query: 293 KAMGQARNDPFSNTYNPGWRNHPNFSWRQGSSGQGNGFQRQFPSQGFQGQSLRQPQERGE 352
MG +N Y P QG QG S +P +G
Sbjct: 188 MYMGN------NNWYRP---------------------------QGGQGWSQLRPYYQG- 213
Query: 353 GEGSGS---KKSLEELVETFINRTENNYKNQEAANKNLENQ-FGQLAKQIAERPQGIFPS 408
G +G+ + SL++LV + T+ K +K LEN QLA + G P
Sbjct: 214 GNNNGNFSNQPSLKDLVFSQAKTTDALSKKLATNDKILENNNLAQLASLVPVNETGRIPG 273
Query: 409 DCIPNPKQENASVVATRSGRVMSELKKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSK 468
P+ EN + R K+ + EK VP +E D T
Sbjct: 274 Q--PDSSIENVKAITMRGASSSDSADKEDQPEK---------TVP---QEYCD----TWL 315
Query: 469 IPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVD 528
+ FP+ + K +D++F++FV+V +K+HI++P DA+ QMP YA+++K IL+ +R L
Sbjct: 316 LLFPQRMRKPLVDEQFARFVEVIQKIHINVPLLDAM-QMPTYARYLKGILNNKRPLP-TT 373
Query: 529 ETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERL 588
E V +TE+ S ++ K+P+K++DPG TI I +ALCDLGAS+++MP +F++L
Sbjct: 374 EVVKLTEQWSNVILHKLPEKKKDPGCPTITCSIGAQQFDQALCDLGASVSVMPKNVFDKL 433
Query: 589 NLGEVTPTMLSLQMAD 604
N + PT + LQ+AD
Sbjct: 434 NFTMLAPTPMHLQLAD 449
>UniRef100_Q7XWD0 OSJNBb0069N01.18 protein [Oryza sativa]
Length = 975
Score = 158 bits (400), Expect = 5e-37
Identities = 97/262 (37%), Positives = 146/262 (55%), Gaps = 17/262 (6%)
Query: 382 AANKNLENQFGQLAKQIAERPQGIFPSDCIPNPKQENASVVATRSGRVMSE--------- 432
+ NK +E Q QLA + + G P P P EN + TR G+ +
Sbjct: 278 SCNKRIETQLAQLAALVPAKETGRIPMQ--PEPSLENVKAITTRGGKSTRDPPYSNIAGT 335
Query: 433 --LKKKTEGEKREEIVGGEVVVPIKVREEVDLTPETSKIPFPKALAKKSLDKKFSKFVDV 490
K+ E+ E V P K+ + +T+ +PFP+ K S+D++ ++F+DV
Sbjct: 336 RQAGKEARSASTTELEENEEVQPDKIPPQEYC--DTTLLPFPQRQRKPSVDEQSARFMDV 393
Query: 491 FKKLHISIPFADALEQMPIYAKFMKDILHKRRRLKGVDETVLMTEECSAILQRKMPKKRR 550
+K+HI+IPF DA+ ++P YA+++KDIL+ +R L E V +TEECS + ++ +K++
Sbjct: 394 TQKIHINIPFVDAM-RVPTYARYLKDILNNKRALP-TTEMVNLTEECSNAILHRLREKKK 451
Query: 551 DPGSFTIPVEIEGMADVEALCDLGASINLMPLTMFERLNLGEVTPTMLSLQMADRSLKTP 610
DPG TI I ALCDLGAS++ M +F++LN + PT + LQ+AD S+ P
Sbjct: 452 DPGCPTISCLIGIQHFDHALCDLGASVSEMRKDVFDKLNYTVLIPTPMRLQLADSSVYYP 511
Query: 611 YGIVEDVMVWVDKYVFPVDFVV 632
GI EDV V V + FPVDFVV
Sbjct: 512 AGIAEDVPVKVRDFFFPVDFVV 533
Score = 81.3 bits (199), Expect = 1e-13
Identities = 55/214 (25%), Positives = 96/214 (44%), Gaps = 25/214 (11%)
Query: 11 ANRSLRDLTSAAMSYDYPGSIVSPDGTGNFELRPALINLVSQNQYGGGALEDPHAHMERF 70
A ++LR+ + + G V+ T +L+ +L+ + + + G ED H+++F
