

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0367.7
(172 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
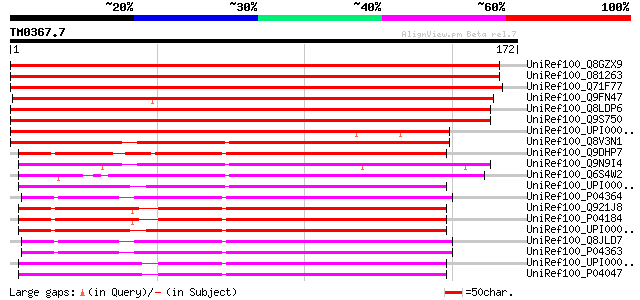
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8GZX9 Putative thymidine kinase [Oryza sativa] 270 1e-71
UniRef100_O81263 Thymidine kinase [Oryza sativa] 270 1e-71
UniRef100_Q71F77 TK1-like deoxyribonucleoside kinase [Lycopersic... 265 3e-70
UniRef100_Q9FN47 Similarity to thymidine kinase [Arabidopsis tha... 249 2e-65
UniRef100_Q8LDP6 Putative thymidine kinase [Arabidopsis thaliana] 243 1e-63
UniRef100_Q9S750 Putative thymidine kinase [Arabidopsis thaliana] 243 1e-63
UniRef100_UPI00002E9B20 UPI00002E9B20 UniRef100 entry 150 2e-35
UniRef100_Q8V3N1 Thymidine kinase [Swinepox virus] 131 6e-30
UniRef100_Q9DHP7 Thymidine kinase [Yaba-like disease virus] 122 3e-27
UniRef100_Q9N9I4 Thymidine kinase [Leishmania major] 119 3e-26
UniRef100_Q6S4W2 Thymidine kinase [Entamoeba histolytica] 119 3e-26
UniRef100_UPI0000361827 UPI0000361827 UniRef100 entry 118 5e-26
UniRef100_P04364 Thymidine kinase [Variola virus] 117 9e-26
UniRef100_Q921J8 Thymidine kinase 1 [Mus musculus] 117 1e-25
UniRef100_P04184 Thymidine kinase, cytosolic [Mus musculus] 117 1e-25
UniRef100_UPI0000428FDB UPI0000428FDB UniRef100 entry 117 2e-25
UniRef100_Q8JLD7 EVM078 [Ectromelia virus] 116 3e-25
UniRef100_P04363 Thymidine kinase [Monkeypox virus] 115 4e-25
UniRef100_UPI00003AD333 UPI00003AD333 UniRef100 entry 115 5e-25
UniRef100_P04047 Thymidine kinase, cytosolic [Gallus gallus] 115 5e-25
>UniRef100_Q8GZX9 Putative thymidine kinase [Oryza sativa]
Length = 270
Score = 270 bits (690), Expect = 1e-71
Identities = 125/166 (75%), Positives = 147/166 (88%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
NVA+IKS KD RYGLDS+VTHDGTK+PCWAL LSSF+ K G +AY+++DVIGIDEAQFF
Sbjct: 97 NVALIKSDKDNRYGLDSVVTHDGTKMPCWALPELSSFQDKLGTEAYDKVDVIGIDEAQFF 156
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
DDL+DFC +AAD DGK VVVAGLDG+Y R FGSVLDIIPLADSVTKLTARCE+CG+ AF
Sbjct: 157 DDLHDFCCKAADRDGKIVVVAGLDGDYKRNKFGSVLDIIPLADSVTKLTARCELCGRRAF 216
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLESQK 166
FTLRKT+ET+ ELIGG DVYMPVCRQHY++GQ+ +E TR+VL+ +K
Sbjct: 217 FTLRKTRETKTELIGGADVYMPVCRQHYLDGQIVIEATRIVLDLEK 262
>UniRef100_O81263 Thymidine kinase [Oryza sativa]
Length = 212
Score = 270 bits (690), Expect = 1e-71
Identities = 125/166 (75%), Positives = 147/166 (88%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
NVA+IKS KD RYGLDS+VTHDGTK+PCWAL LSSF+ K G +AY+++DVIGIDEAQFF
Sbjct: 39 NVALIKSDKDNRYGLDSVVTHDGTKMPCWALPELSSFQDKLGTEAYDKVDVIGIDEAQFF 98
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
DDL+DFC +AAD DGK VVVAGLDG+Y R FGSVLDIIPLADSVTKLTARCE+CG+ AF
Sbjct: 99 DDLHDFCCKAADRDGKIVVVAGLDGDYKRNKFGSVLDIIPLADSVTKLTARCELCGRRAF 158
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLESQK 166
FTLRKT+ET+ ELIGG DVYMPVCRQHY++GQ+ +E TR+VL+ +K
Sbjct: 159 FTLRKTRETKTELIGGADVYMPVCRQHYLDGQIVIEATRIVLDLEK 204
>UniRef100_Q71F77 TK1-like deoxyribonucleoside kinase [Lycopersicon esculentum]
Length = 234
Score = 265 bits (678), Expect = 3e-70
Identities = 122/167 (73%), Positives = 145/167 (86%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
