

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0372b.7
(224 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
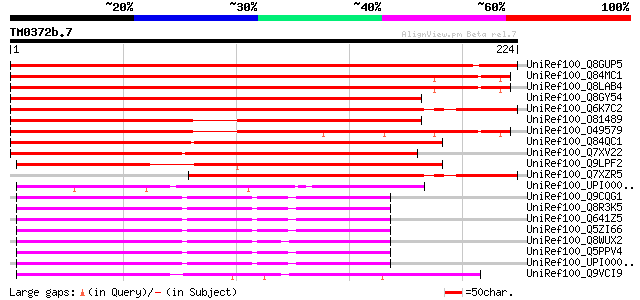
UniRef100_Q8GUP5 Putative cation transporter [Beta procumbens] 343 2e-93
UniRef100_Q84MC1 At4g31290 [Arabidopsis thaliana] 329 3e-89
UniRef100_Q8LAB4 Hypothetical protein [Arabidopsis thaliana] 326 3e-88
UniRef100_Q8GY54 Hypothetical protein At5g26220/T19G15_70 [Arabi... 322 6e-87
UniRef100_Q6K7C2 Putative OsCTTP [Oryza sativa] 300 1e-80
UniRef100_O81489 F9D12.14 protein [Arabidopsis thaliana] 276 4e-73
UniRef100_O49579 Predicted protein [Arabidopsis thaliana] 258 8e-68
UniRef100_Q84QC1 Hypothetical protein At1g44790 [Arabidopsis tha... 189 6e-47
UniRef100_Q7XV22 OSJNBa0064H22.12 protein [Oryza sativa] 188 8e-47
UniRef100_Q9LPF2 T12C22.6 protein [Arabidopsis thaliana] 171 1e-41
UniRef100_Q7XZR5 OsCTTP protein [Oryza sativa] 168 1e-40
UniRef100_UPI000029F00B UPI000029F00B UniRef100 entry 130 3e-29
UniRef100_Q9CQG1 Mus musculus 13 days embryo liver cDNA, RIKEN f... 122 7e-27
UniRef100_Q8R3K5 RIKEN cDNA 2510006C20 [Mus musculus] 122 7e-27
UniRef100_Q641Z5 Hypothetical protein [Rattus norvegicus] 122 7e-27
UniRef100_Q5ZI66 Hypothetical protein [Gallus gallus] 120 2e-26
UniRef100_Q8WUX2 Hypothetical protein [Homo sapiens] 120 3e-26
UniRef100_Q5PPV4 Hypothetical protein [Xenopus laevis] 120 3e-26
UniRef100_UPI00003ACB07 UPI00003ACB07 UniRef100 entry 119 4e-26
UniRef100_Q9VCI9 CG10365-PA, isoform A [Drosophila melanogaster] 117 2e-25
>UniRef100_Q8GUP5 Putative cation transporter [Beta procumbens]
Length = 222
Score = 343 bits (880), Expect = 2e-93
Identities = 162/224 (72%), Positives = 180/224 (80%), Gaps = 2/224 (0%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MVFW+FGYGSLVWNPGF+YDEKV+GYIKDY+RVFDLACIDHRGTPEHPARTCTLE EG
Sbjct: 1 MVFWIFGYGSLVWNPGFQYDEKVLGYIKDYKRVFDLACIDHRGTPEHPARTCTLEYSEGS 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
+CWGA YCVRGGPE E MQYLERRECEYDHK VDFYKEG+ P +TGVIVF STPD
Sbjct: 61 LCWGAAYCVRGGPEIESAAMQYLERRECEYDHKITVDFYKEGEDLEPLVTGVIVFMSTPD 120
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
++NKYYLGPAPLD MA QIATA GPCGNNR+YLFLLEKA+H+I HEDD VIELA EVRK
Sbjct: 121 TISNKYYLGPAPLDKMAWQIATAFGPCGNNREYLFLLEKALHDINHEDDYVIELADEVRK 180
Query: 181 VLGVGNVLPNDKKLSGPVQIQHSPHAPIPALQLLPMPEPIALDS 224
VL + +KK+ + H IP+LQ LP E +ALDS
Sbjct: 181 VLDIVKGATKEKKVITTPHVTVKDH--IPSLQHLPRQEAVALDS 222
>UniRef100_Q84MC1 At4g31290 [Arabidopsis thaliana]
Length = 227
Score = 329 bits (844), Expect = 3e-89
