

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0369c.1
(573 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
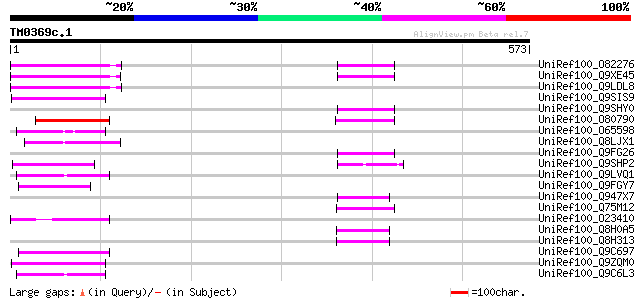
UniRef100_O82276 Putative non-LTR retroelement reverse transcrip... 74 9e-12
UniRef100_Q9XE45 Putative non-LTR retrolelement reverse transcri... 69 3e-10
UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis tha... 69 5e-10
UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcrip... 61 8e-08
UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana] 56 2e-06
UniRef100_O80790 Putative non-LTR retroelement reverse transcrip... 54 9e-06
UniRef100_O65598 Hypothetical protein M3E9.210 [Arabidopsis thal... 54 2e-05
UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor] 54 2e-05
UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like... 53 2e-05
UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcrip... 52 4e-05
UniRef100_Q9LVQ1 Non-LTR retroelement reverse transcriptase-like... 51 8e-05
UniRef100_Q9FGY7 Similarity to non-LTR retroelement reverse tran... 51 1e-04
UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sa... 50 1e-04
UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa] 50 2e-04
UniRef100_O23410 STRONG homology to reverse transcriptase [Arabi... 50 2e-04
UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa] 50 2e-04
UniRef100_Q8H313 Hypothetical protein P0710F09.110 [Oryza sativa] 50 2e-04
UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [A... 49 4e-04
UniRef100_Q9ZQM0 Putative non-LTR retroelement reverse transcrip... 49 4e-04
UniRef100_Q9C6L3 Hypothetical protein F2J7.11 [Arabidopsis thali... 49 5e-04
>UniRef100_O82276 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1231
Score = 74.3 bits (181), Expect = 9e-12
Identities = 44/123 (35%), Positives = 73/123 (58%), Gaps = 6/123 (4%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I ++ G +SH+ D ++LF S Q+++I + L+ F EASG KV+L+KS++F S+
Sbjct: 530 IAVSCGGSKLSHVCFADDLILFAEASVAQIRIIRRVLERFCEASGQKVSLEKSKIFFSHN 589
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPK 120
V + +Q +S GI LGKYL +P L+ R+ KE F ++++ ++ L G K
Sbjct: 590 VSREMEQLISEESGIGCTKELGKYLGMPILQKRMNKETFGEVLERVSARLA------GWK 643
Query: 121 GRA 123
GR+
Sbjct: 644 GRS 646
Score = 51.2 bits (121), Expect = 8e-05
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 2/65 (3%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG-- 420
IWK E+V F+WL ++ + TN++R HL + C C + E + H LR CP
Sbjct: 906 IWKLITPERVRVFIWLVSQNVIMTNVERVRRHLSENAICSVCNGAEETILHVLRDCPAME 965
Query: 421 SVWVR 425
+W R
Sbjct: 966 PIWRR 970
>UniRef100_Q9XE45 Putative non-LTR retrolelement reverse transcriptase [Arabidopsis
thaliana]
Length = 732
Score = 69.3 bits (168), Expect = 3e-10
Identities = 43/122 (35%), Positives = 70/122 (57%), Gaps = 6/122 (4%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I +++ G +SH+ D ++LF S Q+++I + L+ F ASG KV+L+KS++F S
Sbjct: 192 ISLSQGGPKLSHICFADDLILFAEASVAQIRVIRRVLERFCVASGQKVSLEKSKIFFSEN 251
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPK 120
V + + +S GI+ LGKYL +P L+ R+ K+ F I++K + L G K
Sbjct: 252 VSRDLGKLISDESGISSTRELGKYLGMPVLQRRINKDTFGDILEKLTTRLA------GWK 305