NV +IKSSKD RY +D++VTHDGT+ PCW+L +LSSFKQ+FG DAYE++DVIGIDEAQFF
Sbjct: 55 NVVLIKSSKDARYAVDAVVTHDGTRFPCWSLPDLSSFKQRFGKDAYEKVDVIGIDEAQFF 114
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
DLY+FC AAD DGK +VVAGLDG+YLR++FGSVLDIIPLAD+VTKLTARCE+C + AF
Sbjct: 115 GDLYEFCCNAADFDGKIIVVAGLDGDYLRKSFGSVLDIIPLADTVTKLTARCELCNRRAF 174
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLESQKV 167
FT RKT ET+ ELIGG D+YMPVCRQHYVNGQ E+ ++VLES KV
Sbjct: 175 FTFRKTNETETELIGGADIYMPVCRQHYVNGQSVNESAKMVLESHKV 221
>UniRef100_Q9FN47 Similarity to thymidine kinase [Arabidopsis thaliana]
Length = 277
Score = 249 bits (637), Expect = 2e-65
Identities = 117/164 (71%), Positives = 142/164 (86%), Gaps = 1/164 (0%)
Query: 2 VAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYE-QLDVIGIDEAQFF 60
+AIIKS+KDTRY +SIVTHDG K PCW+L +LSSFK++FG D YE +LDVIGIDEAQFF
Sbjct: 103 IAIIKSNKDTRYCTESIVTHDGEKYPCWSLPDLSSFKERFGFDDYENRLDVIGIDEAQFF 162
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
DLY+FCREAAD +GKTV+VAGLDG+++RR FGSVLD+IP+AD+VTKLT+RCE+CGK A
Sbjct: 163 GDLYEFCREAADKEGKTVIVAGLDGDFMRRRFGSVLDLIPIADTVTKLTSRCEVCGKRAL 222
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLES 164
FT+RKT+E + ELIGG +VYMPVCR HYV GQ +ET R VL+S
Sbjct: 223 FTMRKTEEKETELIGGAEVYMPVCRSHYVCGQNVLETARAVLDS 266
>UniRef100_Q8LDP6 Putative thymidine kinase [Arabidopsis thaliana]
Length = 238
Score = 243 bits (621), Expect = 1e-63
Identities = 112/163 (68%), Positives = 137/163 (83%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
+VA++KSSKDTRY DS+VTHDG PCWAL +L SF +KFG+DAY +LDVIGIDEAQFF
Sbjct: 61 SVAMLKSSKDTRYAKDSVVTHDGIGFPCWALPDLMSFPEKFGLDAYNKLDVIGIDEAQFF 120
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
DLY+FC + AD DGK V+VAGLDG+YLRR+FG+VLDIIP+ADSVTKLTARCE+CG AF
Sbjct: 121 GDLYEFCCKVADDDGKIVIVAGLDGDYLRRSFGAVLDIIPIADSVTKLTARCEVCGHKAF 180
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLE 163
FTLRK +T+ ELIGG DVYMPVCR+HY+ + ++ ++ VLE
Sbjct: 181 FTLRKNCDTRTELIGGADVYMPVCRKHYITNHIVIKASKKVLE 223
>UniRef100_Q9S750 Putative thymidine kinase [Arabidopsis thaliana]
Length = 238
Score = 243 bits (621), Expect = 1e-63
Identities = 112/163 (68%), Positives = 137/163 (83%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
+VA++KSSKDTRY DS+VTHDG PCWAL +L SF +KFG+DAY +LDVIGIDEAQFF
Sbjct: 61 SVAMLKSSKDTRYAKDSVVTHDGIGFPCWALPDLMSFPEKFGLDAYNKLDVIGIDEAQFF 120
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
DLY+FC + AD DGK V+VAGLDG+YLRR+FG+VLDIIP+ADSVTKLTARCE+CG AF
Sbjct: 121 GDLYEFCCKVADDDGKIVIVAGLDGDYLRRSFGAVLDIIPIADSVTKLTARCEVCGHKAF 180
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLVLE 163
FTLRK +T+ ELIGG DVYMPVCR+HY+ + ++ ++ VLE
Sbjct: 181 FTLRKNCDTRTELIGGADVYMPVCRKHYITNHIVIKASKKVLE 223
>UniRef100_UPI00002E9B20 UPI00002E9B20 UniRef100 entry
Length = 220
Score = 150 bits (378), Expect = 2e-35
Identities = 77/151 (50%), Positives = 99/151 (64%), Gaps = 2/151 (1%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
+V ++KSSKD RY D IV+HDGT C A+A LS + G + + V+ IDEAQF
Sbjct: 63 SVTLVKSSKDNRYSHDEIVSHDGTSRRCNAVAELSDLRGIIGEEQWNGAHVLAIDEAQFI 122
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICG-KHA 119
DL FC AAD K V+VAGLDG++ R+ FG VLD++PL D+VTKLTA+C +C + A
