

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0378b.2
(736 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
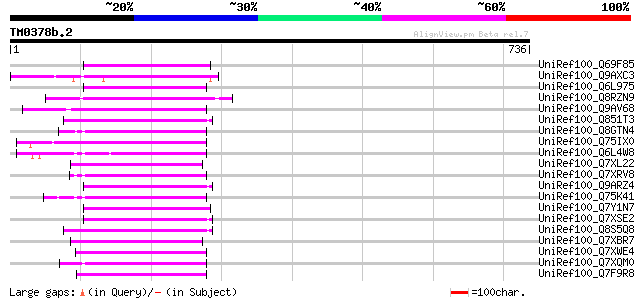
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris] 120 1e-25
UniRef100_Q9AXC3 Hypothetical protein [Antirrhinum hispanicum] 117 1e-24
UniRef100_Q6L975 GAG-POL [Vitis vinifera] 111 9e-23
UniRef100_Q8RZN9 Putative polyprotein [Oryza sativa] 97 2e-18
UniRef100_Q9AV68 Putative gag-pol [Oryza sativa] 97 2e-18
UniRef100_Q851T3 Putative gag-pol [Oryza sativa] 96 4e-18
UniRef100_Q8GTN4 Putative gag-pol [Oryza sativa] 96 5e-18
UniRef100_Q75IX0 Putative retrotransposon gag protein [Oryza sat... 94 1e-17
UniRef100_Q6L4W8 Hypothetical protein OSJNBb0108E17.12 [Oryza sa... 94 2e-17
UniRef100_Q7XL22 OSJNBa0079M09.7 protein [Oryza sativa] 93 3e-17
UniRef100_Q7XRV8 OSJNBb0049I21.2 protein [Oryza sativa] 93 3e-17
UniRef100_Q9ARZ4 Putative polyprotein [Oryza sativa] 92 4e-17
UniRef100_Q75K41 Putative gag-pol protein [Oryza sativa] 92 4e-17
UniRef100_Q7Y1N7 Putative polyprotein [Oryza sativa] 92 4e-17
UniRef100_Q7XSE2 OSJNBb0033G08.12 protein [Oryza sativa] 92 4e-17
UniRef100_Q8S5Q8 Putative retroelement [Oryza sativa] 92 6e-17
UniRef100_Q7XBR7 Putative gypsy-type retrotransposon gag-pol [Or... 92 6e-17
UniRef100_Q7XWE4 OSJNBa0035O13.3 protein [Oryza sativa] 92 6e-17
UniRef100_Q7XQM0 OSJNBa0089E12.4 protein [Oryza sativa] 92 6e-17
UniRef100_Q7F9R8 OSJNBb0054B09.2 protein [Oryza sativa] 92 6e-17
>UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 120 bits (302), Expect = 1e-25
Identities = 54/181 (29%), Positives = 100/181 (54%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 164
PF++D+ + +PD + L ++ Y G +DP +HL F T+M + V C++FP++ +
Sbjct: 211 PFTDDIIATPLPDKWRGLTINLYDGSTDPDEHLNIFRTQMTLYTTDRTVWCKVFPTSLRE 270
Query: 165 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYS 224
+ WF+ LP SI++F KF Q++ ++ + L N++Q+ GESL+ +M R+S
Sbjct: 271 GPLGWFSDLPPNSIASFDALELKFTTQYATSRPHRTSSMSLLNVKQERGESLRTFMNRFS 330
Query: 225 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 284
+ + + P + +LPG L ++P +M+E+R RA+ ++ EE + R
Sbjct: 331 KVCMNIRNLNPEIAMHHLVSAILPGRFTESLIKRPPCNMDELRTRATKFMQIEEHIDYHR 390
Query: 285 K 285
K
Sbjct: 391 K 391
>UniRef100_Q9AXC3 Hypothetical protein [Antirrhinum hispanicum]
Length = 1455
Score = 117 bits (293), Expect = 1e-24
Identities = 89/309 (28%), Positives = 139/309 (44%), Gaps = 18/309 (5%)
Query: 1 RPHRVVLDSPVRTTGVQSPSPPPSPPVPPPPPSPSQVGSLERSPENSPAPEQQPAVTQEQ 60
R R + RT G P PS P + S+ L + + + ++P +
Sbjct: 51 RGARRIRQQEARTRGRWIPPLSPSAPASGGMHARSEPRRLRAEAKQTFSRGERPIGSHRN 110
Query: 61 WRHLMRNIGNIQQRNEHLQAQLDFYRRE----QRD-DGSREADSVAEFRPFSEDVESVAI 115
+ R + N+ + +RE RD DG+ E D + PF+ + + I
Sbjct: 111 GTY--RPFPGMDNNNQQFINMVTELKREVQALNRDRDGTAEEDEMGS--PFAPHILAPEI 166
Query: 116 PDNMKTLVLDSYSGDS---DPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTT 172
P +M+ Y+GD DP+DH+ F M I++ D C +F ST A AWF+
