

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0366.1
(1346 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
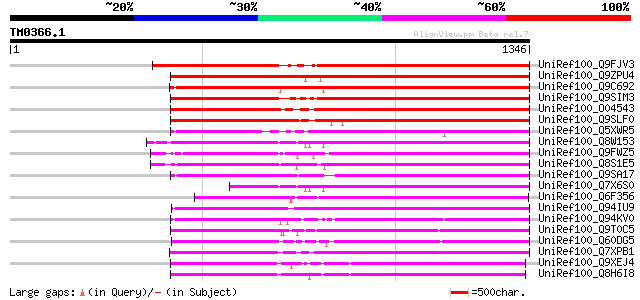
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FJV3 Retroelement pol polyprotein-like [Arabidopsis ... 841 0.0
UniRef100_Q9ZPU4 Putative retroelement pol polyprotein [Arabidop... 812 0.0
UniRef100_Q9C692 Polyprotein, putative [Arabidopsis thaliana] 799 0.0
UniRef100_Q9SIM3 Putative retroelement pol polyprotein [Arabidop... 795 0.0
UniRef100_O04543 F20P5.25 protein [Arabidopsis thaliana] 787 0.0
UniRef100_Q9SLF0 Putative retroelement pol polyprotein [Arabidop... 748 0.0
UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Sol... 747 0.0
UniRef100_Q8W153 Polyprotein [Oryza sativa] 720 0.0
UniRef100_Q9FWZ5 Putative retroelement polyprotein [Arabidopsis ... 693 0.0
UniRef100_Q8S1E5 Putative gag/pol polyprotein [Oryza sativa] 689 0.0
UniRef100_Q9SA17 F28K20.17 protein [Arabidopsis thaliana] 635 e-180
UniRef100_Q7X6S0 OSJNBb0011N17.2 protein [Oryza sativa] 617 e-174
UniRef100_Q6F356 Putative polyprotein [Oryza sativa] 612 e-173
UniRef100_Q94IU9 Copia-like polyprotein [Arabidopsis thaliana] 611 e-173
UniRef100_Q94KV0 Polyprotein [Arabidopsis thaliana] 600 e-169
UniRef100_Q9T0C5 Retrotransposon like protein [Arabidopsis thali... 591 e-167
UniRef100_Q60DG5 Putative polyprotein [Oryza sativa] 568 e-160
UniRef100_Q7XPB1 OSJNBb0026E15.10 protein [Oryza sativa] 566 e-159
UniRef100_Q9XEJ4 Copia-type pol polyprotein [Zea mays] 560 e-158
UniRef100_Q8H6I8 Putative gag-pol polyprotein [Zea mays] 555 e-156
>UniRef100_Q9FJV3 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1475
Score = 841 bits (2172), Expect = 0.0
Identities = 435/986 (44%), Positives = 620/986 (62%), Gaps = 66/986 (6%)
Query: 370 QRRIGLGSLQPDELYHLVVSSSSSTCFSIQHSSTSEINDFGHIIPPGALWHFRLGHLSHD 429
+++IG G+ Q LY V+ +SS C S+ +S+ + + + ALWH RLGH S++
Sbjct: 540 EQKIGRGN-QIGGLY--VLDTSSVECTSVDINSSVTEKQYCNAVVDSALWHSRLGHPSYE 596
Query: 430 RLLALHTVQNSISVSKS--IVCDVCHLAKQKRKMFTVSVSKAQKCFDLVHMDIWGPLAQA 487
+ LH V +K + C +C AKQK F + ++ FDL+H+D WGP A
Sbjct: 597 KNDVLHDVLGLPKRNKEDLVHCSICQKAKQKHLSFPSKNNMSENKFDLIHIDTWGPFATP 656
Query: 488 SVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIVKAIRSDNGPEF 547
+ +KYFLT++DDYSR WV L+ K +V Q +F+ +V+TQ+G +VKA+RSDN PE
Sbjct: 657 TTEGYKYFLTIVDDYSRATWVYLMKAKNDVLQIFPDFLKMVETQYGTLVKAVRSDNAPEL 716
Query: 548 LLPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAV 607
A Y A+GI+ SC TPQQN VERKHQHILN+ARAL+F++ +P + W +L AV
Sbjct: 717 RFEALYQAKGIISYHSCPETPQQNSVVERKHQHILNVARALMFEANMPLEFWGDCILSAV 776
Query: 608 FLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLG 667
FL+NR+P+ LL NKSP+ELL+ + D +KVFG LC+ +T R K PRAR C+FLG
Sbjct: 777 FLINRLPTPLLSNKSPFELLHLKVPDYTSLKVFGCLCYESTSPQQRHKFAPRARACVFLG 836
Query: 668 YKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQTQAESTADKLNVA 727
Y G KGY L+DL T I ISR V+F+E V P+ DK +
Sbjct: 837 YPSGYKGYKLLDLETNTIHISRHVVFYETVFPF--------------------TDKTIIP 876
Query: 728 SEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPNTQTVDEIPQPGSITAEGEVEV 787
++ + D V + + S + P +V
Sbjct: 877 RDV-------------FDLVDPVHENIENPPSTSESAP------------------KVSS 905
Query: 788 RRSTRPLKRPVHLANFQFSALP-IRSSTAHSIVHHYSDERLSTAHRAYALNIAQDQEPST 846
+R +RP P +L ++ +A+P + + + + + +LS AY + + EP T
Sbjct: 906 KRESRP---PGYLQDYFCNAVPDVTKDVRYPLNAYINYTQLSEEFTAYICAVNKYPEPCT 962
Query: 847 FAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKA 906
+A+A K W +AM+ EI ALE TW + LP+G KPIG KWV++VK N DG+L R+KA
Sbjct: 963 YAQAKKIKEWLDAMEIEIDALESTNTWSVCSLPQGKKPIGCKWVFKVKLNADGSLERFKA 1022
Query: 907 RLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDV 966
RLVAKGY Q EGLDY+DTFSPVAK+TTV+ +L++AA + W LHQLD+ NAFL+G+L E++
Sbjct: 1023 RLVAKGYTQREGLDYYDTFSPVAKMTTVKTLLSVAAIKEWSLHQLDISNAFLNGDLKEEI 1082
Query: 967 YMTIPAGVPS-----VKPNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSL 1021
YMT+P G + N V KL KSLYGLKQASR+WY K S+ L+ LGFK++ +DH+L
Sbjct: 1083 YMTLPPGYSMKQGGVLPQNPVLKLQKSLYGLKQASRQWYLKFSSTLKKLGFKKSHADHTL 1142
Query: 1022 FVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHST 1081
F + G ++ LLVYVDD+++ GN + +K L +AF ++DLG +KYFLGLE+A +
Sbjct: 1143 FTRISGKAYIALLVYVDDIVIAGNNDENIEELKKDLAKAFKLRDLGPMKYFLGLEIARTK 1202
Query: 1082 SGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPY-DDPAAYRRLVGRLLY 1140
GIS+CQRKY ++LL++TG LG +P P++PS++LSQ + D+P YRRLVG+L+Y
Sbjct: 1203 EGISVCQRKYTMELLEDTGLLGCRPSTIPMEPSLKLSQHNDEHVIDNPEVYRRLVGKLMY 1262
Query: 1141 LTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSD 1200
LT TRPDI++A +L Q+ S+P + H KAAQ+V+ YLK + GLGL + S + ++ Y+D
Sbjct: 1263 LTITRPDITYAINRLCQFSSSPKNSHLKAAQKVVHYLKGTIGLGLFYSSKSDLCLKAYTD 1322
Query: 1201 ADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYL 1260
ADW C D+RRS SG C ++G SL+SWK+KKQ S SS E+EYRA+A + E+ WL+ L
Sbjct: 1323 ADWGSCVDSRRSTSGICMFLGDSLISWKSKKQNMASSSSAESEYRAMAMGSREIAWLVKL 1382
Query: 1261 LQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPT 1320
L + +V T LFCD+ +A+HIA N VFHERTKH++ DCH+ R++++ G+LK + + T
Sbjct: 1383 LAEFQVKQTKPVPLFCDSTAAIHIANNAVFHERTKHIENDCHITRDRIEQGMLKTMHVDT 1442
Query: 1321 TLQVADVFTKALQPRVFQGFATKLAM 1346
T Q+ADV TK L P +F K+++
Sbjct: 1443 TSQLADVLTKPLFPTLFNSLIGKMSL 1468
>UniRef100_Q9ZPU4 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1501
Score = 812 bits (2097), Expect = 0.0
Identities = 434/967 (44%), Positives = 589/967 (60%), Gaps = 37/967 (3%)
Query: 417 ALWHFRLGHLSHDRLLALHTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLV 476
ALWH RLGH S L +L + S S CDVC AKQ R++F S++K ++CF L+
Sbjct: 528 ALWHQRLGHPSFSVLSSLPLFSKTSSTVTSHSCDVCFRAKQTREVFPESINKTEECFSLI 587
Query: 477 HMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIV 536
H D+WGP + YFLT++DDYSR VW LL K EV+Q + NF+ + QFG+ V
Sbjct: 588 HCDVWGPYRVPASCGAVYFLTIVDDYSRAVWTYLLLEKSEVRQVLTNFLKYAEKQFGKTV 647
Query: 537 KAIRSDNGPEFL-LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLP 595
K +RSDNG EF+ L +++ GI+HQ SCV TPQQNGRVERKH+HILN+ARALLFQ+ LP
Sbjct: 648 KMVRSDNGTEFMCLSSYFRENGIIHQTSCVGTPQQNGRVERKHRHILNVARALLFQASLP 707
Query: 596 KKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSK 655
K W S+L A +L+NR PS +L ++PYE+L+ ++VFGS C+ +T ++ K
Sbjct: 708 IKFWGESILTAAYLINRTPSSILSGRTPYEVLHGSKPVYSQLRVFGSACYVHRVTRDKDK 767
Query: 656 LDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQT 715
R+R CIF+GY G KG+ + D+ E +SRDVIF E V PY G NS L T
Sbjct: 768 FGQRSRSCIFVGYPFGKKGWKVYDIERNEFLVSRDVIFREEVFPYAGVNSSTLASTSLPT 827
Query: 716 QAES---TADKLNVASEIQSAKGDASVSTPCASIEDGV--QHEMPSGASIEDEIP----- 765
+E L V I S + + V T + D E+P+ + D+ P
Sbjct: 828 VSEDDDWAIPPLEVRGSIDSVETERVVCTTDEVVLDTSVSDSEIPNQEFVPDDTPPSSPL 887
Query: 766 --------NTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANF-------------Q 804