Sbjct: 123 PDLVSFCVNAADEHQKKVIVAGLDGDFRRQRFGEVLDLLPLCDTVTKLTAQCGVCSVRPA 182
Query: 120 FFTLRKTQETQV-ELIGGVDVYMPVCRQHYV 149
F+LR T +T EL+GG D Y P CR YV
Sbjct: 183 LFSLRTTGDTSTQELVGGADSYKPACRLCYV 213
>UniRef100_Q8V3N1 Thymidine kinase [Swinepox virus]
Length = 181
Score = 131 bits (330), Expect = 6e-30
Identities = 66/149 (44%), Positives = 96/149 (64%), Gaps = 6/149 (4%)
Query: 1 NVAIIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFF 60
N +IK SKD RYG D++ THD + + +L K D + +D++GIDE QFF
Sbjct: 38 NCRVIKYSKDNRYGNDAVYTHDKCYISAVSTDSLFDIK-----DTLDDVDIVGIDEGQFF 92
Query: 61 DDLYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAF 120
+D+ +FC A+ GK V+VA LDG Y R+ FG++L++IPL++ VTKL A C IC + A
Sbjct: 93 NDIVEFCEYIANK-GKIVIVAALDGTYERKPFGNILNLIPLSEKVTKLNAICMICHRDAS 151
Query: 121 FTLRKTQETQVELIGGVDVYMPVCRQHYV 149
F+ R + E ++ELIGG + Y+ VCR Y+
Sbjct: 152 FSKRLSDEKEIELIGGKEKYLSVCRSCYL 180
>UniRef100_Q9DHP7 Thymidine kinase [Yaba-like disease virus]
Length = 183
Score = 122 bits (307), Expect = 3e-27
Identities = 69/145 (47%), Positives = 91/145 (62%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
+IK SKDTRYG +VTHD +P + +LS + D + DVIGIDE QFF D+
Sbjct: 39 VIKYSKDTRYG-KGLVTHDNNSIPAIPVNSLS----EINCDKIKA-DVIGIDEGQFFPDI 92
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
+FC A+ DGK V+VA LDG +LR FG++L +IP A+ V+KLTA C C A F+
Sbjct: 93 VEFCERMAN-DGKIVIVAALDGTFLREPFGNILKLIPCAEYVSKLTAVCMNCFNSASFSK 151
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R E ++E+IGG D Y VCR+ Y
Sbjct: 152 RIGDEQEIEVIGGKDKYQSVCRKCY 176
>UniRef100_Q9N9I4 Thymidine kinase [Leishmania major]
Length = 284
Score = 119 bits (298), Expect = 3e-26
Identities = 68/166 (40%), Positives = 99/166 (58%), Gaps = 12/166 (7%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWA-LANLSSFKQKFGVDAYEQLDVIGIDEAQFFDD 62
+IK SKDTRY ++ +HD L A ++ L+ + D +++ DV+ IDE QFF D
Sbjct: 37 VIKYSKDTRYDEHNVASHDQLMLRAQAAVSQLTEVR-----DTWKRFDVLAIDEGQFFSD 91
Query: 63 LYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKH-AFF 121
L DFC AAD GK V+V+ LDG+Y R+ FG + +++P ++V KLTA C +C + A F
Sbjct: 92 LVDFCNTAADA-GKVVMVSALDGDYRRKPFGQICELVPYCEAVDKLTAVCMMCHEQPACF 150
Query: 122 TLRKTQETQVELIGGVDVYMPVCRQHYVNGQV----AMETTRLVLE 163
T R Q ELIGG D+Y+ CR+ Y Q+ M T R+ ++
Sbjct: 151 TRRTVNVEQQELIGGADMYIATCRECYSKQQLPSIEEMRTQRMAIK 196
>UniRef100_Q6S4W2 Thymidine kinase [Entamoeba histolytica]
Length = 218
Score = 119 bits (298), Expect = 3e-26
Identities = 67/159 (42%), Positives = 94/159 (58%), Gaps = 7/159 (4%)
Query: 4 IIKSSKDTRYGL-DSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDD 62
+IK SKDTRYG D ++HD W + + K ++ +VIGIDE QFF D
Sbjct: 57 VIKYSKDTRYGSEDEAISHDKES---WKA--IPTMKLMPVLETALNYEVIGIDEGQFFPD 111
Query: 63 LYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFT 122
L +F A + G+ V++A LDG + R+ FG + D+IPL +SV KL+A C CGK A F+
Sbjct: 112 LIEFSEACASY-GRLVIIAALDGTFQRKPFGQITDLIPLCESVKKLSAVCVNCGKKAAFS 170
Query: 123 LRKTQETQVELIGGVDVYMPVCRQHYVNGQVAMETTRLV 161
LR + E +E+IGGVD Y VCR+ + ++T R V
Sbjct: 171 LRTSSEESIEVIGGVDKYCAVCRKCFYKDGATLKTQRSV 209
>UniRef100_UPI0000361827 UPI0000361827 UniRef100 entry
Length = 225
Score = 118 bits (296), Expect = 5e-26
