

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0380.7
(155 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
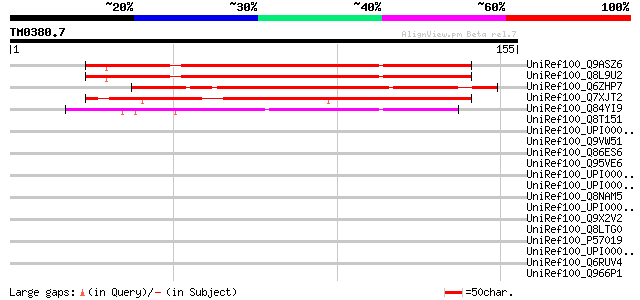
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9ASZ6 AT4g35320/F23E12_120 [Arabidopsis thaliana] 120 8e-27
UniRef100_Q8L9U2 Hypothetical protein [Arabidopsis thaliana] 117 7e-26
UniRef100_Q6ZHP7 Hypothetical protein OJ1191_G08.14 [Oryza sativa] 106 2e-22
UniRef100_Q7XJT2 At2g17300 protein [Arabidopsis thaliana] 102 4e-21
UniRef100_Q84YI9 Hypothetical protein OSJNBa0016N23.133 [Oryza s... 94 1e-18
UniRef100_Q8T151 Similar to Vibrio vulnificus CMCP6. TPR repeat ... 35 0.44
UniRef100_UPI000042CC78 UPI000042CC78 UniRef100 entry 35 0.76
UniRef100_Q9VW51 CG32217-PA [Drosophila melanogaster] 34 0.99
UniRef100_Q86ES6 Clone ZZD1473 mRNA sequence [Schistosoma japoni... 34 0.99
UniRef100_Q95VE6 ELL [Drosophila melanogaster] 34 0.99
UniRef100_UPI00003639A6 UPI00003639A6 UniRef100 entry 33 1.7
UniRef100_UPI000042F81C UPI000042F81C UniRef100 entry 33 1.7
UniRef100_Q8NAM5 Hypothetical protein FLJ35107 [Homo sapiens] 33 1.7
UniRef100_UPI0000363B09 UPI0000363B09 UniRef100 entry 33 2.2
UniRef100_Q9X2V2 Serum opacity factor [Streptococcus pyogenes] 33 2.2
UniRef100_Q8LTG0 GtrC [Bacteriophage P22-pbi] 33 2.2
UniRef100_P57019 Hypothetical 55.3 kDa protein in gtrB 5'region ... 33 2.2
UniRef100_UPI000042C858 UPI000042C858 UniRef100 entry 33 2.9
UniRef100_Q6RUV4 C3 phosphoenolpyruvate carboxylase [Setaria ita... 33 2.9
UniRef100_Q966P1 Hypothetical protein C46C11.1 [Caenorhabditis e... 33 2.9
>UniRef100_Q9ASZ6 AT4g35320/F23E12_120 [Arabidopsis thaliana]
Length = 147
Score = 120 bits (302), Expect = 8e-27
Identities = 62/122 (50%), Positives = 84/122 (68%), Gaps = 8/122 (6%)
Query: 24 LPVSH----SRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGT 79
LPVS + + D +R+RLSS + S +A+ + FPRSKS+S+MG+ AG+
Sbjct: 16 LPVSQRPNAAATADDEPGLIRRRLSSLSLN---LSNQPAAIAARFPRSKSVSAMGEQAGS 72
Query: 80 SIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPP 139
S+K WW+W W+WILSRKPIF RDLELN E K + GS NRG+ HVF+KLRS+IR F+ P
Sbjct: 73 SVKEWWEWGWSWILSRKPIFIRDLELNKDEAKSI-GSQNRGSIMHVFFKLRSQIRNFMGP 131
Query: 140 TN 141
++
Sbjct: 132 SS 133
>UniRef100_Q8L9U2 Hypothetical protein [Arabidopsis thaliana]
Length = 147
Score = 117 bits (294), Expect = 7e-26
Identities = 61/122 (50%), Positives = 83/122 (68%), Gaps = 8/122 (6%)
Query: 24 LPVSH----SRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGT 79
LPVS + + D +R+RLSS + S +A+ + F RSKS+S+MG+ AG+
Sbjct: 16 LPVSQRPNAAATADDEPGLIRRRLSSLSLN---LSNQPAAIAARFARSKSVSAMGEQAGS 72
Query: 80 SIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPP 139
S+K WW+W W+WILSRKPIF RDLELN E K + GS NRG+ HVF+KLRS+IR F+ P
Sbjct: 73 SVKEWWEWGWSWILSRKPIFIRDLELNKDEAKSI-GSQNRGSIMHVFFKLRSQIRNFMGP 131
Query: 140 TN 141
++
Sbjct: 132 SS 133
>UniRef100_Q6ZHP7 Hypothetical protein OJ1191_G08.14 [Oryza sativa]