NT T + S + R+S R P L ++
Sbjct: 888 SVSPSGSPNTPTTPIVVPVASPIPVSPPKQRKSKRATHPPPKLNDYVLYNAMYTPSSIHA 947
Query: 805 FSALPIRSSTAHS-----IVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREA 859
A P +SST + + SD S++HRAY I + EP F EA + W +A
Sbjct: 948 LPADPSQSSTVPGKSLFPLTDYVSDAAFSSSHRAYLAAITDNVEPKHFKEAVQIKVWNDA 1007
Query: 860 MQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGL 919
M E+ ALE N TW +VDLP G IG++WV++ K N DGT+ RYKARLV +G QVEG
Sbjct: 1008 MFTEVDALEINKTWDIVDLPPGKVAIGSQWVFKTKYNSDGTVERYKARLVVQGNKQVEGE 1067
Query: 920 DYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPSVKP 979
DY +TF+PV ++TTVR +L A+ W ++Q+DV NAFLHG+L+E+VYM +P G P
Sbjct: 1068 DYKETFAPVVRMTTVRTLLRNVAANQWEVYQMDVHNAFLHGDLEEEVYMKLPPGFRHSHP 1127
Query: 980 NQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDD 1039
++VC+L KSLYGLKQA R W++KLS L GF Q+ D+SLF + + +L+YVDD
Sbjct: 1128 DKVCRLRKSLYGLKQAPRCWFKKLSDSLLRFGFVQSYEDYSLFSYTRNNIELRVLIYVDD 1187
Query: 1040 VILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQET 1099
+++ GN Q KD L + F +KDLG LKYFLG+EV+ GI L QRKY LD++ ++
Sbjct: 1188 LLICGNDGYMLQKFKDYLSRCFSMKDLGKLKYFLGIEVSRGPEGIFLSQRKYALDVIADS 1247
Query: 1100 GTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYM 1159
G LGS+P TPL+ + L+ + G DP YRRLVGRLLYL TRP++S++ L+Q+M
Sbjct: 1248 GNLGSRPAHTPLEQNHHLASDDGPLLSDPKPYRRLVGRLLYLLHTRPELSYSVHVLAQFM 1307
Query: 1160 SNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFY 1219
NP + HF AA RV+RYLK SPG G+L + + ++ Y D+DW CP TRRSIS Y
Sbjct: 1308 QNPREAHFDAALRVVRYLKGSPGQGILLNADPDLTLEVYCDSDWQSCPLTRRSISAYVVL 1367
Query: 1220 IGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQ 1279
+G S +SWK KKQ TVS SS EAEYRA++YA E++WL LL++L + + L+CD++
Sbjct: 1368 LGGSPISWKTKKQDTVSHSSAEAEYRAMSYALKEIKWLRKLLKELGIEQSTPARLYCDSK 1427
Query: 1280 SALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQG 1339
+A+HIAANPVFHERTKH++ DCH VR+ ++ GI+ + TT Q+ADVFTKAL F
Sbjct: 1428 AAIHIAANPVFHERTKHIESDCHSVRDAVRDGIITTQHVRTTEQLADVFTKALGRNQFLY 1487
Query: 1340 FATKLAM 1346
+KL +
Sbjct: 1488 LMSKLGV 1494
>UniRef100_Q9C692 Polyprotein, putative [Arabidopsis thaliana]
Length = 1468
Score = 799 bits (2064), Expect = 0.0
Identities = 424/952 (44%), Positives = 595/952 (61%), Gaps = 22/952 (2%)
Query: 415 PGALWHFRLGHLSHDRLLALHTVQNSISVSKSI---VCDVCHLAKQKRKMFTVSVSKAQK 471
P LWH RLGH S D+++ L + +S K I VCD C AKQ R F +S +++
Sbjct: 512 PFDLWHRRLGHAS-DKIVNL-LPRELLSSGKEILENVCDTCMRAKQTRDTFPLSDNRSMD 569
Query: 472 CFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQ 531
F L+H D+WGP S +YFLT++DDYSR VWV L+ +K E Q+ +K+FI LV+ Q
Sbjct: 570 SFQLIHCDVWGPYRAPSYSGARYFLTIVDDYSRGVWVYLMTDKSETQKHLKDFIALVERQ 629
Query: 532 FGQIVKAIRSDNGPEFL-LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLF 590
F +K +RSDNG EFL + ++ +GI H+ SCV TP QNGRVERKH+HILNIARAL F
Sbjct: 630 FDTEIKIVRSDNGTEFLCMREYFLHKGIAHETSCVGTPHQNGRVERKHRHILNIARALRF 689
Query: 591 QSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLT 650
QS LP + W +L A +L+NR PS LL+ KSPYE+LY+ A ++VFGSLC+A
Sbjct: 690 QSYLPIQFWGECILSAAYLINRTPSMLLQGKSPYEMLYKTAPKYSHLRVFGSLCYAHNQN 749
Query: 651 NNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPY------KGSN 704
+ K R+R+C+F+GY G KG+ L DL Q+ F+SRDVIF E PY +
Sbjct: 750 HKGDKFAARSRRCVFVGYPHGQKGWRLFDLEEQKFFVSRDVIFQETEFPYSKMSCNEEDE 809
Query: 705 SPAWTCLDLQTQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPS---GASIE 761
C+ E+ + + I A +V+T E + PS S
Sbjct: 810 RVLVDCVGPPFIEEAIGPRTIIGRNIGEATVGPNVATGPIIPEINQESSSPSEFVSLSSL 869
Query: 762 DEIPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRS-------ST 814
D + TV P S T +++RRS+R ++P+ L NF + + + S S+
Sbjct: 870 DPFLASSTVQTADLPLSSTTPAPIQLRRSSRQTQKPMKLKNFVTNTVSVESISPEASSSS 929
Query: 815 AHSIVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWK 874
+ I + R +++H+A+ + EP+T+ EA D WREAM AEI++L N T+
Sbjct: 930 LYPIEKYVDCHRFTSSHKAFLAAVTAGMEPTTYNEAMVDKAWREAMSAEIESLRVNQTFS 989
Query: 875 LVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTV 934
+V+LP G + +GNKWVY++K DG + RYKARLV G Q EG+DY +TF+PVAK++TV
Sbjct: 990 IVNLPPGKRALGNKWVYKIKYRSDGAIERYKARLVVLGNCQKEGVDYDETFAPVAKMSTV 1049
Query: 935 RVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPSVKPNQVCKLLKSLYGLKQ 994
R+ L +AA+++WH+HQ+DV NAFLHG+L E+VYM +P G P++VC+L KSLYGLKQ
Sbjct: 1050 RLFLGVAAARDWHVHQMDVHNAFLHGDLKEEVYMKLPQGFQCDDPSKVCRLHKSLYGLKQ 1109
Query: 995 ASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVK 1054
A R W+ KLS+ L+ GF Q+ SD+SLF F +LVYVDD+I+ G+ K
Sbjct: 1110 APRCWFSKLSSALKQYGFTQSLSDYSLFSYNNDGIFVHVLVYVDDLIISGSCPDAVAQFK 1169
Query: 1055 DSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPS 1114
L F +KDLG+LKYFLG+EV+ + G L QRKY LD++ E G LG++P A PL+ +
Sbjct: 1170 SYLESCFHMKDLGLLKYFLGIEVSRNAQGFYLSQRKYVLDIISEMGLLGARPSAFPLEQN 1229
Query: 1115 IRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVL 1174
+LS D + YRRLVGRL+YL TRP++S++ L+Q+M NP H+ AA RV+
Sbjct: 1230 HKLSLSTSPLLSDSSRYRRLVGRLIYLVVTRPELSYSVHTLAQFMQNPRQDHWNAAIRVV 1289
Query: 1175 RYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTT 1234
RYLK++PG G+L ST+ I G+ D+D+A CP TRRS++GY +G + +SWK KKQ T
Sbjct: 1290 RYLKSNPGQGILLSSTSTLQINGWCDSDYAACPLTRRSLTGYFVQLGDTPISWKTKKQPT 1349
Query: 1235 VSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERT 1294
VSRSS EAEYRA+A+ T EL WL +L DL V+ +F D++SA+ ++ NPV HERT
Sbjct: 1350 VSRSSAEAEYRAMAFLTQELMWLKRVLYDLGVSHVQAMRIFSDSKSAIALSVNPVQHERT 1409
Query: 1295 KHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
KH+++DCH +R+ + GI+ +P+ Q+AD+ TKAL + + F KL +
Sbjct: 1410 KHVEVDCHFIRDAILDGIIATSFVPSHKQLADILTKALGEKEVRYFLRKLGI 1461
>UniRef100_Q9SIM3 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1461
Score = 795 bits (2053), Expect = 0.0
Identities = 423/937 (45%), Positives = 582/937 (61%), Gaps = 57/937 (6%)
Query: 417 ALWHFRLGHLSHDRLLALHTVQNSI--SVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFD 474
++WH RLGH S RL +L V + KS C VCHLAKQK+ F + + F+
Sbjct: 567 SVWHKRLGHPSFSRLDSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFE 626
Query: 475 LVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQ 534
L+H+D+WGP + +V +KYFLT++DD+SR W+ LL +K +V FI LV+ Q+
Sbjct: 627 LLHIDVWGPFSVETVEGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDT 686
Query: 535 IVKAIRSDNGPEFLLPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKL 594
VK++RSDN E FY A+GIV SC TP+QN VERKHQHILN+ARAL+FQS +
Sbjct: 687 RVKSVRSDNAKELAFTEFYKAKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNM 746
Query: 595 PKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRS 654
W VL AVFL+NR PS LL NK+P+E+L + D +K FG LC+++T + R
Sbjct: 747 SLPYWGDCVLTAVFLINRTPSALLSNKTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRH 806