Identities = 64/145 (44%), Positives = 87/145 (59%), Gaps = 6/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
+IK +KDTRY + THD + L + Q+ VIG+DE QFF D+
Sbjct: 46 VIKYAKDTRYSDSGMATHDKNTMEAVPANRLGDVHSQA-----LQVCVIGVDEGQFFPDI 100
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
+FC E A+ GKT++VA LDG + R+ FGS+L+++PLA+SV KL A C C K A +T
Sbjct: 101 VEFCEEMANL-GKTIIVAALDGTFQRKPFGSILNLVPLAESVVKLNAVCMQCFKEAAYTK 159
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR+ Y
Sbjct: 160 RIGAEKEVEVIGGADKYQAVCRKCY 184
>UniRef100_P04364 Thymidine kinase [Variola virus]
Length = 177
Score = 117 bits (294), Expect = 9e-26
Identities = 63/146 (43%), Positives = 87/146 (59%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD L ++A VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFEALEATKLCDV-----LEAITDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + R+ F ++LD+IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRKPFNNILDLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
ET++E+IGG+D+Y VCR+ Y++
Sbjct: 151 LGTETKIEIIGGIDMYQSVCRKCYID 176
>UniRef100_Q921J8 Thymidine kinase 1 [Mus musculus]
Length = 233
Score = 117 bits (293), Expect = 1e-25
Identities = 69/146 (47%), Positives = 89/146 (60%), Gaps = 9/146 (6%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQK-FGVDAYEQLDVIGIDEAQFFDD 62
+IK +KDTRY +S THD + L Q+ GV VIGIDE QFF D
Sbjct: 52 VIKYAKDTRYS-NSFSTHDRNTMDALPACMLRDVTQESLGVA------VIGIDEGQFFPD 104
Query: 63 LYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFT 122
+ DFC A+ +GKTV+VA LDG + R+ FGS+L+++PLA+SV KLTA C C + A +T
Sbjct: 105 IVDFCEMMAN-EGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLTAVCMECFREAAYT 163
Query: 123 LRKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 164 KRLGLEKEVEVIGGADKYHSVCRLCY 189
>UniRef100_P04184 Thymidine kinase, cytosolic [Mus musculus]
Length = 233
Score = 117 bits (293), Expect = 1e-25
Identities = 69/146 (47%), Positives = 89/146 (60%), Gaps = 9/146 (6%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQK-FGVDAYEQLDVIGIDEAQFFDD 62
+IK +KDTRY +S THD + L Q+ GV VIGIDE QFF D
Sbjct: 52 VIKYAKDTRYS-NSFSTHDRNTMDALPACMLRDVTQEALGVA------VIGIDEGQFFPD 104
Query: 63 LYDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFT 122
+ DFC A+ +GKTV+VA LDG + R+ FGS+L+++PLA+SV KLTA C C + A +T
Sbjct: 105 IVDFCEMMAN-EGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLTAVCMECFREAAYT 163
Query: 123 LRKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 164 KRLGLEKEVEVIGGADKYHSVCRLCY 189
>UniRef100_UPI0000428FDB UPI0000428FDB UniRef100 entry
Length = 259
Score = 117 bits (292), Expect = 2e-25
Identities = 67/145 (46%), Positives = 88/145 (60%), Gaps = 7/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
+IK +KDTRY +S THD + L Q+ + VIGIDE QFF D+
Sbjct: 77 VIKYAKDTRYS-NSFSTHDRNTIDALPACMLRDVTQES-----LSVAVIGIDEGQFFPDI 130
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
DFC A+ +GKTV+VA LDG + R+ FGS+L+++PLA+SV KLTA C C + A +T
Sbjct: 131 VDFCEMMAN-EGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLTAVCMECFREAAYTK 189
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 190 RLGLEKEVEVIGGADKYHSVCRLCY 214
>UniRef100_Q8JLD7 EVM078 [Ectromelia virus]
Length = 177