Query: 121 GR 122
GR
Sbjct: 306 GR 307
Score = 57.4 bits (137), Expect = 1e-06
Identities = 25/65 (38%), Positives = 35/65 (53%), Gaps = 2/65 (3%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG-- 420
+W TA E+V FLWL + A+ TN +R+ HL C C + E + H LR CP
Sbjct: 568 VWSVTAPERVRLFLWLVAQQAIMTNQERHRRHLSTTTICQVCKGAKESIIHVLRDCPAME 627
Query: 421 SVWVR 425
+W+R
Sbjct: 628 GIWLR 632
>UniRef100_Q9LDL8 Hypothetical protein AT4g08830 [Arabidopsis thaliana]
Length = 947
Score = 68.6 bits (166), Expect = 5e-10
Identities = 43/123 (34%), Positives = 71/123 (56%), Gaps = 6/123 (4%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I +++ G ISH+ D ++LF S Q+++I + L+ F ASG KV+L KS++F S
Sbjct: 344 IGLSQGGPKISHICFADDLILFAEASVSQIRVIRRILETFCIASGQKVSLDKSKIFFSKN 403
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPK 120
V + ++ +S GI LGKYL +P L+ R+ K+ F ++++ +S L G K
Sbjct: 404 VSRDLEKLISKESGIKSTRELGKYLGMPILQRRINKDTFGEVLERVSSRLA------GWK 457
Query: 121 GRA 123
GR+
Sbjct: 458 GRS 460
>UniRef100_Q9SIS9 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1715
Score = 61.2 bits (147), Expect = 8e-08
Identities = 36/103 (34%), Positives = 52/103 (49%)
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVD 62
ITK+ + ISHL D L F SN ++ + K++ EASG K+N KS + +
Sbjct: 1031 ITKDCLAISHLLFADDSLFFCRASNQNIEQLALIFKKYEEASGQKINYAKSSIIFGQKIP 1090
Query: 63 QRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
++Q L ++GI GKYL LP+ R E F I+ K
Sbjct: 1091 TMRRQRLHRLLGIDNVRGGGKYLGLPEQLGRRKVELFEYIVTK 1133
>UniRef100_Q9SHY0 F1E22.12 [Arabidopsis thaliana]
Length = 1055
Score = 56.2 bits (134), Expect = 2e-06
Identities = 26/65 (40%), Positives = 34/65 (52%), Gaps = 2/65 (3%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG-- 420
+WK E+V FLWL G A+ T +R+ HL + C C VE + H LR CP
Sbjct: 400 LWKVRVPERVKTFLWLVGNQAVMTEEERHRRHLSASNVCQVCKGGVESMLHVLRDCPAQL 459
Query: 421 SVWVR 425
+WVR
Sbjct: 460 GIWVR 464
Score = 46.6 bits (109), Expect = 0.002
Identities = 28/83 (33%), Positives = 47/83 (55%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I +++ G +SH+ D ++LF S Q+Q++ + L+ F ASG KV+L+KS++F S
Sbjct: 107 INLSRGGPKLSHICFADDLILFAEASVEQVQIVRRVLEAFCTASGQKVSLEKSKIFFSKN 166
Query: 61 VDQRKQQELSIIVGITRASYLGK 83
V + + +S GI GK
Sbjct: 167 VSRELGKLISDESGIQSTCDWGK 189
>UniRef100_O80790 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 970
Score = 54.3 bits (129), Expect = 9e-06
Identities = 25/68 (36%), Positives = 34/68 (49%), Gaps = 2/68 (2%)
Query: 360 MKLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP 419
+K IWK A E+V F+WL + TN++R HL C C + E + H LR CP
Sbjct: 644 LKQIWKLVAPERVRVFIWLVSHMVIMTNVERVRRHLSDIATCSVCNGADESILHVLRDCP 703
Query: 420 G--SVWVR 425
+W R
Sbjct: 704 AMTPIWQR 711
Score = 53.1 bits (126), Expect = 2e-05
Identities = 30/82 (36%), Positives = 50/82 (60%)
Query: 29 QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQELSIIVGITRASYLGKYLDLP 88
Q+ +I + L++F EASG V+L+KS++ SN V + ++ +S GI LGKYL +P
Sbjct: 390 QILIIRRVLEQFCEASGQNVSLEKSKIVFSNNVSRDMERLISGESGIGCTRELGKYLGMP 449
Query: 89 QLKARVTKEHFAPIIDKGNSHL 110
L+ R+ KE +++ +S L
Sbjct: 450 ILQKRMNKETSGEVLEHVSSSL 471