Query: 655 KLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQ 714
K PR+R C+FLGY G KGY L+DL + + ISR+V FHE + P
Sbjct: 807 KFLPRSRACVFLGYPFGFKGYKLLDLESNVVHISRNVEFHEELFP--------------- 851
Query: 715 TQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPNTQTVDEIP 774
+AS QSA + V TP + SG SI +P+ Q
Sbjct: 852 -----------LASSQQSATTASDVFTP--------MDPLSSGNSITSHLPSPQ------ 886
Query: 775 QPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIVHHYSDERLSTAHRAY 834
I+ ++ RR T K P HL ++ + +H I S ++S +H Y
Sbjct: 887 ----ISPSTQISKRRIT---KFPAHLQDYH--CYFVNKDDSHPISSSLSYSQISPSHMLY 937
Query: 835 ALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVK 894
NI++ P ++ EA W A+ EI A+E+ TW++ LP G K +G KWV+ VK
Sbjct: 938 INNISKIPIPQSYHEAKDSKEWCGAIDQEIGAMERTDTWEITSLPPGKKAVGCKWVFTVK 997
Query: 895 RNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVD 954
+ DG+L R+KAR+VAKGY Q EGLDY +TFSPVAK+ TV+++L ++AS+ W+L+QLD+
Sbjct: 998 FHADGSLERFKARIVAKGYTQKEGLDYTETFSPVAKMATVKLLLKVSASKKWYLNQLDIS 1057
Query: 955 NAFLHGNLDEDVYMTIPAGVPSVK-----PNQVCKLLKSLYGLKQASRKWYEKLSAHLET 1009
NAFL+G+L+E +YM +P G +K PN VC+L KS+YGLKQASR+W+ K S L
Sbjct: 1058 NAFLNGDLEETIYMKLPDGYADIKGTSLPPNVVCRLKKSIYGLKQASRQWFLKFSNSLLA 1117
Query: 1010 LGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVL 1069
LGF++ DH+LFV+ GS F LLVYVDD+++ T Q + ++L +F +++LG L
Sbjct: 1118 LGFEKQHGDHTLFVRCIGSEFIVLLVYVDDIVIASTTEQAAQSLTEALKASFKLRELGPL 1177
Query: 1070 KYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPA 1129
KYFLGLEVA ++ GISL QRKY L+LL L KP + P+ P+IRLS+ G +D
Sbjct: 1178 KYFLGLEVARTSEGISLSQRKYALELLTSADMLDCKPSSIPMTPNIRLSKNDGLLLEDKE 1237
Query: 1130 AYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPR 1189
YRRLVG+L+YLT TRPDI+ A +L Q+ S P H A +VL+Y+K + G GL +
Sbjct: 1238 MYRRLVGKLMYLTITRPDITFAVNKLCQFSSAPRTAHLAAVYKVLQYIKGTVGQGLFYSA 1297
Query: 1190 NSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAY 1249
+ ++GY+DADW CPD+RRS +G+ ++G SL+SW++KKQ TVSRSS EAEYRALA
Sbjct: 1298 EDDLTLKGYTDADWGTCPDSRRSTTGFTMFVGSSLISWRSKKQPTVSRSSAEAEYRALAL 1357
Query: 1250 ATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQ 1309
A+CE+ WL LL L+V + P+L+ D+ +A++IA NPVFHERTKH++IDCH VREKL
Sbjct: 1358 ASCEMAWLSTLLLALRVH-SGVPILYSDSTAAVYIATNPVFHERTKHIEIDCHTVREKLD 1416
Query: 1310 AGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
G LKLL + T QVAD+ TK L P F +K+++
Sbjct: 1417 NGQLKLLHVKTKDQVADILTKPLFPYQFAHLLSKMSI 1453
>UniRef100_O04543 F20P5.25 protein [Arabidopsis thaliana]
Length = 1315
Score = 787 bits (2033), Expect = 0.0
Identities = 406/937 (43%), Positives = 580/937 (61%), Gaps = 42/937 (4%)
Query: 418 LWHFRLGHLSHDRLLALHTVQNSISVSKS--IVCDVCHLAKQKRKMFTVSVSKAQKCFDL 475
LWH RLGH S +L + ++ + + C VCH++KQK F +K+ + FDL
Sbjct: 406 LWHKRLGHPSVQKLQPMSSLLSFPKQKNNTDFHCRVCHISKQKHLPFVSHNNKSSRPFDL 465
Query: 476 VHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQI 535
+H+D WGP + + ++YFLT++DDYSR WV LL NK +V + F+T+V+ QF
Sbjct: 466 IHIDTWGPFSVQTHDGYRYFLTIVDDYSRATWVYLLRNKSDVLTVIPTFVTMVENQFETT 525
Query: 536 VKAIRSDNGPEFLLPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLP 595
+K +RSDN PE FY ++GIV SC TPQQN VERKHQHILN+AR+L FQS +P
Sbjct: 526 IKGVRSDNAPELNFTQFYHSKGIVPYHSCPETPQQNSVVERKHQHILNVARSLFFQSHIP 585
Query: 596 KKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSK 655
W +L AV+L+NR+P+ +L++K P+E+L + + +KVFG LC+A+T +R K
Sbjct: 586 ISYWGDCILTAVYLINRLPAPILEDKCPFEVLTKTVPTYDHIKVFGCLCYASTSPKDRHK 645
Query: 656 LDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQT 715
PRA+ C F+GY G KGY L+DL T I +SR V+FHE + P+ GS DL
Sbjct: 646 FSPRAKACAFIGYPSGFKGYKLLDLETHSIIVSRHVVFHEELFPFLGS--------DLSQ 697
Query: 716 QAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEI-PNTQTVDEIP 774
+ ++ LN +Q D H PS +S EI P+ + +P
Sbjct: 698 EEQNFFPDLNPTPPMQRQSSD---------------HVNPSDSSSSVEILPSANPTNNVP 742
Query: 775 QPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIVHHYSDERLSTAHRAY 834
+P V+ S R K+P +L ++ + + SST H I S +R++ + +
Sbjct: 743 EPS---------VQTSHRKAKKPAYLQDYYCHS--VVSSTPHEIRKFLSYDRINDPYLTF 791
Query: 835 ALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVK 894
+ + +EPS + EA K WR+AM AE LE TW++ LP + IG +W++++K
Sbjct: 792 LACLDKTKEPSNYTEAEKLQVWRDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRWIFKIK 851
Query: 895 RNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVD 954
N DG++ RYKARLVA+GY Q EG+DY +TFSPVAKL +V+++L +AA L QLD+
Sbjct: 852 YNSDGSVERYKARLVAQGYTQKEGIDYNETFSPVAKLNSVKLLLGVAARFKLSLTQLDIS 911
Query: 955 NAFLHGNLDEDVYMTIPAGVPS-----VKPNQVCKLLKSLYGLKQASRKWYEKLSAHLET 1009
NAFL+G+LDE++YM +P G S + PN VC+L KSLYGLKQASR+WY K S+ L
Sbjct: 912 NAFLNGDLDEEIYMRLPQGYASRQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFSSTLLG 971
Query: 1010 LGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVL 1069
LGF Q+ DH+ F+K F +LVY+DD+I+ N ++K + F ++DLG L
Sbjct: 972 LGFIQSYCDHTCFLKISDGIFLCVLVYIDDIIIASNNDAAVDILKSQMKSFFKLRDLGEL 1031
Query: 1070 KYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPA 1129
KYFLGLE+ S GI + QRKY LDLL ETG LG KP + P+DPS+ + + G + +
Sbjct: 1032 KYFLGLEIVRSDKGIHISQRKYALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGDFVEVG 1091
Query: 1130 AYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPR 1189
YRRL+GRL+YL TRPDI+ A +L+Q+ P H +A ++L+Y+K + G GL +
Sbjct: 1092 PYRRLIGRLMYLNITRPDITFAVNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQGLFYSA 1151
Query: 1190 NSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAY 1249
S + ++ Y++AD+ C D+RRS SGYC ++G SL+ WK++KQ VS+SS EAEYR+L+
Sbjct: 1152 TSELQLKVYANADYNSCRDSRRSTSGYCMFLGDSLICWKSRKQDVVSKSSAEAEYRSLSV 1211
Query: 1250 ATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQ 1309
AT EL WL L++L+V + +LFCDN++A+HIA N VFHERTKH++ DCH VRE+L
Sbjct: 1212 ATDELVWLTNFLKELQVPLSKPTLLFCDNEAAIHIANNHVFHERTKHIESDCHSVRERLL 1271
Query: 1310 AGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
G+ +L I T LQ+AD FTK L P F +K+ +
Sbjct: 1272 KGLFELYHINTELQIADPFTKPLYPSHFHRLISKMGL 1308
>UniRef100_Q9SLF0 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1333
Score = 748 bits (1931), Expect = 0.0
Identities = 411/955 (43%), Positives = 582/955 (60%), Gaps = 45/955 (4%)
Query: 418 LWHFRLGHLSHDRLLALHTVQNSISVSKSIV---CDVCHLAKQKRKMFTVSVSKAQKCFD 474
LWH R+GH + R+++L ++S+SVS + + CDVCH AKQ R F +S++K + F+
Sbjct: 391 LWHSRMGHPAA-RVVSL-IPESSVSVSSTHLNKACDVCHRAKQTRNSFPLSINKTLRIFE 448
Query: 475 LVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQ 534
L++ D+WGP S +YFLT++DDYSR VW+ LLN+K E +KNF + QF
Sbjct: 449 LIYCDLWGPYRTPSHTGARYFLTIIDDYSRGVWLYLLNDKSEAPCHLKNFFAMTDRQFNV 508
Query: 535 IVKAIRSDNGPEFL-LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSK 593
+K +RSDNG EFL L F+ QG++H++SCV+TP++N RVERKH+H+LN+ARAL FQ+
Sbjct: 509 KIKTVRSDNGTEFLCLTKFFQEQGVIHERSCVATPERNDRVERKHRHLLNVARALRFQAN 568