Identities = 162/225 (72%), Positives = 183/225 (81%), Gaps = 5/225 (2%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSLVWNPGF YDEKV+G+IK Y+RVFDLACIDHRGTPEHPARTCTLE+ E
Sbjct: 1 MVMWVFGYGSLVWNPGFHYDEKVLGFIKGYKRVFDLACIDHRGTPEHPARTCTLEKAEEA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWG +CVRGGPEKE+L M+YLERRECEYD KT VDFYKE D PA+TGVIVFTSTPD
Sbjct: 61 ICWGTAFCVRGGPEKERLAMEYLERRECEYDLKTSVDFYKEDDPLKPAVTGVIVFTSTPD 120
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KV+NKYYLGPAPL+ MARQIATA+GPCGNNRDYLFLLEKAMH+IGHE+D VIELA+EVRK
Sbjct: 121 KVSNKYYLGPAPLEDMARQIATANGPCGNNRDYLFLLEKAMHDIGHEEDYVIELANEVRK 180
Query: 181 VLGVGN---VLPNDKKLSGPVQIQHSPHAPIPALQLLP-MPEPIA 221
VL + V P + + V + + P A Q+LP PE +A
Sbjct: 181 VLAESSTKKVTPVKESRASRVANKSKNNVP-TAHQILPHHPEAVA 224
>UniRef100_Q8LAB4 Hypothetical protein [Arabidopsis thaliana]
Length = 227
Score = 326 bits (835), Expect = 3e-88
Identities = 161/225 (71%), Positives = 182/225 (80%), Gaps = 5/225 (2%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSLVWNPGF YDEKV+G+IK Y+RVFDLACIDHRGTPEHPARTCTLE+ E
Sbjct: 1 MVMWVFGYGSLVWNPGFHYDEKVLGFIKGYKRVFDLACIDHRGTPEHPARTCTLEKAEEA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWG +CVRGGPEKE+L M+YLERRECEYD KT VDFYKE PA+TGVIVFTSTPD
Sbjct: 61 ICWGTAFCVRGGPEKERLAMEYLERRECEYDLKTSVDFYKEDYPLKPAVTGVIVFTSTPD 120
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KV+NKYYLGPAPL+ MARQIATA+GPCGNNRDYLFLLEKAMH+IGHE+D VIELA+EVRK
Sbjct: 121 KVSNKYYLGPAPLEDMARQIATANGPCGNNRDYLFLLEKAMHDIGHEEDYVIELANEVRK 180
Query: 181 VLGVGN---VLPNDKKLSGPVQIQHSPHAPIPALQLLP-MPEPIA 221
VL + V P + + V + + P A Q+LP PE +A
Sbjct: 181 VLAESSTKKVTPVKESRASRVANKSKNNVP-TAHQILPHHPEAVA 224
>UniRef100_Q8GY54 Hypothetical protein At5g26220/T19G15_70 [Arabidopsis thaliana]
Length = 216
Score = 322 bits (824), Expect = 6e-87
Identities = 147/182 (80%), Positives = 164/182 (89%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSL+WNPGF++DEK+IGYIKDY+RVFDLACIDHRGTPEHPARTCTLE+ G
Sbjct: 1 MVLWVFGYGSLIWNPGFDFDEKLIGYIKDYKRVFDLACIDHRGTPEHPARTCTLEQSTGA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWGA YCVRGGPEKEKL M+YLERRECEYD KTLV+FY E D+S P +TGVIVFTSTPD
Sbjct: 61 ICWGAAYCVRGGPEKEKLAMEYLERRECEYDSKTLVEFYTENDTSTPIVTGVIVFTSTPD 120
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KV+NKYYLGPAPL+ MARQIATA GPCGNNR+YLF LEKAM +I HE++ VIELA+EVRK
Sbjct: 121 KVSNKYYLGPAPLEEMARQIATASGPCGNNREYLFKLEKAMFDIEHEEEYVIELANEVRK 180
Query: 181 VL 182
L
Sbjct: 181 QL 182
>UniRef100_Q6K7C2 Putative OsCTTP [Oryza sativa]
Length = 216
Score = 300 bits (769), Expect = 1e-80
Identities = 144/224 (64%), Positives = 170/224 (75%), Gaps = 9/224 (4%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSL+WNPGF++DEK++G++K Y+R F+LACIDHRGTPEHPARTCTLE E