Sbjct: 2 AEKTLREFAAPSADNVTVGPAVNIGDT-ILDLKCSLVTMAQASPFCGKPNEDASTHLQQF 60
Query: 71 IRNCNTYRVQNVSTDTIRLSLFPFSLRDTAEEWLNSQPQGSITSWEDLAEKFTTRFVPRA 130
+ C+TY V+ V D ++ +W+ + F ++F P
Sbjct: 61 LEICSTYTVEGVIPDA-----------------------ATVNTWDRCSTAFLSKFFPMD 97
Query: 131 LLRKLKNDIMTFTQSTDENLYEAWERFKKLLRRCPQHNLTQAEQVAKFYDGLQYSSRFGL 190
L+ I +F Q+ DE + EAWE ++ + CP H + + FY+ L SSR L
Sbjct: 98 KTNALRGRISSFQQTRDEMIPEAWEHLQEYVAACPYHGMDDWLILQSFYNELTPSSRDHL 157
Query: 191 DAASSGEFDALLPQVGYELIEKMAIRAMNSSNDR 224
DA + G F + +LIEKM + M S +R
Sbjct: 158 DATTRGAFFLKTVRAAIDLIEKM-VSNMGWSEER 190
>UniRef100_Q6UUL4 Hypothetical protein [Oryza sativa]
Length = 731
Score = 155 bits (393), Expect = 3e-36
Identities = 81/187 (43%), Positives = 123/187 (65%), Gaps = 3/187 (1%)
Query: 465 ETSKIPFPKALAKKSLDKKFSKFVDVFKKLHISIPFADALEQMPIYAKFMKDILHKRRRL 524
+T +PF + K S+D++F++FV+V +K+HI++P D + QMP YA+++KD L+ +R L
Sbjct: 160 DTRLLPFSERSRKPSVDEQFARFVEVIQKIHINVPLLDTI-QMPTYARYLKDKLNNKRPL 218
Query: 525 KGVDETVLMTEECSAILQRKMPKKRRDPGSFTIPVEIEGMADVEALCDLGASINLMPLTM 584
E V +TE+CS I+ K+PKK+ DPG TI I +ALCDLGA++++MP +
Sbjct: 219 P--TEVVKLTEQCSNIILHKLPKKKEDPGCPTITCSIGAQQFDQALCDLGANVSVMPKDV 276
Query: 585 FERLNLGEVTPTMLSLQMADRSLKTPYGIVEDVMVWVDKYVFPVDFVVLDMEEDEKIPLI 644
F++LN PT + LQ+A+ S++ P GI EDV V + + PV FVVLD + ++ PLI
Sbjct: 277 FDKLNFTVWAPTPMHLQLANSSVRYPAGIAEDVPVKIRDFFIPVGFVVLDTDTGKETPLI 336
Query: 645 LGRPFLA 651
LG LA
Sbjct: 337 LGALSLA 343
Score = 48.5 bits (114), Expect = 7e-04
Identities = 19/35 (54%), Positives = 28/35 (79%)
Query: 61 EDPHAHMERFIRNCNTYRVQNVSTDTIRLSLFPFS 95
ED +AH+++F+ C+TY ++ VS DT+RL LFPFS
Sbjct: 13 EDANAHLQQFLEICSTYTIKGVSPDTVRLRLFPFS 47
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.133 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,230,299,972
Number of Sequences: 2790947
Number of extensions: 54337455
Number of successful extensions: 155233
Number of sequences better than 10.0: 728
Number of HSP's better than 10.0 without gapping: 455
Number of HSP's successfully gapped in prelim test: 278
Number of HSP's that attempted gapping in prelim test: 153902
Number of HSP's gapped (non-prelim): 1332
length of query: 728
length of database: 848,049,833
effective HSP length: 135
effective length of query: 593
effective length of database: 471,271,988
effective search space: 279464288884
effective search space used: 279464288884
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0388a.1