Length = 183
Score = 106 bits (264), Expect = 2e-22
Identities = 56/112 (50%), Positives = 74/112 (66%), Gaps = 7/112 (6%)
Query: 38 GLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNWWDWTWAWILSRKP 97
GLRKRLSSF+ S SASA W+ F RSKS S+G +AG +K WWDW W++S+KP
Sbjct: 45 GLRKRLSSFSDKIQPIS-SASAEWA-FRRSKSAPSLGAFAGGPLKRWWDWGVGWLMSKKP 102
Query: 98 IFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPPTNSHHHLPRT 149
FA DLE+N++E LG +RG+W H+ YK+RS +RR + + H LP T
Sbjct: 103 GFATDLEMNEEEVAALGRG-SRGSWGHILYKMRSGVRRLV----TSHSLPTT 149
>UniRef100_Q7XJT2 At2g17300 protein [Arabidopsis thaliana]
Length = 139
Score = 102 bits (253), Expect = 4e-21
Identities = 56/120 (46%), Positives = 78/120 (64%), Gaps = 11/120 (9%)
Query: 24 LPVSHSRSVDSSEHGL-RKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIK 82
LP+S S + E GL R+RLSS + + S + RSKS+S MG+ G+S+K
Sbjct: 17 LPIS---STTTDEPGLLRRRLSSLSLNLSRNQPSTVS------RSKSVSDMGEQGGSSVK 67
Query: 83 NWWDWTWAWILSRK-PIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSEIRRFIPPTN 141
WW+W+W+WIL +K PIF DLE+N ETK G+ RG++ HVF+KLRSEIRR + P++
Sbjct: 68 EWWEWSWSWILLKKLPIFFTDLEVNKNETKSSLGNQQRGSFIHVFFKLRSEIRRLLRPSS 127
>UniRef100_Q84YI9 Hypothetical protein OSJNBa0016N23.133 [Oryza sativa]
Length = 173
Score = 94.0 bits (232), Expect = 1e-18
Identities = 49/125 (39%), Positives = 75/125 (59%), Gaps = 7/125 (5%)
Query: 18 TSNNKQLPVSHSRSVD-SSEH--GLRKRLSSFNTT--PAAASASASAVWSSFPRSKSLSS 72
++ + LPV+++ +H GLR+R+SSF+ P +SA A + S+ +
Sbjct: 15 SAERENLPVTNATGKGRGGDHLIGLRRRMSSFSVRIQPLMSSAGAGGAFRRATSMPSVKA 74
Query: 73 MGDYAGTSIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLRSE 132
+ AG +++ WW W W+++R+P FARDLE+ND E L G H RGTWRHVFY+LR+
Sbjct: 75 LAAQAG-AVRRWWGWGLGWVMNRRPAFARDLEMNDDEAAAL-GCHCRGTWRHVFYRLRAG 132
Query: 133 IRRFI 137
RR +
Sbjct: 133 ARRLL 137
>UniRef100_Q8T151 Similar to Vibrio vulnificus CMCP6. TPR repeat containing protein
[Dictyostelium discoideum]
Length = 856
Score = 35.4 bits (80), Expect = 0.44
Identities = 25/78 (32%), Positives = 39/78 (49%)
Query: 1 MSSIGHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAV 60
MSS G + SFF +NN S S SS +K SS +++ +++S+S++A+
Sbjct: 1 MSSSGKTEKMSFFKKMGNNNNSGSSGISSGSSGSSSSSKKKSSSSSSSSSSSSSSSSTAI 60
Query: 61 WSSFPRSKSLSSMGDYAG 78
+S KS S D G
Sbjct: 61 PTSITNIKSRSQSTDGLG 78
>UniRef100_UPI000042CC78 UPI000042CC78 UniRef100 entry
Length = 591
Score = 34.7 bits (78), Expect = 0.76
Identities = 17/38 (44%), Positives = 26/38 (67%)
Query: 32 VDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKS 69
+D++ GL+KR S +TT +AAS+SAS ++ P KS
Sbjct: 112 IDAANAGLKKRSDSVSTTQSAASSSASPSSAASPNKKS 149
>UniRef100_Q9VW51 CG32217-PA [Drosophila melanogaster]
Length = 1059
Score = 34.3 bits (77), Expect = 0.99
Identities = 19/64 (29%), Positives = 31/64 (47%)
Query: 18 TSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYA 77
+ NN S S S +S+ H + S FN +S+S+SA S P +L ++G