Query: 594 LPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNR 653
LP + W VL A +L+NR PS +L + +PYE L+++ + ++VFGSLC+A
Sbjct: 569 LPIQFWGECVLTAAYLINRTPSSVLNDSTPYERLHKKQPRFDHLRVFGSLCYAHNRNRGG 628
Query: 654 SKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDL 713
K R+R+C+F+GY G KG+ L DL E F+SRDV+F E P++ S+ ++
Sbjct: 629 DKFAERSRRCVFVGYPHGQKGWRLFDLEQNEFFVSRDVVFSELEFPFRISHEQ--NVIEE 686
Query: 714 QTQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPNTQTVDE- 772
+ +A A ++ E + G + TP + + PS S E + +D
Sbjct: 687 EEEA-LWAPIVDGLIEEEVHLGQNAGPTPPICVSSPIS---PSATSSRSEHSTSSPLDTE 742
Query: 773 -IPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALP-IRSSTAHSIVHHY--SDERLS 828
+P P + ST P N QF L + +TA ++ R S
Sbjct: 743 VVPTPAT-----------STTSASSPSSPTNLQFLPLSRAKPTTAQAVAPPAVPPPRRQS 791
Query: 829 TAHRA-------YALNIAQDQE-PSTFAEANKDLHWREAMQ---------AEIKALEKNG 871
T ++A + +N QE PS L R+ + I A E+N
Sbjct: 792 TRNKAPPVTLKDFVVNTTVCQESPSKLNSILYQLQKRDDTRRFSASHTTYVAIDAQEENH 851
Query: 872 TWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKL 931
TW + DLP G + IG++WVY+VK N DG++ RYKARLVA G Q EG DY +TF+PVAK+
Sbjct: 852 TWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARLVALGNKQKEGEDYGETFAPVAKM 911
Query: 932 TTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPSVKPNQVCKLLKSLYG 991
TVR+ L +A +NW +HQ+DV NAFLHG+L E+VYM +P G + PN+VC+L K+LYG
Sbjct: 912 ATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYMKLPPGFEASHPNKVCRLRKALYG 971
Query: 992 LKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQ 1051
LKQA R W+EKL+ L+ GF+Q+ +D+SLF +GS +L+YVDD+I+ GN+ Q
Sbjct: 972 LKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGSVRIKILIYVDDLIITGNSQRATQ 1031
Query: 1052 LVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPL 1111
K+ L F +KDLG LKYFLG+EVA ST+GI +CQRKY LD++ ETG LG KP PL
Sbjct: 1032 QFKEYLASCFHMKDLGPLKYFLGIEVARSTTGIYICQRKYALDIISETGLLGVKPANFPL 1091
Query: 1112 DPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQ 1171
+ + +L DP YRRLVGRL+YL TR D++ + L+++M P + H+ AA
Sbjct: 1092 EQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTRLDLAFSVHILARFMQEPREDHWAAAL 1151
Query: 1172 RVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKK 1231
RV+RYLKA PG G+ R+ I G+ D+DWAG P +RRS++GY G S +SWK KK
Sbjct: 1152 RVVRYLKADPGQGVFLRRSGDFQITGWCDSDWAGDPMSRRSVTGYFVQFGDSPISWKTKK 1211
Query: 1232 QTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFH 1291
Q TVS+SS EAEYRA+++ EL WL LL L V+ ++ CD++SA++IA NPVFH
Sbjct: 1212 QDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLGVSHVQPMIMCCDSKSAIYIATNPVFH 1271
Query: 1292 ERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
ERTKH++ID H VR++ G++ + TT Q+AD+FTK L F F KL +
Sbjct: 1272 ERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLADIFTKPLGRDCFSAFRIKLGI 1326
>UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Solanum tuberosum]
Length = 1476
Score = 747 bits (1929), Expect = 0.0
Identities = 408/950 (42%), Positives = 572/950 (59%), Gaps = 71/950 (7%)
Query: 418 LWHFRLGHLSHDRLLALHTVQNSISVSKSIV--CDVCHLAKQKRKMFTVSVSKAQKCFDL 475
+WH RLGH+ + L ++ S K ++ CDVC LA+Q R F +S S+++ CFDL
Sbjct: 552 MWHKRLGHIP---MSVLRKIKMFDSPQKLVLPSCDVCPLARQVRLPFPISQSRSENCFDL 608
Query: 476 VHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQI 535
+H+D+WGP A+ + +YFLTV+DD+SR+ W+ L++ K +V ++NFI ++ TQFGQ
Sbjct: 609 IHLDVWGPYKAATHNKMRYFLTVVDDHSRWTWIFLMHLKSDVSTVLQNFILMIDTQFGQK 668
Query: 536 VKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQS 592
+K RSDNG EF + + GIVHQ SC TPQQNG VER+H+HIL ARAL FQ
Sbjct: 669 IKIFRSDNGTEFFNAQCDGLFKSHGIVHQSSCPHTPQQNGVVERRHKHILETARALRFQG 728
Query: 593 KLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNN 652
LP + W VL AV ++NRIPS +L NKSP+EL+Y+ + DL M+V G LC AT L N
Sbjct: 729 HLPIRFWGECVLSAVHIINRIPSSVLHNKSPFELMYKRSPDLSYMRVIGCLCHATNLVNT 788
Query: 653 RSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLD 712
++ KGY L DL Q F+SRD++F+E V P+
Sbjct: 789 STQ-----------------KGYKLYDLEHQHFFVSRDMVFNEAVFPF------------ 819
Query: 713 LQTQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPNTQTVDE 772
Q+ + AD + + S PC+S + P+ + E+ IP
Sbjct: 820 ---QSPALADPHDTPVFL--------ASPPCSSHTEDADAVQPAIITSEEIIP------- 861
Query: 773 IPQPGSITAEGEV----EVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIVHHYSDERLS 828
+ P S ++ + E RRS R K P+ +F ++ + + I + LS
Sbjct: 862 VASPPSAVSDDHLHPPPERRRSYRTGKPPIWQKDFITTSTSRSNHCLYPISDNIDYSCLS 921
Query: 829 TAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNK 888
+ ++ Y + + + EP + +A D W AM+ EI+ALE N TW++V LP+G K IG K
Sbjct: 922 STYQCYIASSSVETEPQFYYQAANDCRWVHAMKEEIQALEDNKTWEVVSLPKGKKAIGCK 981
Query: 889 WVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHL 948
WVY++K G + R+KARLVAKGYNQ EGLDY +TFSPV K+ T+R +L LA S+ W +
Sbjct: 982 WVYKIKYKASGEIERFKARLVAKGYNQKEGLDYQETFSPVVKMVTLRTVLTLAVSKGWDI 1041
Query: 949 HQLDVDNAFLHGNLDEDVYMTIPAGVPSVKPN--QVCKLLKSLYGLKQASRKWYEKLSAH 1006
Q+DV NAFL G+L E+VYM +P G K +VC+LLKSLYGLKQASR+W KL+
Sbjct: 1042 QQMDVYNAFLQGDLIEEVYMQLPQGFQYDKTGDPKVCRLLKSLYGLKQASRQWNVKLTTA 1101
Query: 1007 LETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDL 1066
L GF+Q+ D+SL +K +L+YVDD+++ G+++ K L F IKDL
Sbjct: 1102 LLAAGFQQSHLDYSLMLKRTADGIVIVLIYVDDLLITGSSLQLIDDAKQVLKANFKIKDL 1161
Query: 1067 GVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPY- 1125
G L+YFLG+E A + SG+ + QRKY L+L+ + G GSKP TP++ ++L+ + +
Sbjct: 1162 GTLRYFLGMEFARNASGMLMHQRKYALELISDLGLGGSKPSVTPVELHLKLTTREFDLHV 1221
Query: 1126 ---------DDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRY 1176
DP Y+RLVGRLLYLT TRPDIS A Q LSQ+M P H +AA RV++Y
Sbjct: 1222 GSSGADSLLADPTEYQRLVGRLLYLTITRPDISFAVQHLSQFMHAPKVSHMEAAIRVVKY 1281
Query: 1177 LKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVS 1236
+K +PGLGL + +Q Y DADW C +TR+SI+GY G +L+SWK+KKQ T+S
Sbjct: 1282 VKQAPGLGLYMAVQTADTLQAYCDADWGSCINTRKSITGYMIQFGSALLSWKSKKQPTIS 1341
Query: 1237 RSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKH 1296
RSS EAEYR+LA EL WL L ++L + + L+CD+++A+ IAANPVFHERTKH
Sbjct: 1342 RSSAEAEYRSLASTVAELVWLTGLFKELDMPLSLPVSLYCDSKAAIQIAANPVFHERTKH 1401
Query: 1297 LDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
+DIDCH +REK+QAG++ + +PT Q AD+ TK L +KL +
Sbjct: 1402 IDIDCHFIREKVQAGLVMIHYLPTQEQPADILTKGLSSAQHSYLVSKLGL 1451
>UniRef100_Q8W153 Polyprotein [Oryza sativa]
Length = 1472
Score = 720 bits (1858), Expect = 0.0
Identities = 412/1032 (39%), Positives = 599/1032 (57%), Gaps = 52/1032 (5%)
Query: 354 RLLLAKEMLQDMEVQTQRRIGLGSLQPDELYHLVVSSSSSTCFSIQHSSTSEINDFGHII 413
R+ L +E E +T +++G+G ++ D L++L ++ ++ S++ E+ +
Sbjct: 447 RVSLDRENCLIQERRTGKKLGIG-IRRDGLWYLDRRGTNEDVCALMASTSKEVTEV---- 501
Query: 414 PPGALWHFRLGHLSHDRLLALHTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCF 473
L H RLGH+S + + + V+ S ++CD C K R + ++ F