Score = 116 bits (290), Expect = 3e-25
Identities = 63/146 (43%), Positives = 86/146 (58%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD L +DA VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFEALEATKLCDV-----MDAVTDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + R+ F ++LD+IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRKPFNNILDLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
ET++E+IGG D+Y +CR+ Y++
Sbjct: 151 LGTETEIEIIGGNDMYQSMCRKCYID 176
>UniRef100_P04363 Thymidine kinase [Monkeypox virus]
Length = 177
Score = 115 bits (289), Expect = 4e-25
Identities = 63/146 (43%), Positives = 87/146 (59%), Gaps = 7/146 (4%)
Query: 5 IKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDLY 64
IK S D RYG + THD + L ++A VIGIDE QFF D+
Sbjct: 38 IKYSNDNRYGT-GLWTHDKNNFAALEVTKLCDV-----LEAITDFSVIGIDEGQFFPDIV 91
Query: 65 DFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTLR 124
+FC A+ +GK V+VA LDG + RR F ++L++IPL++ V KLTA C C K A F+ R
Sbjct: 92 EFCERMAN-EGKIVIVAALDGTFQRRPFNNILNLIPLSEMVVKLTAVCMKCFKEASFSKR 150
Query: 125 KTQETQVELIGGVDVYMPVCRQHYVN 150
ET++E+IGG D+Y VCR+ Y++
Sbjct: 151 LGAETEIEIIGGNDMYQSVCRKCYID 176
>UniRef100_UPI00003AD333 UPI00003AD333 UniRef100 entry
Length = 204
Score = 115 bits (288), Expect = 5e-25
Identities = 64/145 (44%), Positives = 86/145 (59%), Gaps = 6/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
++K +KDTRY + THD + L Q+ A VIGIDE QFF D+
Sbjct: 32 LVKYAKDTRYCTTGVSTHDRNTMEARPACALQDVYQEALGSA-----VIGIDEGQFFPDI 86
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
+FC + A+ GKTV+VA LDG + R+ FGS+L+++PLA+SV KL A C C + A +T
Sbjct: 87 VEFCEKMAN-TGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLNAVCMECYREASYTK 145
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 146 RLGAEREVEVIGGADKYHSVCRACY 170
>UniRef100_P04047 Thymidine kinase, cytosolic [Gallus gallus]
Length = 224
Score = 115 bits (288), Expect = 5e-25
Identities = 64/145 (44%), Positives = 86/145 (59%), Gaps = 6/145 (4%)
Query: 4 IIKSSKDTRYGLDSIVTHDGTKLPCWALANLSSFKQKFGVDAYEQLDVIGIDEAQFFDDL 63
++K +KDTRY + THD + L Q+ A VIGIDE QFF D+
Sbjct: 52 LVKYAKDTRYCTTGVSTHDRNTMEARPACALQDVYQEALGSA-----VIGIDEGQFFPDI 106
Query: 64 YDFCREAADHDGKTVVVAGLDGNYLRRNFGSVLDIIPLADSVTKLTARCEICGKHAFFTL 123
+FC + A+ GKTV+VA LDG + R+ FGS+L+++PLA+SV KL A C C + A +T
Sbjct: 107 VEFCEKMAN-TGKTVIVAALDGTFQRKAFGSILNLVPLAESVVKLNAVCMECYREASYTK 165
Query: 124 RKTQETQVELIGGVDVYMPVCRQHY 148
R E +VE+IGG D Y VCR Y
Sbjct: 166 RLGAEREVEVIGGADKYHSVCRACY 190
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 270,596,327
Number of Sequences: 2790947
Number of extensions: 10586027
Number of successful extensions: 22618
Number of sequences better than 10.0: 295
Number of HSP's better than 10.0 without gapping: 171
Number of HSP's successfully gapped in prelim test: 124
Number of HSP's that attempted gapping in prelim test: 22190
Number of HSP's gapped (non-prelim): 297
length of query: 172
length of database: 848,049,833
effective HSP length: 118
effective length of query: 54
effective length of database: 518,718,087
effective search space: 28010776698
effective search space used: 28010776698
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0367.7