Sbjct: 167 PVSMRLSNPVVYTGDRNARDPRDHINQFLAVMDILSTPDHALCWIFRSTLTGRAQAWFSQ 226
Query: 173 LPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVED 232
LPRGSI +F + F+ QF+ N+ P TI L + + EGESL+EYM R++ + V
Sbjct: 227 LPRGSIRSFDELRELFVRQFACNRRYPKTIGHLMTMVEAEGESLREYMRRFANTVIDVRH 286
Query: 233 EEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFK------RKR 286
P + G + L +PARS+EE+ RA Y+L E+ K R
Sbjct: 287 VGPHILSEFVAQGTRSVEFANSLVGRPARSLEELMHRAERYMLQEDTRLAKLGIQGHEDR 346
Query: 287 AKLEKGDTS 295
++ +GD +
Sbjct: 347 SRNRRGDNT 355
>UniRef100_Q6L975 GAG-POL [Vitis vinifera]
Length = 549
Score = 111 bits (277), Expect = 9e-23
Identities = 56/176 (31%), Positives = 95/176 (53%), Gaps = 1/176 (0%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 164
PF + + P +Y G SDP DH++++ M + +DA+ C++FP++ +
Sbjct: 5 PFCSHIINYEPPRGFLVPKFSTYDGTSDPFDHIMHYRQLMTLDIGNDALLCKVFPASLQG 64
Query: 165 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYS 224
A++WF LP S+ NFRD S F+ Q+ + I+ L NI+ Q+ ESL+E++ R+
Sbjct: 65 QALSWFHRLPPNSVGNFRDLSEAFVGQYLCSARHKQNISTLQNIKMQDNESLREFVKRFG 124
Query: 225 AASVKVEDEEPRACALAFKNGLLPG-GLNSKLTRKPARSMEEMRARASTYILDEED 279
A ++VE A FK + PG L +KP +M+++ RA+ Y + E+D
Sbjct: 125 QAVLQVEACSMDAVLQIFKRSICPGTPFFESLAKKPPTTMDDLFRRANKYSMLEDD 180
>UniRef100_Q8RZN9 Putative polyprotein [Oryza sativa]
Length = 2001
Score = 97.1 bits (240), Expect = 2e-18
Identities = 61/265 (23%), Positives = 120/265 (45%), Gaps = 7/265 (2%)
Query: 52 QQPAVTQEQWRHLMRNIGNIQQRNEHLQAQLDFYRREQRDDGSREADSVAEFRPFSEDVE 111
QQP + EQ ++ + +IQ + F + +R D+ P S++++
Sbjct: 293 QQPEMMFEQG-YIPQMANHIQYAGQIPNQAFQFQPTQPYFQPTRAIDTK---NPLSQNLQ 348
Query: 112 SVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFT 171
P N K + Y GD+DP ++L + T + D K ++ P+ + A++W+T
Sbjct: 349 MAPWPINFKLSNITKYKGDTDPNEYLRVYETAVEAAGGDDTTKAKILPTMLEGVALSWYT 408
Query: 172 TLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVE 231
T+P +I ++ F F +P +DLY ++Q GE+L+ ++ ++S ++
Sbjct: 409 TIPPMTIYSWEHMRDTFRAGFIGAYEEPKEADDLYAMKQLPGETLRSFIVKFSRVRCQIR 468
Query: 232 DEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKRKRAKLEK 291
+ A K LLPG L L R ++ +++ R ++ EED+ ++ +
Sbjct: 469 HVDDEMLIAAAKRALLPGPLRFDLARNRPKTTKDLFERMESFARGEEDELRVQEEEAVLL 528
Query: 292 GDTSPKQR*KEEFQIGGKAEGKEQG 316
G KQ ++ G + +G+ G
Sbjct: 529 G---KKQSKNKQISQGEEQKGENTG 550
>UniRef100_Q9AV68 Putative gag-pol [Oryza sativa]
Length = 1720
Score = 96.7 bits (239), Expect = 2e-18
Identities = 67/260 (25%), Positives = 109/260 (41%), Gaps = 6/260 (2%)
Query: 19 PSPPPSPPVPPPPPSPSQVGSLERSPENSPAPEQQPAVTQEQWRHLMRNIGNIQQRNEHL 78
P+ PS + S G ERS + P ++ V + + R + N+H
Sbjct: 220 PANSPSLSITGTGSSRRSRGHDERSVHSPPERHRERRVERPRSPPHQRPVDLRDMINQHR 279
Query: 79 QAQLDFYRREQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLL 138
A R R +R D V F+ D+ V P K ++ Y+G ++P+ L
Sbjct: 280 AA------RGHRHSPNRYDDDVDRVATFTNDLRRVDWPVGFKPTGIEKYNGTTNPESWLT 333