Sbjct: 502 ---LLLHCRLGHISFEIMSKMFPVEFSKVDKHMLICDACEYGKHTRTSYVSRGLRSILPF 558
Query: 474 DLVHMDIW-GPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQF 532
L+H D+W P+ S+ KYF+T +D YSR W+ L+ +K EV + +NF +K F
Sbjct: 559 MLIHSDVWTSPVV--SMSGMKYFVTFIDCYSRMTWLYLMRHKDEVLKCFQNFYAYIKNHF 616
Query: 533 GQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALL 589
V+ IR+DNG E++ F S +GI+HQ SC TP QNG ERK++H+L IAR+L+
Sbjct: 617 NARVQFIRTDNGGEYMNSEFGHFLSLEGILHQTSCPDTPPQNGVAERKNRHLLEIARSLM 676
Query: 590 FQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTL 649
+ +PK +W +V+ A +L+NR PS++L K+PYE+++ + + +VFG CF
Sbjct: 677 YTMNVPKFLWSEAVMTAAYLINRTPSRILGMKTPYEMIFGKNEFVVPPRVFGCTCFVRDH 736
Query: 650 TNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSN---SP 706
+ KLDPRA KCIF+GY KGY + F+S DV F E V P+ G S
Sbjct: 737 RPSIGKLDPRAVKCIFIGYSSSQKGYKCWSPSERRTFVSMDVTFRESV-PFYGEKTDISS 795
Query: 707 AWTCLDLQTQAESTADK---LNVASEIQSAKGDASVST-PCASIEDGVQHEMPSGASIED 762
+ LD T+ + K + E + +KG V PCA I D VQ + E+
Sbjct: 796 LFVDLDDLTRGDHDQQKEGEILGLKENEQSKGKIVVGEIPCA-IGDPVQEQEWRKPHEEE 854
Query: 763 EI---------PNTQTV---DEIP------QPGSITAEGEVEVRRSTRPLKRPVHLANFQ 804
+ P TQ V D++ Q S + ++E+R L +
Sbjct: 855 NLQVYTRRMRLPTTQQVEVDDQVSDDLTHVQVSSESGGEQIEIREEESNLPIAIRKGMRS 914
Query: 805 FSALPIR---------SSTAHSIVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLH 855
+ P + S + I ++ S LS+ +RA+ ++ P + EA +D
Sbjct: 915 NAGKPPQRYGFEIGDESGDENDIANYVSYTSLSSTYRAFVASLNSAIIPKDWKEAKQDPR 974
Query: 856 WREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQ 915
W +AM E++ALEKN TW LV P G K + KWVY VK+N DG + RYKARLVAKGY+Q
Sbjct: 975 WHQAMLDELEALEKNKTWDLVSYPNGKKVVNCKWVYAVKQNPDGKVERYKARLVAKGYSQ 1034
Query: 916 VEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVP 975
G+DY +TF+PVAK++TVR I++ A + +W LHQLDV NAFLHG+L E+VYM IP G
Sbjct: 1035 TYGIDYDETFAPVAKMSTVRTIISCAVNFDWPLHQLDVKNAFLHGDLQEEVYMEIPPGFA 1094
Query: 976 SVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLL 1034
+++ +V +L KSLYGLKQ+ R W+++ + +G+KQ DH++F G T L
Sbjct: 1095 TLQTKGKVLRLKKSLYGLKQSPRAWFDRFRRAMCAMGYKQCNGDHTVFYHHSGDHITILA 1154
Query: 1035 VYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLD 1094
VYVDD+I+ GN +E +K +L + F +KDLG LKYFLG+E+A S GI L QRKY LD
Sbjct: 1155 VYVDDMIITGNDCSEITRLKQNLSKEFEVKDLGQLKYFLGIEIARSPRGIVLSQRKYALD 1214
Query: 1095 LLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQ 1154
LL +TG LG +P +TP+D + +L E G P + Y+RLVGRL+YL TRPDI++A
Sbjct: 1215 LLSDTGMLGCRPASTPVDQNHKLCAESGNPVNKER-YQRLVGRLIYLCHTRPDITYAVSM 1273
Query: 1155 LSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSIS 1214
+S+YM +P GH A R+LRYLK SPG GL F +N + ++GY DA WA CPD RRS S
Sbjct: 1274 VSRYMHDPRSGHMDAVYRILRYLKGSPGKGLWFKKNGHLEVEGYCDAHWASCPDDRRSTS 1333
Query: 1215 GYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVL 1274
GYC ++G +LVSW++KKQ VSRS+ EAEYRA++ + EL WL LL +L + L
Sbjct: 1334 GYCVFVGGNLVSWRSKKQPVVSRSTAEAEYRAMSVSLSELLWLRNLLSELMLPVDTPMKL 1393
Query: 1275 FCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQP 1334
+CDN+SA+ IA NPV H+RTKH+++D ++EKL G+L+L + + QVAD FTK L
Sbjct: 1394 WCDNKSAISIANNPVQHDRTKHVELDRFFIKEKLDEGVLELEFVMSGGQVADCFTKGLGV 1453
Query: 1335 RVFQGFATKLAM 1346
+ K+ M
Sbjct: 1454 KECNSSCDKMGM 1465
>UniRef100_Q9FWZ5 Putative retroelement polyprotein [Arabidopsis thaliana]
Length = 1404
Score = 693 bits (1789), Expect = 0.0
Identities = 405/1013 (39%), Positives = 586/1013 (56%), Gaps = 62/1013 (6%)
Query: 366 EVQTQRRIGLGSLQPDELYHLV-VSSSSSTCFSIQHSSTSEINDFGHIIPPGALWHFRLG 424
+++T + IG G + ELY L +S +SS+CFS + N LWH RLG
Sbjct: 416 DIETGKVIGEGGSK-GELYVLEDLSPNSSSCFSSKSHLGISFN---------TLWHARLG 465
Query: 425 HLSHDRLLALHTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLVHMDIWGPL 484
H H R L L S + C+ C L K + +F S++ +KCFDLVH D+W
Sbjct: 466 H-PHTRALKLMLPNISFDHTS---CEACILGKHCKSVFPKSLTIYEKCFDLVHSDVWTSP 521
Query: 485 AQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIVKAIRSDNG 544
S N+KYF+T +++ S++ W+ LL +K V + NF T V QF +K R+DNG
Sbjct: 522 C-VSRDNNKYFVTFINEKSKYTWITLLPSKDRVFEAFTNFETYVTNQFNAKIKVFRTDNG 580
Query: 545 PEFLLPAF---YSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKKMWCY 601
E+ F + +GI+HQ SC TPQQNG ERK++H++ +AR+++F + +PK+ W
Sbjct: 581 GEYTSQKFRDHLAKRGIIHQTSCPYTPQQNGVAERKNRHLMEVARSMMFHTSVPKRFWGD 640
Query: 602 SVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLDPRAR 661
+VL A +L+NR P+K+L + SP+E+L ++ ++VFG +CF RSKLD ++
Sbjct: 641 AVLTACYLINRTPTKVLSDLSPFEVLNNTKPFIDHLRVFGCVCFVLIPGEQRSKLDAKST 700
Query: 662 KCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQTQAESTA 721
KC+FLGY KGY D FISRDV F E+ +N W +L+ ST+
Sbjct: 701 KCMFLGYSTTQKGYKCFDPTKNRTFISRDVKFLEN---QDYNNKKDWE--NLKDLTHSTS 755
Query: 722 DKLNVASEIQSAKGDASVSTPC--------------ASIEDGVQHEMPSGASIEDEIPNT 767
D++ + G+ S ST + E +QH+ + ++++ PNT
Sbjct: 756 DRVETLKFLLDHLGNDSTSTTQHQPEMTQDQEDLNQENEEVSLQHQ-ENLTHVQEDPPNT 814
Query: 768 QT----VDEIPQPGSITAEGEV------EVRRSTRPLKRPVHLANFQFSALPIRSSTAHS 817
Q V EI S E +RRSTR ++R N A P +++ + +
Sbjct: 815 QEHSEHVQEIQDDSSEDEEPTQVLPPPPPLRRSTR-IRRKKEFFNSNAVAHPFQATCSLA 873
Query: 818 IVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVD 877
+V H+A+ I++ P T+ EA + WR+A+ EI A+++N TW D
Sbjct: 874 LV--------PLDHQAFLSKISEHWIPQTYEEAMEVKEWRDAIADEINAMKRNHTWDEDD 925
Query: 878 LPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVI 937
LP+G K + ++WV+ +K +G + RYK RLVA+G+ Q G DY +TF+PVAKL TVRV+
Sbjct: 926 LPKGKKTVSSRWVFTIKYKSNGDIERYKTRLVARGFTQTYGSDYMETFAPVAKLHTVRVV 985
Query: 938 LALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPSVKP-NQVCKLLKSLYGLKQAS 996
LALA + +W L Q+DV NAFL G L++DVYMT P G+ P ++V +L K++YGLKQ+
Sbjct: 986 LALATNLSWGLWQMDVKNAFLQGELEDDVYMTPPPGLEDTIPCDKVLRLRKAIYGLKQSP 1045
Query: 997 RKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDS 1056
R WY KLS L+ GFK++ SDH+LF +L+YVDD+I+ G+ K
Sbjct: 1046 RAWYHKLSRTLKDHGFKKSESDHTLFTLQSPQGIVVVLIYVDDLIITGDNKDGIDSTKTF 1105
Query: 1057 LHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIR 1116
L F IKDLG LKYFLG+EV S +G+ L QRKY LDLL ETG + +KP TPL+ +
Sbjct: 1106 LKSCFDIKDLGELKYFLGIEVCRSNAGLFLSQRKYTLDLLNETGFMDAKPARTPLEDGYK 1165
Query: 1117 LS---QEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRV 1173
++ +++ + + D YR+LVG+L+YLT TRPDI A Q+SQ+M PM H+ +R+
Sbjct: 1166 VNRKGEKEDEKFGDAPLYRKLVGKLIYLTNTRPDICFAVNQVSQHMKVPMVYHWNMVERI 1225
Query: 1174 LRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQT 1233
LRYLK S G G+ +NS+ I GY DAD+AG RRS +GYC +IG +L +WK KKQ
Sbjct: 1226 LRYLKGSSGQGIWMGKNSSTEIVGYCDADYAGDRGDRRSKTGYCTFIGGNLATWKTKKQK 1285
Query: 1234 TVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHER 1293
VS SS E+EYRA+ T EL WL LL+DL + + CDN++A++IA+N VFHER
Sbjct: 1286 VVSCSSAESEYRAMRKLTNELTWLKALLKDLGIEQHMPITMHCDNKAAIYIASNSVFHER 1345