Sbjct: 223 SGNNNNRSSSSSNSFNSNNHSRKLGSSPFNGLGVGSSSSSSAFASRSPNPSTLGAIGTVN 282
Query: 78 GTSI 81
G+ +
Sbjct: 283 GSGV 286
>UniRef100_Q86ES6 Clone ZZD1473 mRNA sequence [Schistosoma japonicum]
Length = 271
Score = 34.3 bits (77), Expect = 0.99
Identities = 35/136 (25%), Positives = 55/136 (39%), Gaps = 5/136 (3%)
Query: 11 SFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSL 70
S F+ + N+ P S S +D E GL + S++ P SA S + SK
Sbjct: 111 SVFSASSLRFNRHRPASVSGYLD--ECGLNR---SYDLYPVPLSARPSTSFGQLHGSKRY 165
Query: 71 SSMGDYAGTSIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSHNRGTWRHVFYKLR 130
S+ TS +N ++ I +P+ + L ++ + L + RH
Sbjct: 166 KSLDRGCYTSCQNLSSKPFSTIKMNRPLNRQPLTIHPHSDRRLRELQHMNYLRHGNRVDG 225
Query: 131 SEIRRFIPPTNSHHHL 146
R +IP T S HHL
Sbjct: 226 EFERTYIPQTYSSHHL 241
>UniRef100_Q95VE6 ELL [Drosophila melanogaster]
Length = 1060
Score = 34.3 bits (77), Expect = 0.99
Identities = 19/64 (29%), Positives = 31/64 (47%)
Query: 18 TSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYA 77
+ NN S S S +S+ H + S FN +S+S+SA S P +L ++G
Sbjct: 223 SGNNNNSSSSSSNSFNSNNHSRKLGSSPFNGLGVGSSSSSSAFASRSPNPSTLGAIGTVN 282
Query: 78 GTSI 81
G+ +
Sbjct: 283 GSGV 286
>UniRef100_UPI00003639A6 UPI00003639A6 UniRef100 entry
Length = 184
Score = 33.5 bits (75), Expect = 1.7
Identities = 23/79 (29%), Positives = 40/79 (50%)
Query: 2 SSIGHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVW 61
SS H S + ++S+N S S S SS R+ S+ +++ +++S+S+S+
Sbjct: 88 SSSSSSSHSSNISSSSSSSNSSSSSSSSSSSSSSSSSSRRSNSNSSSSSSSSSSSSSSSS 147
Query: 62 SSFPRSKSLSSMGDYAGTS 80
SS S SS G + +S
Sbjct: 148 SSSSSCSSSSSSGSSSSSS 166
>UniRef100_UPI000042F81C UPI000042F81C UniRef100 entry
Length = 443
Score = 33.5 bits (75), Expect = 1.7
Identities = 25/61 (40%), Positives = 32/61 (51%), Gaps = 1/61 (1%)
Query: 23 QLPVSHSRSVDSSEHG-LRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSI 81
Q P S S + DSS L S+ T+ AAA++SA+A SS S S +S G Y I
Sbjct: 142 QQPSSSSSTEDSSTAASLTSVASAAATSSAAATSSAAATSSSAATSSSSNSTGSYTPNGI 201
Query: 82 K 82
K
Sbjct: 202 K 202
>UniRef100_Q8NAM5 Hypothetical protein FLJ35107 [Homo sapiens]
Length = 258
Score = 33.5 bits (75), Expect = 1.7
Identities = 24/68 (35%), Positives = 38/68 (55%), Gaps = 3/68 (4%)
Query: 14 TIPTTSNNKQLPVS-HSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSS 72
+IP++S++ P S HS S SS H SS +T+ ++S+S+S+ SS P S + SS
Sbjct: 54 SIPSSSSSSSSPSSSHSSSSPSSSHSSSSPSSSSSTSSPSSSSSSSS--SSSPSSSNSSS 111
Query: 73 MGDYAGTS 80
+ S
Sbjct: 112 SSSSSSPS 119
>UniRef100_UPI0000363B09 UPI0000363B09 UniRef100 entry
Length = 490
Score = 33.1 bits (74), Expect = 2.2
Identities = 23/77 (29%), Positives = 41/77 (52%), Gaps = 1/77 (1%)
Query: 11 SFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSL 70
S+ + +S++ S+S S DSS SS++ T A++S+S +SS+ S S
Sbjct: 253 SYSSSSYSSSSYSTSYSYSSSSDSSYSYSSSYSSSYSYTTVTAASSSSYSYSSY-SSSSY 311
Query: 71 SSMGDYAGTSIKNWWDW 87
SS G Y+ +S + + +
Sbjct: 312 SSRGSYSSSSYSSSYSY 328