Sbjct: 1 MVLWVFGYGSLIWNPGFDFDEKILGFVKGYKRTFNLACIDHRGTPEHPARTCTLESDEEA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWG YCV+GG +KE+ M+YLERRECEYD K VDFYKEGDS PA+TGV+VF STPD
Sbjct: 61 ICWGIAYCVKGGLKKEQEAMKYLERRECEYDQKISVDFYKEGDSLKPAVTGVLVFVSTPD 120
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
V NKYYLGPAPL+ MARQIATA+GP GNNRDYLF +EKA+ NI HEDD +IELA+EVRK
Sbjct: 121 PVGNKYYLGPAPLEDMARQIATANGPNGNNRDYLFSMEKALSNICHEDDSIIELANEVRK 180
Query: 181 VLGVGNVLPNDKKLSGPVQIQHSPHAPIPALQLLPMPEPIALDS 224
VL P +K + SP + L +PE +DS
Sbjct: 181 VLS----RPKEK-----ITGSDSPQKSHALVHLSALPEGTVVDS 215
>UniRef100_O81489 F9D12.14 protein [Arabidopsis thaliana]
Length = 197
Score = 276 bits (705), Expect = 4e-73
Identities = 131/182 (71%), Positives = 147/182 (79%), Gaps = 19/182 (10%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSL+WNPGF++DEK+IGYIKDY+RVFDLACIDHRGTPEHPARTCTLE+ G
Sbjct: 1 MVLWVFGYGSLIWNPGFDFDEKLIGYIKDYKRVFDLACIDHRGTPEHPARTCTLEQSTGA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWGA YCVRGGPEKEKL M+ E D+S P +TGVIVFTSTPD
Sbjct: 61 ICWGAAYCVRGGPEKEKLAME-------------------ENDTSTPIVTGVIVFTSTPD 101
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
KV+NKYYLGPAPL+ MARQIATA GPCGNNR+YLF LEKAM +I HE++ VIELA+EVRK
Sbjct: 102 KVSNKYYLGPAPLEEMARQIATASGPCGNNREYLFKLEKAMFDIEHEEEYVIELANEVRK 161
Query: 181 VL 182
L
Sbjct: 162 QL 163
>UniRef100_O49579 Predicted protein [Arabidopsis thaliana]
Length = 250
Score = 258 bits (659), Expect = 8e-68
Identities = 145/267 (54%), Positives = 166/267 (61%), Gaps = 66/267 (24%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
MV WVFGYGSLVWNPGF YDEKV+G+IK Y+RVFDLACIDHRGTPEHPARTCTLE+ E
Sbjct: 1 MVMWVFGYGSLVWNPGFHYDEKVLGFIKGYKRVFDLACIDHRGTPEHPARTCTLEKAEEA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
ICWG +CVRGGPEKE+L M+ E D PA+TGVIVFTSTPD
Sbjct: 61 ICWGTAFCVRGGPEKERLAME-------------------EDDPLKPAVTGVIVFTSTPD 101
Query: 121 KVNNKYYLGPAPLDVMA--------------RQIATAHGPCGNNRDYLFLLEKAMHNI-- 164
KV+NKYYLGPAPL+ MA RQIATA+GPCGNNRDYLFLLEKAMH+I
Sbjct: 102 KVSNKYYLGPAPLEDMARFLSSHLAVILYIQRQIATANGPCGNNRDYLFLLEKAMHDIGF 161
Query: 165 --------------------------GHEDDMVIELAHEVRKVLGVGN---VLPNDKKLS 195
GHE+D VIELA+EVRKVL + V P + +
Sbjct: 162 VVKGSDLDGFKRDIVFDVLSHFLFHTGHEEDYVIELANEVRKVLAESSTKKVTPVKESRA 221
Query: 196 GPVQIQHSPHAPIPALQLLP-MPEPIA 221
V + + P A Q+LP PE +A
Sbjct: 222 SRVANKSKNNVP-TAHQILPHHPEAVA 247
>UniRef100_Q84QC1 Hypothetical protein At1g44790 [Arabidopsis thaliana]
Length = 199
Score = 189 bits (479), Expect = 6e-47