>UniRef100_O65598 Hypothetical protein M3E9.210 [Arabidopsis thaliana]
Length = 1141
Score = 53.5 bits (127), Expect = 2e-05
Identities = 31/98 (31%), Positives = 54/98 (54%), Gaps = 2/98 (2%)
Query: 8 IGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQ 67
+ ISHL D V++FF + L I +TL++F SG+KVN KS FC+ ++Q ++
Sbjct: 668 LSISHLMFADDVMIFFDGGSSSLHGICETLEDFASWSGLKVNNDKSHFFCAG-LEQAERN 726
Query: 68 ELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
L+ G + +YL LP + ++ + P+++K
Sbjct: 727 SLA-AYGFPQGCLPIRYLGLPLMCRKLRIAEYEPLLEK 763
>UniRef100_Q8LJX1 Putative reverse transcriptase [Sorghum bicolor]
Length = 1998
Score = 53.5 bits (127), Expect = 2e-05
Identities = 33/106 (31%), Positives = 55/106 (51%), Gaps = 1/106 (0%)
Query: 17 DGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQELSIIVGIT 76
D L+ QL ++ + F EA+G+KVNLQKS M N +D+ + L+ G +
Sbjct: 1494 DDTLIIAEGDTRQLLILKSIINTFSEATGLKVNLQKSMMLPIN-MDEERLDTLARTFGCS 1552
Query: 77 RASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHLGRISF*IGPKGR 122
+ S+ YL LP + T + + P+I+K + LG ++ + GR
Sbjct: 1553 KGSFPFTYLGLPLGITKPTIQDYQPLINKCEARLGSVATFLSEAGR 1598
>UniRef100_Q9FG26 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 676
Score = 53.1 bits (126), Expect = 2e-05
Identities = 22/65 (33%), Positives = 32/65 (48%), Gaps = 2/65 (3%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG-- 420
+W+ E+ FLWL G + TN +R H+ D C C + E + H LR CP
Sbjct: 347 VWRVMVPERARIFLWLVGNQVVLTNAERVRRHMADSDVCPLCKGASESLIHVLRDCPAMM 406
Query: 421 SVWVR 425
+W+R
Sbjct: 407 GIWMR 411
>UniRef100_Q9SHP2 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 773
Score = 52.4 bits (124), Expect = 4e-05
Identities = 29/90 (32%), Positives = 46/90 (50%)
Query: 4 TKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQ 63
++ G ISHL L+F + + + LK + +ASG VN QKS + +D
Sbjct: 125 SRNGPAISHLLFAHDSLVFCKATLEECMTLVNVLKLYEKASGQAVNFQKSAILFGKGLDF 184
Query: 64 RKQQELSIIVGITRASYLGKYLDLPQLKAR 93
R ++LS ++GI + G+YL LP+ R
Sbjct: 185 RTSEQLSQLLGIYKTEGFGRYLGLPEFVGR 214
Score = 52.0 bits (123), Expect = 5e-05
Identities = 31/76 (40%), Positives = 41/76 (53%), Gaps = 7/76 (9%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPT--NMKRYHSHLVKFDACFRCGASVEDVEHALRRCPG 420
IWK A K+ F W + +ALPT N+KR L+ D C RCG + EDV H L +C
Sbjct: 501 IWKQNAPPKIKHFWWRSAHNALPTAGNLKR--RRLITDDTCQRCGEASEDVNHLLFQCRV 558
Query: 421 S--VWVRSRFAKALPG 434
S +W ++ K PG
Sbjct: 559 SKEIWEQAHI-KLCPG 573
>UniRef100_Q9LVQ1 Non-LTR retroelement reverse transcriptase-like [Arabidopsis
thaliana]
Length = 1072
Score = 51.2 bits (121), Expect = 8e-05
Identities = 29/103 (28%), Positives = 56/103 (54%), Gaps = 2/103 (1%)
Query: 8 IGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQ 67
+ ISHL D V++FF + + I +TL +F + SG+KVN KS++F + + ++
Sbjct: 550 LSISHLMFADDVMIFFDGGSSSMHGICETLDDFADWSGLKVNKDKSQLFQAGL--DLSER 607
Query: 68 ELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHL 110
S G ++ +YL LP + ++ + P+++K ++ L
Sbjct: 608 ITSAAYGFPAGTFPIRYLGLPLMCRKLRIADYGPLLEKLSARL 650
>UniRef100_Q9FGY7 Similarity to non-LTR retroelement reverse transcriptase
[Arabidopsis thaliana]
Length = 171
Score = 50.8 bits (120), Expect = 1e-04