>UniRef100_Q9X2V2 Serum opacity factor [Streptococcus pyogenes]
Length = 1029
Score = 33.1 bits (74), Expect = 2.2
Identities = 19/70 (27%), Positives = 35/70 (49%)
Query: 25 PVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAVWSSFPRSKSLSSMGDYAGTSIKNW 84
P + V + SS NT A+A A+AS ++ P + + +S G+ AG+ + +
Sbjct: 25 PTVLGQEVSTGVSNTEASASSTNTNTASADATASGTAATTPSAGTSTSTGEAAGSGLSSE 84
Query: 85 WDWTWAWILS 94
+W+ A + S
Sbjct: 85 ANWSDAAVAS 94
>UniRef100_Q8LTG0 GtrC [Bacteriophage P22-pbi]
Length = 486
Score = 33.1 bits (74), Expect = 2.2
Identities = 13/35 (37%), Positives = 21/35 (59%)
Query: 86 DWTWAWILSRKPIFARDLELNDQETKLLGGSHNRG 120
DW W+ +L ++ +F+R+ L D+E KL G G
Sbjct: 434 DWMWSEVLMQRNVFSRNYRLYDKEVKLENGWKKSG 468
>UniRef100_P57019 Hypothetical 55.3 kDa protein in gtrB 5'region [Bacteriophage P22]
Length = 485
Score = 33.1 bits (74), Expect = 2.2
Identities = 13/35 (37%), Positives = 21/35 (59%)
Query: 86 DWTWAWILSRKPIFARDLELNDQETKLLGGSHNRG 120
DW W+ +L ++ +F+R+ L D+E KL G G
Sbjct: 433 DWMWSEVLMQRNVFSRNYRLYDKEVKLENGWKKSG 467
>UniRef100_UPI000042C858 UPI000042C858 UniRef100 entry
Length = 1409
Score = 32.7 bits (73), Expect = 2.9
Identities = 27/72 (37%), Positives = 38/72 (52%), Gaps = 9/72 (12%)
Query: 2 SSIGHQQHRSFF--TIPTTSNNKQLPVSHSRSVD--SSEHGLRKRLSSFNTTPAAASASA 57
+S GH FF +IP+TS ++Q P S +V+ SS S +T AAA+ S+
Sbjct: 72 TSTGH-----FFGPSIPSTSTHQQTPGETSNNVNTKSSSQNQSPSTSPTSTVAAAAATSS 126
Query: 58 SAVWSSFPRSKS 69
S V S+ P S S
Sbjct: 127 SPVASTRPASTS 138
>UniRef100_Q6RUV4 C3 phosphoenolpyruvate carboxylase [Setaria italica]
Length = 961
Score = 32.7 bits (73), Expect = 2.9
Identities = 14/45 (31%), Positives = 24/45 (53%)
Query: 73 MGDYAGTSIKNWWDWTWAWILSRKPIFARDLELNDQETKLLGGSH 117
+G YA S + DW + + ++P+F DL L ++ +LG H
Sbjct: 466 IGSYAEWSEEKRQDWLLSELSGKRPLFGSDLPLTEETADVLGAFH 510
>UniRef100_Q966P1 Hypothetical protein C46C11.1 [Caenorhabditis elegans]
Length = 943
Score = 32.7 bits (73), Expect = 2.9
Identities = 19/60 (31%), Positives = 32/60 (52%)
Query: 1 MSSIGHQQHRSFFTIPTTSNNKQLPVSHSRSVDSSEHGLRKRLSSFNTTPAAASASASAV 60
+SS+ ++ R T P+T + P+SHS S++ + KR S + AAS +A A+
Sbjct: 737 LSSVTSRESRDTQTAPSTPLQQPKPLSHSSSMNGFNPPVHKRSLSQSLADTAASTAAYAL 796
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.316 0.127 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 270,489,760
Number of Sequences: 2790947
Number of extensions: 9981275
Number of successful extensions: 30979
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 25
Number of HSP's successfully gapped in prelim test: 24
Number of HSP's that attempted gapping in prelim test: 30924
Number of HSP's gapped (non-prelim): 67
length of query: 155
length of database: 848,049,833
effective HSP length: 116
effective length of query: 39
effective length of database: 524,299,981
effective search space: 20447699259
effective search space used: 20447699259
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0380.7