Identities = 91/191 (47%), Positives = 125/191 (64%), Gaps = 1/191 (0%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
M WVFGYGSL+W GF +DE + G+IK YRRVF DHRGTP+ P RT TLE E
Sbjct: 1 MAMWVFGYGSLIWKTGFPFDESLPGFIKGYRRVFHQGSTDHRGTPDFPGRTVTLEAAHEE 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
+C G Y + +K ++ +LE RE +YD K +DF+ + ++S PA+ GV+V+ ++PD
Sbjct: 61 VCCGVAYKITKEEDKRDALL-HLEVREKQYDQKEYLDFFTDSNASEPAVAGVMVYIASPD 119
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
K +N YLGPAPL+ +A+QI A GP G NRDYLF LE+A+ +G +D V +LA++VR
Sbjct: 120 KKSNNNYLGPAPLEDIAKQIVKAKGPSGPNRDYLFNLEEALAQLGFKDKHVTDLANQVRH 179
Query: 181 VLGVGNVLPND 191
+L L D
Sbjct: 180 ILSESEELDID 190
>UniRef100_Q7XV22 OSJNBa0064H22.12 protein [Oryza sativa]
Length = 187
Score = 188 bits (478), Expect = 8e-47
Identities = 89/180 (49%), Positives = 121/180 (66%), Gaps = 1/180 (0%)
Query: 1 MVFWVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGE 60
M WVFGYGSLVWNPGF +D +++G+++DYRRVF DHRGTPE P RT TLE + G
Sbjct: 1 MAMWVFGYGSLVWNPGFAHDARLVGFVRDYRRVFYQGSTDHRGTPEFPGRTVTLEHQPGA 60
Query: 61 ICWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPD 120
CWG Y + +K+ +++LE RE +YD K +D Y + PA+ V+V+ +T +
Sbjct: 61 TCWGVAYKISTEQDKQ-TALEHLEVREKQYDEKIYLDLYMDSSPKTPAVKNVMVYLATTN 119
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
K +N+ YLGPAPL+ MA+QI A GP G N++YLF LE A++ IG D V +LA+ VRK
Sbjct: 120 KQSNQNYLGPAPLEEMAKQIYLAEGPSGPNKEYLFKLEDALNKIGVVDPHVQDLANAVRK 179
>UniRef100_Q9LPF2 T12C22.6 protein [Arabidopsis thaliana]
Length = 180
Score = 171 bits (433), Expect = 1e-41
Identities = 87/189 (46%), Positives = 118/189 (62%), Gaps = 20/189 (10%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W GF +DE + G+IK YRRVF DHRGTP+ P RT TLE E+C
Sbjct: 2 WVFGYGSLIWKTGFPFDESLPGFIKGYRRVFHQGSTDHRGTPDFPGRTVTLEAAHEEVC- 60
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFY-KEGDSSHPALTGVIVFTSTPDKV 122
+LE RE +YD K +DF+ ++ ++S PA+ GV+V+ ++PDK
Sbjct: 61 ------------------HLEVREKQYDQKEYLDFFTQDSNASEPAVAGVMVYIASPDKK 102
Query: 123 NNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRKVL 182
+N YLGPAPL+ +A+QI A GP G NRDYLF LE+A+ +G +D V +LA++VR +L
Sbjct: 103 SNNNYLGPAPLEDIAKQIVKAKGPSGPNRDYLFNLEEALAQLGFKDKHVTDLANQVRHIL 162
Query: 183 GVGNVLPND 191
L D
Sbjct: 163 SESEELDID 171
>UniRef100_Q7XZR5 OsCTTP protein [Oryza sativa]
Length = 137
Score = 168 bits (425), Expect = 1e-40
Identities = 89/145 (61%), Positives = 103/145 (70%), Gaps = 9/145 (6%)
Query: 80 MQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVNNKYYLGPAPLDVMARQ 139
M+YLERRECEYD K VDFYKEGDS PA+TGV+VF STPD V NKYYLGPAPL+ MARQ
Sbjct: 1 MKYLERRECEYDQKISVDFYKEGDSLKPAVTGVLVFVSTPDPVGNKYYLGPAPLEDMARQ 60