Identities = 27/80 (33%), Positives = 41/80 (50%)
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
++HL VD L F S Q +I + LK + SG ++N KS + + VD + E+
Sbjct: 10 VTHLLFVDDSLFFCKASKKQCGVILEILKHYETVSGQQINFTKSSIQFGHKVDDSLKAEI 69
Query: 70 SIIVGITRASYLGKYLDLPQ 89
+GIT +G YL LP+
Sbjct: 70 KATLGITNLGGMGSYLGLPE 89
>UniRef100_Q947X7 Hypothetical protein OSJNBa0067N01.20 [Oryza sativa]
Length = 387
Score = 50.4 bits (119), Expect = 1e-04
Identities = 20/57 (35%), Positives = 31/57 (54%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP 419
IW++ A +V FF WL ++ LPT + + ++ C C A++ED H RCP
Sbjct: 229 IWRSAAPPRVKFFAWLMSKNRLPTRVNLHKKTILPTPTCELCNANLEDTYHIFLRCP 285
>UniRef100_Q75M12 Hypothetical protein P0676G05.2 [Oryza sativa]
Length = 1936
Score = 50.1 bits (118), Expect = 2e-04
Identities = 25/66 (37%), Positives = 33/66 (49%), Gaps = 2/66 (3%)
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC-- 418
+LIWK +KV F W ++L T + +L +FD C C ED HAL RC
Sbjct: 1682 ELIWKCNVPQKVRIFAWRVASNSLATMENKKKRNLERFDVCGICDREKEDAGHALCRCVH 1741
Query: 419 PGSVWV 424
S+WV
Sbjct: 1742 ANSLWV 1747
Score = 41.6 bits (96), Expect = 0.064
Identities = 21/97 (21%), Positives = 44/97 (44%)
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVD 62
I ++ GIS+L D LLFF + +++ + L + + +G +N K +
Sbjct: 1309 ICRQAPGISYLLFADDTLLFFKAEKKEAEVVKEVLTNYAQGTGQLINPAKCSILFGEASP 1368
Query: 63 QRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHF 99
+++ + + R ++ +YL P + R+ K F
Sbjct: 1369 SSVSEDIRNTLQVERDNFEDRYLGFPTPEGRMHKGRF 1405
>UniRef100_O23410 STRONG homology to reverse transcriptase [Arabidopsis thaliana]
Length = 929
Score = 50.1 bits (118), Expect = 2e-04
Identities = 33/110 (30%), Positives = 56/110 (50%), Gaps = 17/110 (15%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I I++ G +SH+ D ++LF S Q KV+L+KS++F SN
Sbjct: 482 ISISQGGPKVSHVCFADDLILFAEASVAQ-----------------KVSLEKSKIFFSNN 524
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHL 110
V + + ++ GI LGKYL +P L+ R+ K+ F ++++ +S L
Sbjct: 525 VSRDLEGLITAETGIGSTRELGKYLGMPVLQKRINKDTFGEVLERVSSRL 574
Score = 40.8 bits (94), Expect = 0.11
Identities = 17/43 (39%), Positives = 24/43 (55%)
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFR 403
K IW A E+V FLWL G+ + TN++R H+ + C R
Sbjct: 822 KRIWGVIAPERVRVFLWLVGQQVIMTNVERVRRHIGDIEVCQR 864
>UniRef100_Q8H0A5 Putative reverse transcriptase [Oryza sativa]
Length = 1557
Score = 49.7 bits (117), Expect = 2e-04
Identities = 24/59 (40%), Positives = 29/59 (48%)
Query: 361 KLIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP 419
+L+WK KV F W A + LPT + +L D C CG ED HAL RCP
Sbjct: 1411 ELLWKCNVPSKVKIFTWRATSNCLPTWDNKKKRNLEISDTCVICGMEKEDTMHALCRCP 1469
Score = 42.7 bits (99), Expect = 0.029
Identities = 27/105 (25%), Positives = 45/105 (42%)
Query: 1 ICITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNV 60
I + + GISHL D LLFF Q Q I + + ++G +N K + N
Sbjct: 1036 IRVCHQAPGISHLLFADDTLLFFKADLSQSQAIKEVFGSYATSTGQLINPTKCSILFGNS 1095
Query: 61 VDQRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
+ + ++ + I + KYL LP R+ K F + ++
Sbjct: 1096 LPIASRDAITNCLQIASTEFEDKYLGLPTPGGRMHKGRFQSLRER 1140