Query: 1294 TKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
TKH+++DCH VREK+ G+ + Q+AD+FTKA +V KL +
Sbjct: 1346 TKHIEVDCHKVREKIIEGVTLPCYTRSEDQLADIFTKAASLKVCNFIHGKLGL 1398
>UniRef100_Q8S1E5 Putative gag/pol polyprotein [Oryza sativa]
Length = 1090
Score = 689 bits (1779), Expect = 0.0
Identities = 403/1011 (39%), Positives = 558/1011 (54%), Gaps = 59/1011 (5%)
Query: 366 EVQTQRRIGLGSLQPDELYHLVVSSSSSTCFSIQHSSTSEINDFGHIIPPGALWHFRLGH 425
++QT RR+ L ELY L ++ SS + +S++ LWH RLGH
Sbjct: 86 DLQT-RRVILRCNSRGELYTLPAATPSSAAHGLLATSST-------------LWHCRLGH 131
Query: 426 LSHDRLLALHTVQNSISVS----KSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLVHMDIW 481
A+H ++N S+S + +C C L K R F S S+ F+LVH D+W
Sbjct: 132 PGP---AAIHGLRNIASISCNKIDTSLCHACQLGKHTRLPFHNSSSRTSVPFELVHCDVW 188
Query: 482 -GPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIVKAIR 540
P+ S KY+L VLDD+S F W LL K +V + + F+ V TQFG +K+ +
Sbjct: 189 TSPVMSTS--GFKYYLVVLDDFSHFCWTFLLRLKSDVHRHIVEFVEYVSTQFGLPLKSFQ 246
Query: 541 SDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKK 597
+DNG EF+ + F +++G + SC T QNG+ ER + I N R LL Q+ +P
Sbjct: 247 ADNGREFVNTAITTFLASRGTQLRLSCPYTSPQNGKAERMLRTINNSIRTLLIQASMPPS 306
Query: 598 MWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLD 657
W ++ A +L+NR PS + P++LL+R D ++VFG LC+ KL
Sbjct: 307 YWAEALATATYLLNRRPSSSIHQSLPFQLLHRTIPDFSHLRVFGCLCYPNLSATTPHKLS 366
Query: 658 PRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQTQA 717
PR+ C+FLGY KGY +DL T I ISR V+F E P+ + PA + D Q
Sbjct: 367 PRSTACVFLGYPTSHKGYRCLDLSTHRIIISRHVVFDESQFPF-AATPPAASSFDFLLQG 425
Query: 718 ESTADKLNVASEIQSAKGDASVST---------PCASIEDGVQHEMPSGASIEDEIPNTQ 768
S AD ++ E Q + ST P + G S + T
Sbjct: 426 LSPADAPSLEVE-QPRPLTVAPSTEVEQPYLPLPSRRLSAGTVTVASEAPSAGAPLVGTS 484
Query: 769 TVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTA----------HSI 818
+ D P PGS T + +RPV + +P SSTA HS+
Sbjct: 485 SADATP-PGSATRASTIVSPFRHVYTRRPV-------TTVPPSSSTAVTNAVAAPQPHSM 536
Query: 819 VHHYSDERLSTAHRAY--ALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLV 876
V L R A A P+ + A D +WR AM E K L NGTW+LV
Sbjct: 537 VTRSQSGSLRPVDRLTYTATQAAASPVPANYHSALADPNWRAAMADEYKELVDNGTWRLV 596
Query: 877 DLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRV 936
P KW+++ K + DG+LARYKAR V +GY+Q G+DY +TFSPV KL T+RV
Sbjct: 597 SRPPRANIATGKWIFKHKFHSDGSLARYKARWVVRGYSQQHGIDYDETFSPVVKLATIRV 656
Query: 937 ILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQA 995
+L++AAS+ W +HQLDV NAFLHG+L E VY P+G V P+ VC L KSLYGLKQA
Sbjct: 657 VLSIAASRAWPIHQLDVKNAFLHGHLKETVYCQQPSGFVDPTAPDAVCLLQKSLYGLKQA 716
Query: 996 SRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKD 1055
R WY++ + ++ +GF +ASD SLFV G LL+YVDD+IL +T T Q +
Sbjct: 717 PRAWYQRFATYIRQMGFMPSASDTSLFVYKDGDRIAYLLLYVDDIILTASTTTLLQQLTA 776
Query: 1056 SLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSI 1115
LH F + DLG L +FLG+ V S G+ L QR+Y +DLLQ G +TP+D
Sbjct: 777 RLHSEFAMTDLGDLHFFLGISVKRSPDGLFLSQRQYAVDLLQRAGMAECHSTSTPVDTHA 836
Query: 1116 RLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLR 1175
+LS G P DP+AYR + G L YLT TRPD+++A QQ+ +M +P + H +R+LR
Sbjct: 837 KLSATDGLPVADPSAYRSIAGALQYLTLTRPDLAYAVQQVCLFMHDPREPHLALVKRILR 896
Query: 1176 YLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTV 1235
Y+K S +GL ++ YSDADWAGCP++RRS SGYC Y+G +LVSW +K+QTTV
Sbjct: 897 YVKGSLSIGLHIGSGPIQSLTAYSDADWAGCPNSRRSTSGYCVYLGDNLVSWSSKRQTTV 956
Query: 1236 SRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTK 1295
SRSS EAEYRA+A+A E WL LLQ+L V + +++CDN SA+++ ANPV H RTK
Sbjct: 957 SRSSAEAEYRAVAHAVAECCWLRQLLQELHVPIASATIVYCDNVSAVYMTANPVHHRRTK 1016
Query: 1296 HLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
H++ID H VREK+ G +++L +P++ Q AD+ TK L ++F F + L +
Sbjct: 1017 HIEIDIHFVREKVALGQVRVLYVPSSHQFADIMTKGLPVQLFTDFRSSLCV 1067
>UniRef100_Q9SA17 F28K20.17 protein [Arabidopsis thaliana]
Length = 1415
Score = 635 bits (1638), Expect = e-180
Identities = 355/942 (37%), Positives = 525/942 (55%), Gaps = 48/942 (5%)
Query: 418 LWHFRLGHLSHDRLLALHTVQNS--ISVSKSI---VCDVCHLAKQKRKMFTVSVSKAQKC 472
+WH RLGH + AL +QNS I ++KS VC+ C + K R F +S S+
Sbjct: 455 VWHHRLGHANSK---ALQHLQNSKAIQINKSRTSPVCEPCQMGKSSRLPFLISDSRVLHP 511
Query: 473 FDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQF 532
D +H D+WGP S KY+ +DDYSR+ W L+NK E +F LV+ Q
Sbjct: 512 LDRIHCDLWGPSPVVSNQGLKYYAIFVDDYSRYSWFYPLHNKSEFLSVFISFQKLVENQL 571
Query: 533 GQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALL 589
+K +SD G EF+ L S GI H+ SC TPQQNG ERKH+H++ + ++L
Sbjct: 572 NTKIKVFQSDGGGEFVSNKLKTHLSEHGIHHRISCPYTPQQNGLAERKHRHLVELGLSML 631
Query: 590 FQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTL 649
F S P+K W S A +++NR+PS +LKN SPYE L+ E D ++VFGS C+
Sbjct: 632 FHSHTPQKFWVESFFTANYIINRLPSSVLKNLSPYEALFGEKPDYSSLRVFGSACYPCLR 691
Query: 650 TNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKG---SNSP 706
++K DPR+ +C+FLGY KGY T +++ISR+VIF+E LP+K S P
Sbjct: 692 PLAQNKFDPRSLQCVFLGYNSQYKGYRCFYPPTGKVYISRNVIFNESELPFKEKYQSLVP 751
Query: 707 AWTCLDLQTQAESTADKLNV-ASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIP 765
++ LQ + +++V A+ +Q ++T S V ++ +
Sbjct: 752 QYSTPLLQAWQHNKISEISVPAAPVQLFSKPIDLNTYAGS---QVTEQLTDPEPTSNNEG 808
Query: 766 NTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIVHHYSDE 825
+ + V+ + + + E + T K + N +++ + R +TA
Sbjct: 809 SDEEVNPVAEEIAANQEQVINSHAMTTRSKAGIQKPNTRYALITSRMNTA---------- 858
Query: 826 RLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPI 885
EP T A A K W EA+ EI + TW LV + + +
Sbjct: 859 -----------------EPKTLASAMKHPGWNEAVHEEINRVHMLHTWSLVPPTDDMNIL 901
Query: 886 GNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQN 945
+KWV++ K + DG++ + KARLVAKG++Q EG+DY +TFSPV + T+R++L ++ S+
Sbjct: 902 SSKWVFKTKLHPDGSIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDVSTSKG 961
Query: 946 WHLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQASRKWYEKLS 1004
W + QLDV NAFLHG L E V+M P+G + KP VC+L K++YGLKQA R W++ S
Sbjct: 962 WPIKQLDVSNAFLHGELQEPVFMYQPSGFIDPQKPTHVCRLTKAIYGLKQAPRAWFDTFS 1021
Query: 1005 AHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIK 1064
L GF + SD SLFV Q LL+YVDD++L G+ + + + +L F +K
Sbjct: 1022 NFLLDYGFVCSKSDPSLFVCHQDGKILYLLLYVDDILLTGSDQSLLEDLLQALKNRFSMK 1081
Query: 1065 DLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKP 1124
DLG +YFLG+++ +G+ L Q Y D+LQ+ G P+ TPL +L +
Sbjct: 1082 DLGPPRYFLGIQIEDYANGLFLHQTAYATDILQQAGMSDCNPMPTPLPQ--QLDNLNSEL 1139
Query: 1125 YDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLG 1184
+ +P +R L G+L YLT TRPDI A + Q M +P F +R+LRY+K + G+G
Sbjct: 1140 FAEPTYFRSLAGKLQYLTITRPDIQFAVNFICQRMHSPTTSDFGLLKRILRYIKGTIGMG 1199
Query: 1185 LLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEY 1244