Query: 139 YFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQ 198
+ + P +A +W LPR +I ++ + F+ F +
Sbjct: 334 VYGIAIRAAGGDSKAMANYLPVALADSARSWLHGLPRRTIGSWAELRDHFIANFQGTFER 393
Query: 199 PVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRK 258
P T DLYNI Q+ GESL++Y+ R+S K+ D AF G+ L K RK
Sbjct: 394 PGTQFDLYNIVQKSGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEDLVGKFGRK 453
Query: 259 PARSMEEMRARASTYILDEE 278
P R++++M +A+ Y E+
Sbjct: 454 PPRTVKQMFEKANEYAKAED 473
>UniRef100_Q851T3 Putative gag-pol [Oryza sativa]
Length = 971
Score = 95.9 bits (237), Expect = 4e-18
Identities = 59/211 (27%), Positives = 97/211 (45%), Gaps = 2/211 (0%)
Query: 77 HLQAQLDFYRREQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDH 136
H QA + + G+ + P S +++ IP K L + YSG++DPK+
Sbjct: 329 HAQAVISPFATPYPQQGTAGRAGGEKGLPLSGGIKNRPIPPQFKFLPVPRYSGETDPKEF 388
Query: 137 LLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANK 196
L + + + + K ++ A +W+ LP SI ++ F++ F
Sbjct: 389 LSIYESAIEAAHGDENTKAKVIHLALDGIARSWYFNLPANSIYSWEQLRDVFVLNFRGTY 448
Query: 197 NQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLT 256
+P T L IRQ+ GES++EYM R+S A +VED + A GLL G L K+
Sbjct: 449 EEPKTQQHLLGIRQRPGESIREYMRRFSQARCQVEDITEASVINAASAGLLEGELTRKIA 508
Query: 257 RKPARSMEEMRARASTYILDEEDDAFKRKRA 287
K +++E + + EED KR++A
Sbjct: 509 NKEPQTLEHLVRIIDGFAKSEEDS--KRRQA 537
>UniRef100_Q8GTN4 Putative gag-pol [Oryza sativa]
Length = 792
Score = 95.5 bits (236), Expect = 5e-18
Identities = 58/209 (27%), Positives = 96/209 (45%), Gaps = 5/209 (2%)
Query: 70 NIQQRNEHLQAQLDFYRREQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSG 129
N +++H +A YR R D + D VA F ++D+ V P K ++ Y G
Sbjct: 294 NELNQSDHRRAARGHYRSPDRHDD--DLDGVAAF---TDDLRRVDWPAGFKPTGIEKYDG 348
Query: 130 DSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFL 189
++P+ L + + + P +A +W LPRG+I ++ + F+
Sbjct: 349 TTNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFI 408
Query: 190 VQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPG 249
F +P T DLYN+ Q+ GESL++Y+ R+S K+ D AF G+
Sbjct: 409 ANFQGTFERPGTQYDLYNVIQKSGESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHE 468
Query: 250 GLNSKLTRKPARSMEEMRARASTYILDEE 278
L K RKP R+++ M +A+ Y E+
Sbjct: 469 ELVGKFGRKPPRTVKLMFEKANEYAKAED 497
>UniRef100_Q75IX0 Putative retrotransposon gag protein [Oryza sativa]
Length = 776
Score = 94.4 bits (233), Expect = 1e-17
Identities = 71/276 (25%), Positives = 115/276 (40%), Gaps = 8/276 (2%)
Query: 10 PVRTTGVQSPSPPPSPPVP-----PPPPSPSQVGSLERSPENSPAPEQQPAVTQEQWRHL 64
PV + V++P+ P + PP S VGS RS + P + R +
Sbjct: 184 PVNSGSVRTPTGSQVPHLRSHDYHPPSLSIVGVGSSRRSRGHDERSIHSPPERHRE-RRV 242
Query: 65 MRNIGNIQQRNEHLQAQLDFYR--REQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTL 122
R +QR L+ ++ R R R R D V F+ D+ V P K
Sbjct: 243 ERPRSPPRQRPVDLRDTINQRRAARGHRHSPDRYDDDVDGVAAFTNDLRQVDWPAGFKPT 302
Query: 123 VLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFR 182
++ Y G ++P+ L + + P +A +W LPRG+I ++