Query: 140 IATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRKVLGVGNVLPNDKKLSGPVQ 199
IATA+GP GNNRDYLF +EKA+ NI HEDD +IELA+EVRKVL P +K +
Sbjct: 61 IATANGPNGNNRDYLFSMEKALSNICHEDDSIIELANEVRKVLS----RPKEK-----IT 111
Query: 200 IQHSPHAPIPALQLLPMPEPIALDS 224
SP + L +PE +DS
Sbjct: 112 GSDSPQKSHALVHLSALPEGTVVDS 136
>UniRef100_UPI000029F00B UPI000029F00B UniRef100 entry
Length = 220
Score = 130 bits (327), Expect = 3e-29
Identities = 77/183 (42%), Positives = 105/183 (57%), Gaps = 8/183 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYI-KDYRRVFDLACIDHRGTPEHPARTCTLEEKEG-EI 61
WVFGYGS+VW GFE+D+ V + +RR F DHRGTPE P RT TLE + E+
Sbjct: 42 WVFGYGSIVWRVGFEHDDVVAPVCARGFRRRFFQGSTDHRGTPEFPGRTATLERCDASEV 101
Query: 62 CWGAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDS-SHPALTGVIVFTSTPD 120
CWGA Y V E+ ++YLE RE +YD + +D Y D+ + P + + + +TP
Sbjct: 102 CWGAAYRVPA--ERRAETLEYLEIREKQYDERVEMDLYAADDAGAAPVIENAVTYIATPA 159
Query: 121 KVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHEDDMVIELAHEVRK 180
+N + LG A D +A QIA A GP G N DYL+ L +A+ + EDD + L VR
Sbjct: 160 DINLNW-LGDA--DDVAEQIARAVGPSGANSDYLYNLCEALRALDIEDDDLYALEASVRA 216
Query: 181 VLG 183
+ G
Sbjct: 217 IRG 219
>UniRef100_Q9CQG1 Mus musculus 13 days embryo liver cDNA, RIKEN full-length enriched
library, clone:2510006C20 product:hypothetical protein,
full insert sequence [Mus musculus]
Length = 178
Score = 122 bits (306), Expect = 7e-27
Identities = 70/165 (42%), Positives = 90/165 (54%), Gaps = 7/165 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W F Y +K++GYI +Y R F DHRG P P R TL E G W
Sbjct: 2 WVFGYGSLIWKVDFPYQDKLVGYITNYSRRFWQGSTDHRGVPGKPGRVVTLVEDPGGSVW 61
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + G E+E V YL+ RE T V FY + ++ P V+++ T D N
Sbjct: 62 GVAYKLPVGKEEE--VKTYLDFREKGGYRTTTVIFYPKDSTTKP--FSVLLYIGTCDNPN 117
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHED 168
YLGPAPL+ +A QI A GP G N +YLF L ++ + ED
Sbjct: 118 ---YLGPAPLEDIAEQIFNAAGPSGRNTEYLFELADSVRKLVPED 159
>UniRef100_Q8R3K5 RIKEN cDNA 2510006C20 [Mus musculus]
Length = 178
Score = 122 bits (306), Expect = 7e-27
Identities = 70/165 (42%), Positives = 90/165 (54%), Gaps = 7/165 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W F Y +K++GYI +Y R F DHRG P P R TL E G W
Sbjct: 2 WVFGYGSLIWKVDFPYQDKLVGYITNYSRRFWQGSTDHRGVPGKPGRVVTLVEDPGGSVW 61
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + G E+E V YL+ RE T V FY + ++ P V+++ T D N
Sbjct: 62 GVAYKLPVGKEEE--VKTYLDFREKGGYRTTTVIFYPKDSTTKP--FSVLLYIGTYDNPN 117
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHED 168