>UniRef100_Q8H313 Hypothetical protein P0710F09.110 [Oryza sativa]
Length = 195
Score = 49.7 bits (117), Expect = 2e-04
Identities = 22/58 (37%), Positives = 29/58 (49%)
Query: 362 LIWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRCP 419
++WK +KV F+W A + LPT + L + C CG ED HAL RCP
Sbjct: 1 MVWKCNVPQKVKIFVWRAVTNCLPTLENKKKRKLEVSNVCSICGVETEDTGHALYRCP 58
>UniRef100_Q9C697 Reverse transcriptase, putative; 16838-20266 [Arabidopsis thaliana]
Length = 1142
Score = 48.9 bits (115), Expect = 4e-04
Identities = 27/101 (26%), Positives = 50/101 (48%)
Query: 10 ISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQEL 69
+SHL D L F + Q +I + LK++ SG ++N KS + + V+ + ++
Sbjct: 460 VSHLLFADDSLFFCKANKEQCGIILEILKQYESVSGQQINFSKSSIQFGHKVEDSIKADI 519
Query: 70 SIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDKGNSHL 110
+I+GI +G YL LP+ + F+ + D+ S +
Sbjct: 520 KLILGIHNLGGMGSYLGLPESLGGSKTKVFSFVRDRLQSRI 560
Score = 38.9 bits (89), Expect = 0.41
Identities = 22/75 (29%), Positives = 32/75 (42%), Gaps = 3/75 (4%)
Query: 363 IWKATAAEKVMFFLWLAGRDALPTNMKRYHSHLVKFDACFRCGASVEDVEHALRRC--PG 420
IWK K+ FLW +P + ++ C CGAS E + H L +C
Sbjct: 828 IWKVQCPPKLRHFLWQILSGCVPVSENLRKRGILCDKGCVSCGASEESINHTLFQCHPAR 887
Query: 421 SVWVRSRFAKALPGL 435
+W S+ A PG+
Sbjct: 888 QIWALSQIPTA-PGI 901
>UniRef100_Q9ZQM0 Putative non-LTR retroelement reverse transcriptase [Arabidopsis
thaliana]
Length = 1044
Score = 48.9 bits (115), Expect = 4e-04
Identities = 30/103 (29%), Positives = 48/103 (46%)
Query: 3 ITKEGIGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVD 62
++K ++HL D + F + LK++ EASG +N KS + S
Sbjct: 495 VSKGNPRVNHLLFADDTIFFCRSDLKSCKTFLCILKKYEEASGQMINKSKSAITFSRKTP 554
Query: 63 QRKQQELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
+ E I+GI LGKYL LP++ R ++ F I+D+
Sbjct: 555 DHIKTEAQQILGIQLVGGLGKYLGLPKMFGRKKRDLFNQIVDR 597
>UniRef100_Q9C6L3 Hypothetical protein F2J7.11 [Arabidopsis thaliana]
Length = 1213
Score = 48.5 bits (114), Expect = 5e-04
Identities = 27/98 (27%), Positives = 50/98 (50%), Gaps = 2/98 (2%)
Query: 8 IGISHLFCVDGVLLFFPVSN*QLQLITQTLKEFFEASGMKVNLQKSRMFCSNVVDQRKQQ 67
+ ISHL D V++FF + L I +TL +F SG+KVN KS ++ + + + +
Sbjct: 690 LSISHLMFADDVMIFFDGGSFSLHGICETLDDFASWSGLKVNKDKSHLYLAGL--NQLES 747
Query: 68 ELSIIVGITRASYLGKYLDLPQLKARVTKEHFAPIIDK 105
+ G + +YL LP + ++ + P+++K
Sbjct: 748 NANAAYGFPIGTLPIRYLGLPLMNRKLRIAEYEPLLEK 785
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.339 0.147 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 877,229,091
Number of Sequences: 2790947
Number of extensions: 32784547
Number of successful extensions: 81311
Number of sequences better than 10.0: 156
Number of HSP's better than 10.0 without gapping: 67
Number of HSP's successfully gapped in prelim test: 89
Number of HSP's that attempted gapping in prelim test: 81097
Number of HSP's gapped (non-prelim): 249
length of query: 573
length of database: 848,049,833
effective HSP length: 133
effective length of query: 440
effective length of database: 476,853,882
effective search space: 209815708080
effective search space used: 209815708080
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0369c.1