L RNST+ + YSD+D AGC +TRRS +G+C +G +L+SW AK+Q TVS SS EAEY
Sbjct: 1200 LPIKRNSTLTLSAYSDSDHAGCKNTRRSTTGFCILLGSNLISWSAKRQPTVSNSSTEAEY 1259
Query: 1245 RALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVV 1304
RAL YA E+ W+ +LL+DL + ++CDN SA++++ANP H R+KH D D H +
Sbjct: 1260 RALTYAAREITWISFLLRDLGIPQYLPTQVYCDNLSAVYLSANPALHNRSKHFDTDYHYI 1319
Query: 1305 REKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
RE++ G+++ I T Q+ADVFTK+L R F +KL +
Sbjct: 1320 REQVALGLIETQHISATFQLADVFTKSLPRRAFVDLRSKLGV 1361
>UniRef100_Q7X6S0 OSJNBb0011N17.2 protein [Oryza sativa]
Length = 1262
Score = 617 bits (1590), Expect = e-174
Identities = 344/812 (42%), Positives = 486/812 (59%), Gaps = 38/812 (4%)
Query: 570 QNGRVERKHQHILNIARALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYR 629
+NG ERK++H+L IAR+L++ +PK +W +V+ A +L+NR PS++L K+PYE+++
Sbjct: 447 ENGVAERKNRHLLEIARSLMYTMNVPKFLWSEAVMTAAYLINRTPSRILGMKTPYEMIFG 506
Query: 630 EAVDLEMMKVFGSLCFATTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISR 689
+ + +VFG CF + KLDPRA KCIF+GY KGY + F+S
Sbjct: 507 KNEFVVPPRVFGCTCFVRDHRPSIGKLDPRAVKCIFIGYSSSQKGYKCWSPSERRTFVSM 566
Query: 690 DVIFHEHVLPYKGSN---SPAWTCLDLQTQAESTADK---LNVASEIQSAKGDASVST-P 742
DV F E V P+ G S + LD T+ + K + E + +KG V P
Sbjct: 567 DVTFRESV-PFYGEKTDISSLFVDLDDLTRGDHDQQKEGEILGLKENEQSKGKIVVGEIP 625
Query: 743 CASIEDGVQHEMPSGASIEDEI---------PNTQTV---DEIP------QPGSITAEGE 784
CA I D VQ + E+ + P TQ V D++ Q S + +
Sbjct: 626 CA-IGDPVQEQEWRKPHEEENLQVYTRRMRLPTTQQVEVDDQVSDDLTHVQVSSESGGEQ 684
Query: 785 VEVRRSTRPLKRPVHLANFQFSALPIR---------SSTAHSIVHHYSDERLSTAHRAYA 835
+E+R L + + P + S + I ++ S LS+ ++A+
Sbjct: 685 IEIREEESNLPIAIRKGMRSNAGKPPQRYGFEIGDESGDENDIANYVSYTSLSSTYKAFV 744
Query: 836 LNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKR 895
++ P + EA +D W +AM E++ALEKN TW LV P G K + KWVY VK+
Sbjct: 745 ASLNSAIIPKDWKEAKQDPRWHQAMLDELEALEKNKTWDLVSYPNGKKVVNCKWVYAVKQ 804
Query: 896 NVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDN 955
N DG + RYKARLVAKGY+Q G+DY +TF+PVAK++TVR I++ A + +W LHQLDV N
Sbjct: 805 NPDGKVERYKARLVAKGYSQTYGIDYDETFAPVAKMSTVRTIISCAVNFDWPLHQLDVKN 864
Query: 956 AFLHGNLDEDVYMTIPAGVPSVKPN-QVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQ 1014
AFLHG+L E+VYM IP G +++ +V +L KSLYGLKQ+ R W+++ + +G+KQ
Sbjct: 865 AFLHGDLQEEVYMEIPPGFATLQTKGKVLRLKKSLYGLKQSPRAWFDRFRRAMCAMGYKQ 924
Query: 1015 TASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLG 1074
DH++F G T L VYVDD+I+ GN +E +K +L + F +KDLG LKYFLG
Sbjct: 925 CNGDHTVFYHHSGDHITILAVYVDDMIITGNDCSEITRLKQNLSKEFEVKDLGQLKYFLG 984
Query: 1075 LEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRL 1134
+E+A S GI L QRKY LDLL +TG LG +P +TP+D + +L E G P + Y+RL
Sbjct: 985 IEIARSPRGIVLSQRKYALDLLSDTGMLGCRPASTPVDQNHKLCAESGNPVNKER-YQRL 1043
Query: 1135 VGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTIN 1194
VGRL+YL TRPDI++A +S+YM +P GH A R+LRYLK SPG GL F +N +
Sbjct: 1044 VGRLIYLCHTRPDITYAVSMVSRYMHDPRSGHMDAVYRILRYLKGSPGKGLWFKKNGHLE 1103
Query: 1195 IQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCEL 1254
++GY DADWA CPD RRS SGYC ++G +LVSW++KKQ VSRS+ EAEYRA++ + EL
Sbjct: 1104 VEGYCDADWASCPDDRRSTSGYCVFVGGNLVSWRSKKQPVVSRSTAEAEYRAMSVSLSEL 1163
Query: 1255 QWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILK 1314
WL LL +L + L+CDN+SA+ IA NPV H+RTKH+++D ++EKL G+L+
Sbjct: 1164 LWLRNLLSELMLPVDTPMKLWCDNKSAISIANNPVQHDRTKHVELDRFFIKEKLDEGVLE 1223
Query: 1315 LLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
L + + QVAD FTK L + K+ M
Sbjct: 1224 LEFVMSGGQVADCFTKGLGVKECNSSCDKMGM 1255
>UniRef100_Q6F356 Putative polyprotein [Oryza sativa]
Length = 1256
Score = 612 bits (1577), Expect = e-173
Identities = 367/907 (40%), Positives = 507/907 (55%), Gaps = 46/907 (5%)
Query: 479 DIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIVKA 538
D+WGP A + KY+++ +D +SRF W+ L K EV Q+ + F LV+ F + + +
Sbjct: 350 DVWGP-APVAGGGKKYYVSFIDAFSRFTWIYFLKYKSEVFQKFQEFQCLVERMFNRKIIS 408
Query: 539 IRSDNGPEFL-LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSKLPKK 597
+++D G E+ L F++ GI H SC T QQNG ERKH+HI+ + +LL + LP K
Sbjct: 409 MQTDWGGEYQKLNTFFNKIGITHHVSCPHTHQQNGSAERKHRHIVEVGLSLLAHASLPLK 468
Query: 598 MWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNNRSKLD 657
W + V+L+NR+P+K+L SP E L+++ D + ++ FG C+ N K+
Sbjct: 469 FWDEAYQSGVYLINRMPTKVLGYSSPLECLFKQTPDYQALRTFGCACWPDLRPYNSHKMH 528
Query: 658 PRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTCLDLQTQA 717
R+++C FLGY KG+ +D T I+ISRDV+F E V P+ N A T L +
Sbjct: 529 FRSKRCTFLGYSPLHKGFRCLDSSTGRIYISRDVVFDESVFPFAELNPNAGTNLRKEVNL 588
Query: 718 ESTADKLN--------------------VASE---------IQSAKGDASVSTPCASIED 748
AD LN VA+E + S + DA+ S S
Sbjct: 589 -LPADMLNPGDVQLNDHVNYYPTAPANMVAAENPVENTEENLASTRDDAADSG--GSDTG 645
Query: 749 GVQHEMPSGASIEDEIPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSAL 808
+ + P+ + D N D SI+A + S L N Q
Sbjct: 646 TISNADPADDAAGDMTANPNLNDSSTHESSISASASPASQSSVATAPEAT-LPNPQQHQQ 704
Query: 809 PIRSST----AHSIVHHYSD----ERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAM 860
+RSST A V H E++ T + EP +A D +W+ AM
Sbjct: 705 ALRSSTPEGEASRPVTHLQKGIRKEKIYTDRTVKYGCLTTTGEPRDLHDALHDTNWKHAM 764
Query: 861 QAEIKALEKNGTWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLD 920
A+ AL N TW LV +G I KWVY++KR DG+L RYKARLVAKG+ Q G+D
Sbjct: 765 DAKFTALLHNKTWHLVPPQKGRNIIDYKWVYKIKRKQDGSLDRYKARLVAKGFKQRYGID 824
Query: 921 YFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVPS-VKP 979
Y DTFSPV K T+R+IL++A S+ W L QLDV NAFLHG L+E+VYM P G V P
Sbjct: 825 YEDTFSPVVKAATIRIILSIAVSRGWTLRQLDVQNAFLHGILEEEVYMKQPPGYEDKVHP 884
Query: 980 NQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDD 1039
+ VCKL K+LYGLKQA R WY KLS L+ LGF+ + +D SLF +G +LVYVDD
Sbjct: 885 DYVCKLDKALYGLKQAPRAWYAKLSQKLQHLGFQGSKADTSLFFYNKGGLIIFVLVYVDD 944
Query: 1040 VILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQET 1099
+I+ + + L + F +KDLG L YFLG+EV ++SGI L Q KY DLL+
Sbjct: 945 IIVASSRQDAVPALLKDLQKDFALKDLGDLHYFLGIEVNKASSGIVLTQEKYVTDLLRRV 1004
Query: 1100 GTLGSKPVATPLDPSIRLSQEQGKPY--DDPAAYRRLVGRLLYLTTTRPDISHATQQLSQ 1157
G KPV+TPL S +L+ +G +D + YR +VG L YLT TRPDI ++ Q
Sbjct: 1005 GMTDCKPVSTPLSTSEKLTLHEGDLLGPNDASNYRSVVGALQYLTLTRPDIYFPVNKVCQ 1064
Query: 1158 YMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYC 1217
++ P H+ A +R+LRYLK LGL ++ ++ + GYSDADWAG D RRS G+
Sbjct: 1065 FLHAPTIVHWAAMKRILRYLKQCTKLGLKISKSKSMLVSGYSDADWAGNIDDRRSTGGFA 1124
Query: 1218 FYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCD 1277
++G +LVSW AKKQ TV RSS E+EY+ALA AT E+ W+ LL++L V L+CD
Sbjct: 1125 VFLGDNLVSWNAKKQATVPRSSTESEYKALANATAEIMWIQTLLEELSVPSPPMARLWCD 1184
Query: 1278 NQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVF 1337
N A ++++NPVFH RTKH+++D H VRE++Q +L++ IPT QVAD FTKAL R