YLGPAPL+ +A QI A GP G N +YLF L ++ + ED
Sbjct: 118 ---YLGPAPLEDIAEQIFNAAGPSGRNTEYLFELADSVRKLVPED 159
>UniRef100_Q641Z5 Hypothetical protein [Rattus norvegicus]
Length = 178
Score = 122 bits (306), Expect = 7e-27
Identities = 70/165 (42%), Positives = 90/165 (54%), Gaps = 7/165 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W F Y +K++GYI +Y R F DHRG P P R TL E G W
Sbjct: 2 WVFGYGSLIWKVDFPYQDKLVGYITNYSRRFWQGSTDHRGVPGKPGRVVTLVEDPGGSVW 61
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + G E+E V YL+ RE T V FY + ++ P V+++ T D N
Sbjct: 62 GVAYKLPVGKEEE--VKTYLDFREKGGYRTTTVIFYPKDSTTKP--FSVLLYIGTCDNPN 117
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHED 168
YLGPAPL+ +A QI A GP G N +YLF L ++ + ED
Sbjct: 118 ---YLGPAPLEDIAEQIFNAAGPSGRNTEYLFELADSIRKLVPED 159
>UniRef100_Q5ZI66 Hypothetical protein [Gallus gallus]
Length = 186
Score = 120 bits (302), Expect = 2e-26
Identities = 72/165 (43%), Positives = 86/165 (51%), Gaps = 7/165 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W F Y EK++G I+ Y R F DHRG P P R TL E W
Sbjct: 2 WVFGYGSLIWKVDFPYQEKMVGRIRGYSRRFWQGSTDHRGVPGKPGRVVTLVEDPEGCVW 61
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + G E E V YL+ RE T V FY + S P V+++ T D N
Sbjct: 62 GVAYRLPAGQECE--VKAYLDFREKGGYRTTTVVFYPKDSSIKP--FDVLLYIGTRDNPN 117
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHED 168
YLGPAPL +A QI A GP G N +YLF L +M N+ ED
Sbjct: 118 ---YLGPAPLQEIAEQIIDAVGPSGRNTEYLFELANSMRNLVPED 159
>UniRef100_Q8WUX2 Hypothetical protein [Homo sapiens]
Length = 184
Score = 120 bits (301), Expect = 3e-26
Identities = 69/165 (41%), Positives = 90/165 (53%), Gaps = 7/165 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W F Y +K++GYI +Y R F DHRG P P R TL E W
Sbjct: 2 WVFGYGSLIWKVDFPYQDKLVGYITNYSRRFWQGSTDHRGVPGKPGRVVTLVEDPAGCVW 61
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + G E+E V YL+ RE T V FY + ++ P V+++ T D
Sbjct: 62 GVAYRLPVGKEEE--VKAYLDFREKGGYRTTTVIFYPKDPTTKP--FSVLLYIGTCD--- 114
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHED 168
N YLGPAPL+ +A QI A GP G N +YLF L ++ N+ E+
Sbjct: 115 NPDYLGPAPLEDIAEQIFNAAGPSGRNTEYLFELANSIRNLVPEE 159
>UniRef100_Q5PPV4 Hypothetical protein [Xenopus laevis]
Length = 184
Score = 120 bits (300), Expect = 3e-26
Identities = 71/165 (43%), Positives = 88/165 (53%), Gaps = 7/165 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W F Y EK++GYI Y R F DHRG P P R TL E W
Sbjct: 2 WVFGYGSLIWKVDFPYVEKLVGYIMCYSRRFWQGSTDHRGVPGKPGRVVTLVEDPEGCVW 61
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + G E+E V YL+ RE + V FY + S P V+++ T D N
Sbjct: 62 GVAYRLPEGKEEE--VKAYLDFREKGGYRTSTVVFYPKDPSIQP--FNVLLYIGTCDNPN 117