Sbjct: 303 GIEKYDGTTNPESWLTVYGLAIRTAGGDSKAMANYLPVALADSARSWLHGLPRGTIGSWA 362
Query: 183 DFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAF 242
+ F+ F +P T DLYN+ Q+ ESL++Y+ R+S K+ D AF
Sbjct: 363 ELRDHFIANFQGTFERPGTQFDLYNVVQKSEESLRDYIRRFSEQRNKISDITDDVIIAAF 422
Query: 243 KNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEE 278
G+ L K RKP + +++M +A+ Y E+
Sbjct: 423 TKGIRHEDLVGKFGRKPPKMVKQMFEKANEYAKAED 458
>UniRef100_Q6L4W8 Hypothetical protein OSJNBb0108E17.12 [Oryza sativa]
Length = 1315
Score = 93.6 bits (231), Expect = 2e-17
Identities = 77/285 (27%), Positives = 129/285 (45%), Gaps = 24/285 (8%)
Query: 10 PVRTTGVQSPSPPPSPPVPPPP---PSPSQVGSL---------ERSPENSPAPEQQPAVT 57
PV + V++P+ PP+ PS S VG+ E+S + P ++
Sbjct: 153 PVNSASVRTPTGSRVPPLQSQDYHQPSLSVVGAGTSRRSRDHDEQSVHSPPDRHRERRAE 212
Query: 58 QEQWRHLMRNIGNIQQRNEHLQAQ--LDFYRREQRDDGSREADSVAEFRPFSEDVESVAI 115
+ H R I + N+ A+ + + ++ DD + D VA F + D+ V
Sbjct: 213 CPRSPHRRRPIDLRETINQRRAARGYIPHHSPDRYDD---DVDGVAAF---TSDLRRVDW 266
Query: 116 PDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKC--RMFPSTFKSTAMAWFTTL 173
P K ++ Y G ++P+ L ++ + I AA +K P +A +W L
Sbjct: 267 PAGFKPTGIEKYDGTTNPETWLTVYS--LAIRAAGGDIKAMANYLPVALADSARSWLHGL 324
Query: 174 PRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDE 233
PRG+I ++ + F+ F +P T DLYNI Q+ GESL++Y+ R+S K+ D
Sbjct: 325 PRGTIGSWAELRDHFIANFQGTFERPGTQFDLYNIVQKSGESLRDYIRRFSEQRNKISDI 384
Query: 234 EPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEE 278
AF G+ L K RKP R++++M +A+ Y E+
Sbjct: 385 TDDVIIAAFTKGIHHDLLVGKFGRKPPRTVKQMFEKANEYAKAED 429
>UniRef100_Q7XL22 OSJNBa0079M09.7 protein [Oryza sativa]
Length = 877
Score = 92.8 bits (229), Expect = 3e-17
Identities = 51/187 (27%), Positives = 84/187 (44%)
Query: 87 REQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVI 146
R R R D V F+ D+ V P K ++ Y G ++P+ L + +
Sbjct: 314 RGHRHSPDRYDDDVDRVAAFTNDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRA 373
Query: 147 IAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLY 206
P +A +W LPRG+I ++ + F+ F +P T DLY
Sbjct: 374 AGGDSKAMANYLPVALADSAWSWLHGLPRGTIGSWAELRDHFITNFQGTFERPGTQFDLY 433
Query: 207 NIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEM 266
N+ Q+ GESL++Y+ R+S K+ D +AF G+ L K RKP +++++M
Sbjct: 434 NVIQKSGESLRDYIRRFSEQRNKISDITDDVIIVAFTKGIRHEDLVGKFGRKPPKTVKQM 493
Query: 267 RARASTY 273
+A+ Y
Sbjct: 494 FEKANEY 500
>UniRef100_Q7XRV8 OSJNBb0049I21.2 protein [Oryza sativa]
Length = 1843
Score = 92.8 bits (229), Expect = 3e-17
Identities = 55/194 (28%), Positives = 90/194 (46%), Gaps = 6/194 (3%)
Query: 85 YRREQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKM 144
Y ++ DD + D VA F ++D+ V P K ++ Y G ++P+ L + +
Sbjct: 412 YSPDRHDD---DLDGVAAF---TDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAI 465
Query: 145 VIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTIND 204
P +A +W LPRG+I ++ + +F+ F +P T D