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHED 168
YLGPAPL+ +A QI A GP G N +YLF L ++ N+ ED
Sbjct: 118 ---YLGPAPLEDIAEQILNAVGPSGRNTEYLFELANSLRNLVPED 159
>UniRef100_UPI00003ACB07 UPI00003ACB07 UniRef100 entry
Length = 184
Score = 119 bits (299), Expect = 4e-26
Identities = 72/165 (43%), Positives = 86/165 (51%), Gaps = 7/165 (4%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL+W F Y EK++G I+ Y R F DHRG P P R TL E W
Sbjct: 2 WVFGYGSLIWKVDFPYQEKMVGRIRGYSRRFWQGSTDHRGVPGKPGRVVTLVEDPEGCVW 61
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVDFYKEGDSSHPALTGVIVFTSTPDKVN 123
G Y + G E E V YL+ RE T V FY + S P V+++ T D N
Sbjct: 62 GVHYRLPAGQECE--VKAYLDFREKGGYRTTTVVFYPKDSSIKP--FDVLLYIGTRDNPN 117
Query: 124 NKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHNIGHED 168
YLGPAPL +A QI A GP G N +YLF L +M N+ ED
Sbjct: 118 ---YLGPAPLQEIAEQIIDAVGPSGRNTEYLFELANSMRNLVPED 159
>UniRef100_Q9VCI9 CG10365-PA, isoform A [Drosophila melanogaster]
Length = 284
Score = 117 bits (294), Expect = 2e-25
Identities = 74/213 (34%), Positives = 105/213 (48%), Gaps = 16/213 (7%)
Query: 4 WVFGYGSLVWNPGFEYDEKVIGYIKDYRRVFDLACIDHRGTPEHPARTCTLEEKEGEICW 63
WVFGYGSL W+PGF Y + + GYI+ Y R F + HRG E P R TL E + I W
Sbjct: 58 WVFGYGSLCWHPGFNYTKCITGYIRGYVRRFWQGNVTHRGCEEKPGRVATLVEDKEGITW 117
Query: 64 GAVYCVRGGPEKEKLVMQYLERRECEYDHKTLVD--FYKEGDSSHPALTG----VIVFTS 117
G Y + G + YL++REC +D F+ S +G V+V+ +
Sbjct: 118 GCAYRITG-----STALDYLKQRECTLGGYATIDTKFFPRVASQDTPFSGEAVEVLVYVA 172
Query: 118 TPDKVNNKYYLGPAPLDVMARQIATAHGPCGNNRDYLFLLEKAMHN--IGHEDDMVIELA 175
TP+ N Y+LG P++ +A+QI + GP G+N +YL L MH G DD + EL
Sbjct: 173 TPE---NIYWLGDDPVEEIAQQIVSCRGPSGHNAEYLLRLALFMHEEIPGVRDDHLFELE 229
Query: 176 HEVRKVLGVGNVLPNDKKLSGPVQIQHSPHAPI 208
V + L + + P +I+ H I
Sbjct: 230 QLVLEELYRRQIPLSSVMGRNPDRIRRDSHEDI 262
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.320 0.141 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 440,480,091
Number of Sequences: 2790947
Number of extensions: 19846959
Number of successful extensions: 38905
Number of sequences better than 10.0: 134
Number of HSP's better than 10.0 without gapping: 128
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 38646
Number of HSP's gapped (non-prelim): 142
length of query: 224
length of database: 848,049,833
effective HSP length: 123
effective length of query: 101
effective length of database: 504,763,352
effective search space: 50981098552
effective search space used: 50981098552
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 72 (32.3 bits)
Lotus: description of TM0372b.7