Sbjct: 466 RAAGGDSKAMANYLPVALADSARSWLHGLPRGTIGSWAELRDRFIANFQGTFERPGTQYD 525
Query: 205 LYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSME 264
LYN+ Q+ GESL+EY+ R+S K+ D AF G+ L K RKP R+++
Sbjct: 526 LYNVIQKSGESLREYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVK 585
Query: 265 EMRARASTYILDEE 278
M +A+ Y E+
Sbjct: 586 LMFEKANEYAKAED 599
>UniRef100_Q9ARZ4 Putative polyprotein [Oryza sativa]
Length = 1579
Score = 92.4 bits (228), Expect = 4e-17
Identities = 53/183 (28%), Positives = 90/183 (48%), Gaps = 2/183 (1%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 164
P S +++ IP K + YSG++DPK+ L + + + + K ++ T
Sbjct: 49 PLSGGIKTRPIPPQFKFPPVPCYSGETDPKEFLSIYESAIEAAHGDENTKAKVIHLTLDG 108
Query: 165 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYS 224
A +W+ LP SI ++ F++ F +P T L IRQ+ GES++EYM R+S
Sbjct: 109 IARSWYFNLPANSIYSWEQLRDVFVLNFRGTYEEPKTQQHLLGIRQRPGESIREYMRRFS 168
Query: 225 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 284
A +V+D + A GLL G L K+ K ++++++ + EED KR
Sbjct: 169 QARCQVQDITEASVINAASAGLLEGELTRKIANKEPQTLKQLLRIIDGFARGEEDS--KR 226
Query: 285 KRA 287
++A
Sbjct: 227 RQA 229
>UniRef100_Q75K41 Putative gag-pol protein [Oryza sativa]
Length = 1653
Score = 92.4 bits (228), Expect = 4e-17
Identities = 59/231 (25%), Positives = 106/231 (45%), Gaps = 7/231 (3%)
Query: 48 PAPEQQPAVTQEQWRHLMRNIGNIQQRNEHLQAQLDFYRREQRDDGSREADSVAEFRPFS 107
P P+ + V + H R + +++ L+A + ++ DD + D VA F +
Sbjct: 285 PHPDWERRVERPHSPHRRRPV-DLRDTINQLRAARGNHSPDRYDD---DMDGVAAF---T 337
Query: 108 EDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAM 167
++ V P + K ++ Y G ++P+ L + + + P +A
Sbjct: 338 SELRRVDWPADFKPTGIEKYDGTTNPESWLTVYGLAVRAVGGDSKAMANYLPVALADSAR 397
Query: 168 AWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAAS 227
+W LPRG+I ++ + F+ F +P T DLYNI Q+ GESL++Y+ R+S
Sbjct: 398 SWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTHFDLYNIVQKSGESLRDYIRRFSKQR 457
Query: 228 VKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEE 278
K+ D AF G+ L K RKP +++++M +A+ Y E+
Sbjct: 458 NKISDITDDVIIAAFTKGIRHEDLVGKFGRKPPKTVKQMFEKANEYAKAED 508
>UniRef100_Q7Y1N7 Putative polyprotein [Oryza sativa]
Length = 518
Score = 92.4 bits (228), Expect = 4e-17
Identities = 53/181 (29%), Positives = 86/181 (47%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 164
P S +++ IP K + YSG++D K+ L + + + + K ++
Sbjct: 282 PLSWGIKNHPIPPQFKYPPVPQYSGETDSKEFLSIYESAIEAAHGDENTKAKLIHLVLDG 341
Query: 165 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYS 224
A +W+ LP SI ++ F++ F +P T DL I Q++GES +EY+ R+
Sbjct: 342 IARSWYFNLPVNSIYSWEQLRDVFVLNFRGTYEEPKTQQDLVGIHQRQGESTREYIRRFL 401
Query: 225 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 284
A +V+D + A GLL G L KL RK R++E + Y DEED +R
Sbjct: 402 QARCQVQDITEASVIHAASAGLLEGELTRKLVRKEPRTLEHLFRIIDGYARDEEDTKRRR 461
Query: 285 K 285
+
Sbjct: 462 E 462
>UniRef100_Q7XSE2 OSJNBb0033G08.12 protein [Oryza sativa]
Length = 1847
Score = 92.4 bits (228), Expect = 4e-17
Identities = 53/183 (28%), Positives = 89/183 (47%), Gaps = 2/183 (1%)
Query: 105 PFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKS 164
P S +++ IP K + YSG++DPK+ L + + + + K ++
Sbjct: 417 PLSGGIKTRPIPPQFKFPPVPRYSGETDPKEFLSIYESAIEAAHGDENTKAKVIHLALDG 476
Query: 165 TAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGESLKEYMARYS 224
A +W+ LP SI ++ F++ F +P T L IRQ++GES++EYM R+S
Sbjct: 477 IARSWYFNLPANSIYSWEQLRDVFVLNFRGTYKEPKTQQHLLGIRQKQGESIREYMRRFS 536
Query: 225 AASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYILDEEDDAFKR 284
A +V+D + A GLL G L K+ K +++E + + EED KR
Sbjct: 537 QARCQVQDITKASVINAASAGLLEGELTRKIANKEPQTLEHLLRIIDGFARGEEDS--KR 594
Query: 285 KRA 287
++A
Sbjct: 595 RQA 597
>UniRef100_Q8S5Q8 Putative retroelement [Oryza sativa]
Length = 1292
Score = 92.0 bits (227), Expect = 6e-17
Identities = 57/211 (27%), Positives = 96/211 (45%), Gaps = 2/211 (0%)
Query: 77 HLQAQLDFYRREQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDH 136
H+QA + + G+ + P S +++ IP K + YSG++DPK+
Sbjct: 21 HVQAAISPFATPYPQQGAVNRAGGEKGLPLSGGIKTRPIPPQFKFPPVPRYSGETDPKEF 80
Query: 137 LLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANK 196
L + + + + K ++ A +W+ LP SI ++ F++ F
Sbjct: 81 LSIYESAIEAAHGDENTKAKVIHLALDGIARSWYFNLPDNSIYSWEQLRDVFVLNFRGTY 140
Query: 197 NQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLT 256
+P T L IRQ+ GES++EYM R+S A +V+D + A GLL G L K+
Sbjct: 141 EEPKTQQHLLGIRQRPGESIREYMRRFSQARCQVQDITEASVINAASAGLLEGELTRKIA 200
Query: 257 RKPARSMEEMRARASTYILDEEDDAFKRKRA 287
K ++E + + EED KR++A
Sbjct: 201 NKEPLTLEHLLCIIDGFARGEEDS--KRRQA 229
>UniRef100_Q7XBR7 Putative gypsy-type retrotransposon gag-pol [Oryza sativa]
Length = 785
Score = 92.0 bits (227), Expect = 6e-17
Identities = 52/187 (27%), Positives = 84/187 (44%)
Query: 87 REQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVI 146
R R +R D V F+ D+ V P K ++ Y G ++P+ L + +
Sbjct: 255 RGHRHSPNRYDDDVDGVAAFTNDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRA 314
Query: 147 IAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLY 206
P ++A +W LPRG+I ++ + F+ F +P T DLY
Sbjct: 315 AGGDSKAMANYLPVALANSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQFDLY 374
Query: 207 NIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEM 266
N+ Q+ GESL++Y+ R+S K+ D AF G+ L K RKP R++++M
Sbjct: 375 NVVQKSGESLRDYIRRFSEQRNKISDITDDIIIAAFTKGIRHEDLVGKFGRKPPRTVKQM 434
Query: 267 RARASTY 273
+A Y
Sbjct: 435 FEKAHEY 441
>UniRef100_Q7XWE4 OSJNBa0035O13.3 protein [Oryza sativa]
Length = 2008
Score = 92.0 bits (227), Expect = 6e-17
Identities = 51/185 (27%), Positives = 85/185 (45%)
Query: 94 SREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAV 153
+R D + E F++D+ V P K ++ Y G ++P L + + +
Sbjct: 409 NRHDDDLDEVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPGSWLTVYGLAIRAAGGDNKA 468
Query: 154 KCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEG 213
P +A +W LPRG+I ++ + F+ F +P T DLYN+ Q+ G
Sbjct: 469 MANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSG 528
Query: 214 ESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTY 273
ESL++Y+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 529 ESLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEY 588
Query: 274 ILDEE 278
E+
Sbjct: 589 AKAED 593
>UniRef100_Q7XQM0 OSJNBa0089E12.4 protein [Oryza sativa]
Length = 1965
Score = 92.0 bits (227), Expect = 6e-17
Identities = 57/208 (27%), Positives = 94/208 (44%), Gaps = 3/208 (1%)
Query: 71 IQQRNEHLQAQLDFYRREQRDDGSREADSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGD 130
I R+ +Q + R D + D VA F ++D+ V P + K ++ Y G
Sbjct: 394 IDLRDTIIQRRAARGRHHSPDRYDDDLDGVAAF---TDDLRRVDWPASFKPTGIEKYDGT 450
Query: 131 SDPKDHLLYFNTKMVIIAASDAVKCRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLV 190
++P+ L + + + P +A +W LPRG+I ++ + F+
Sbjct: 451 TNPESWLTVYGLAIRAAGGDNKAMANYLPVALADSARSWLHGLPRGTIGSWAELRDHFIA 510
Query: 191 QFSANKNQPVTINDLYNIRQQEGESLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGG 250
F +P T DLYN+ Q+ GESL+EY+ R+S K+ D AF G+
Sbjct: 511 NFQGTFERPGTQYDLYNVIQKSGESLREYIRRFSEQRNKISDITDDVIIAAFTKGIRHEE 570
Query: 251 LNSKLTRKPARSMEEMRARASTYILDEE 278
L K RKP R+++ M +A+ Y E+
Sbjct: 571 LVGKFGRKPPRTVKLMFEKANEYAKAED 598
>UniRef100_Q7F9R8 OSJNBb0054B09.2 protein [Oryza sativa]
Length = 1992
Score = 92.0 bits (227), Expect = 6e-17
Identities = 50/184 (27%), Positives = 85/184 (46%)
Query: 95 READSVAEFRPFSEDVESVAIPDNMKTLVLDSYSGDSDPKDHLLYFNTKMVIIAASDAVK 154
R D + F++D+ V P K ++ Y G ++P+ L + + +
Sbjct: 409 RHDDDLDGVAAFTDDLRRVDWPAGFKPTGIEKYDGTTNPESWLTVYGLAIRAVGGDSKAM 468
Query: 155 CRMFPSTFKSTAMAWFTTLPRGSISNFRDFSSKFLVQFSANKNQPVTINDLYNIRQQEGE 214
P + +A +W LPRG+I ++ + F+ F +P T DLYN+ Q+ GE
Sbjct: 469 ANYLPVALEDSARSWLHGLPRGTIGSWAELRDHFIANFQGTFERPGTQYDLYNVIQKSGE 528
Query: 215 SLKEYMARYSAASVKVEDEEPRACALAFKNGLLPGGLNSKLTRKPARSMEEMRARASTYI 274
SL++Y+ R+S K+ D AF G+ L K RKP R+++ M +A+ Y
Sbjct: 529 SLRDYIRRFSEQRNKISDITDDVIIAAFTKGIRHEELVGKFGRKPPRTVKLMFEKANEYA 588
Query: 275 LDEE 278
E+
Sbjct: 589 KAED 592
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.336 0.147 0.491
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,185,342,634
Number of Sequences: 2790947
Number of extensions: 49605354
Number of successful extensions: 694707
Number of sequences better than 10.0: 3088
Number of HSP's better than 10.0 without gapping: 1726
Number of HSP's successfully gapped in prelim test: 1537
Number of HSP's that attempted gapping in prelim test: 595238
Number of HSP's gapped (non-prelim): 34990
length of query: 736
length of database: 848,049,833
effective HSP length: 135
effective length of query: 601
effective length of database: 471,271,988
effective search space: 283234464788
effective search space used: 283234464788
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 79 (35.0 bits)
Lotus: description of TM0378b.2