Sbjct: 1185 NLGAKYLSSNPVFHARTKHIEVDYHFVRERMQRKLLEVEFIPTGDQVADGFTKALSARQL 1244
Query: 1338 QGFATKL 1344
+ F L
Sbjct: 1245 ENFKYNL 1251
>UniRef100_Q94IU9 Copia-like polyprotein [Arabidopsis thaliana]
Length = 1466
Score = 611 bits (1575), Expect = e-173
Identities = 350/941 (37%), Positives = 512/941 (54%), Gaps = 32/941 (3%)
Query: 419 WHFRLGHLSHDRLLALHTVQNSISVSKSI---VCDVCHLAKQKRKMFTVSVSKAQKCFDL 475
WH RLGH S+ ++L + I V+KS VC+ C + K R F S +A K D
Sbjct: 458 WHHRLGH-SNSKILQQLLTRKEIQVNKSRTSPVCEPCQMGKSTRLQFFSSDFRALKPLDR 516
Query: 476 VHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQI 535
VH D+WGP S KY+ +DD+SRF W L K + + LV+ Q G
Sbjct: 517 VHCDLWGPSPVVSNQGFKYYAVFVDDFSRFSWFFPLRMKSKFISVFIAYQKLVENQLGTK 576
Query: 536 VKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQS 592
+K +SD G EF L + GI H+ SC TPQQNG ERKH+H++ + ++L+ S
Sbjct: 577 IKEFQSDGGGEFTSNKLKEHFREHGIHHRISCPYTPQQNGVAERKHRHLVELGLSMLYHS 636
Query: 593 KLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTNN 652
P K W + A +L N +PS +LK SPYE L+++ VD ++VFG+ C+
Sbjct: 637 HTPLKFWVEAFFTANYLSNLLPSSVLKEISPYETLFQQKVDYTPLRVFGTACYPCLRPLA 696
Query: 653 RSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKG---SNSPAWT 709
++K DPR+ +C+FLGY KGY + T +++ISR VIF E P+K S P +
Sbjct: 697 KNKFDPRSLQCVFLGYHNQYKGYRCLYPPTGKVYISRHVIFDEAQFPFKEKYHSLVPKYQ 756
Query: 710 CLDLQTQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPNTQT 769
LQ + S P + ++ + P S + N +T
Sbjct: 757 TTLLQAWQHTDL---------------TPPSVPSSQLQPLARQMTPMATSENQPMMNYET 801
Query: 770 VDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRS-STAHSIVHHYSDERLS 828
+ + +++ E E PV + +AL S H ++ S + +
Sbjct: 802 EEAVNVNMETSSDEETESNDEFDHEVAPVLNDQNEDNALGQGSLENLHPMITR-SKDGIQ 860
Query: 829 TAHRAYALNIAQDQ--EPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIG 886
+ YAL +++ EP T A K W A+ EI + TW LV E + +
Sbjct: 861 KPNPRYALIVSKSSFDEPKTITTAMKHPSWNAAVMDEIDRIHMLNTWSLVPATEDMNILT 920
Query: 887 NKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNW 946
+KWV++ K DGT+ + KARLVAKG++Q EG+DY +TFSPV + T+R++L A + W
Sbjct: 921 SKWVFKTKLKPDGTIDKLKARLVAKGFDQEEGVDYLETFSPVVRTATIRLVLDTATANEW 980
Query: 947 HLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQASRKWYEKLSA 1005
L QLDV NAFLHG L E V+M P+G V KPN VC+L K+LYGLKQA R W++ S
Sbjct: 981 PLKQLDVSNAFLHGELQEPVFMFQPSGFVDPNKPNHVCRLTKALYGLKQAPRAWFDTFSN 1040
Query: 1006 HLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKD 1065
L GF+ + SD SLFV Q LL+YVDD++L G+ + +L+ F +KD
Sbjct: 1041 FLLDFGFECSTSDPSLFVCHQNGQSLILLLYVDDILLTGSDQLLMDKLLQALNNRFSMKD 1100
Query: 1066 LGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPY 1125
LG +YFLG+E+ +G+ L Q Y D+L + G P+ TPL L +P+
Sbjct: 1101 LGPPRYFLGIEIESYNNGLFLHQHAYASDILHQAGMTECNPMPTPLPQ--HLEDLNSEPF 1158
Query: 1126 DDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGL 1185
++P +R L G+L YLT TRPDI +A + Q M P + F +R+LRY+K + +GL
Sbjct: 1159 EEPTYFRSLAGKLQYLTITRPDIQYAVNFICQRMHAPTNSDFGLLKRILRYVKGTINMGL 1218
Query: 1186 LFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYR 1245
++ + G+ D+D+AGC DTRRS +G+C +G +L+SW AK+Q T+S SS EAEYR
Sbjct: 1219 PIRKHHNPVLSGFCDSDYAGCKDTRRSTTGFCILLGSTLISWSAKRQPTISHSSTEAEYR 1278
Query: 1246 ALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVR 1305
AL+ E+ W+ LL+DL ++ +FCDN SA++++ANP H+R+KH D D H +R
Sbjct: 1279 ALSDTAREITWISSLLRDLGISQHQPTRVFCDNLSAVYLSANPALHKRSKHFDKDFHYIR 1338
Query: 1306 EKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
E++ G+++ IP T+Q+ADVFTK+L R F KL +
Sbjct: 1339 ERVALGLIETQHIPATIQLADVFTKSLPRRPFITLRAKLGV 1379
>UniRef100_Q94KV0 Polyprotein [Arabidopsis thaliana]
Length = 1453
Score = 600 bits (1546), Expect = e-169
Identities = 352/954 (36%), Positives = 505/954 (52%), Gaps = 49/954 (5%)
Query: 418 LWHFRLGHLSHDRLLALHTVQNSISVSKSI---VCDVCHLAKQKRKMFTVSVSKAQKCFD 474
+WH RLGH S+ R+L IS +KS VC+ C + K + F S S+
Sbjct: 458 IWHHRLGH-SNSRILQQLKSSKEISFNKSRMSPVCEPCQMGKSSKLQFFSSNSRELDLLG 516
Query: 475 LVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQ 534
+H D+WGP S KY++ +DDYSR+ W L K + F LV+ QF
Sbjct: 517 RIHCDLWGPSPVVSKQGFKYYVVFVDDYSRYSWFYPLKAKSDFFAVFVAFQNLVENQFNT 576
Query: 535 IVKAIRSDNGPEF---LLPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQ 591
+K +SD G EF L+ + GI H+ SC TPQQNG ERKH+H + + +++F
Sbjct: 577 KIKVFQSDGGGEFTSNLMKKHLTDCGIQHRISCPYTPQQNGIAERKHRHFVELGLSMMFH 636
Query: 592 SKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLTN 651
S P + W + A FL N +PS L N SP E L ++ + M++VFG+ C+
Sbjct: 637 SHTPLQFWVEAFFTASFLSNMLPSPSLGNVSPLEALLKQKPNYAMLRVFGTACYPCLRPL 696
Query: 652 NRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYK---------- 701
K +PR+ +C+FLGY KGY + T ++ISR VIF E P+K
Sbjct: 697 GEHKFEPRSLQCVFLGYNSQYKGYRCLYPPTGRVYISRHVIFDEETFPFKQKYQFLVPQY 756
Query: 702 -GSNSPAWTCLDLQTQA-------ESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHE 753
S AW Q E + L IQ + + P A + +GV +E
Sbjct: 757 ESSLLSAWQSSIPQADQSLIPQAEEGKIESLAKPPSIQKNTIQDTTTQP-AILTEGVLNE 815
Query: 754 MPSGASIEDEIPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSS 813
ED T+T + + E EV V + H P+ +
Sbjct: 816 EEE----EDSFEETETESLNEETHTQNDEAEVTVEEEVQQEPENTH---------PMTTR 862
Query: 814 TAHSIVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTW 873
+ I H S+ R + +++ +EP + EA W A+ E++ + TW
Sbjct: 863 SKAGI--HKSNTRYALLTSKFSV-----EEPKSIDEALNHPGWNNAVNDEMRTIHMLHTW 915
Query: 874 KLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTT 933
LV E + +G +WV++ K DG++ + KARLVAKG++Q EGLDY +TFSPV + T
Sbjct: 916 SLVQPTEDMNILGCRWVFKTKLKPDGSVDKLKARLVAKGFHQEEGLDYLETFSPVVRTAT 975
Query: 934 VRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGL 992
+R++L +A ++ W++ QLDV NAFLHG L E VYM P G V KP+ VC+L K+LYGL
Sbjct: 976 IRLVLDVATAKGWNIKQLDVSNAFLHGELKEPVYMLQPPGFVDQEKPSYVCRLTKALYGL 1035
Query: 993 KQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQL 1052
KQA R W++ +S +L GF + SD SLF + LL+YVDD++L G+ Q
Sbjct: 1036 KQAPRAWFDTISNYLLDFGFSCSKSDPSLFTYHKNGKTLVLLLYVDDILLTGSDHNLLQE 1095
Query: 1053 VKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLD 1112
+ SL++ F +KDLG YFLG+E+ S G+ L Q Y D+L + + TPL
Sbjct: 1096 LLMSLNKRFSMKDLGAPSYFLGVEIESSPEGLFLHQTAYAKDILHQAAMSNCNSMPTPLP 1155
Query: 1113 PSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQR 1172
I P +P +R L G+L YLT TRPDI A + Q M +P F +R
Sbjct: 1156 QHIENLNSDLFP--EPTYFRSLAGKLQYLTITRPDIQFAVNFICQRMHSPTTADFGLLKR 1213
Query: 1173 VLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQ 1232
+LRY+K + LGL +N +++ YSD+DWAGC +TRRS +G+C +G +L+SW AK+Q
Sbjct: 1214 ILRYVKGTIHLGLHIKKNQNLSLVAYSDSDWAGCKETRRSTTGFCTLLGCNLISWSAKRQ 1273
Query: 1233 TTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHE 1292
TVS+SS EAEYRAL EL WL +LL+D+ VT T ++ CDN SA++++ANP H
Sbjct: 1274 ETVSKSSTEAEYRALTAVAQELTWLSFLLRDIGVTQTHPTLVKCDNLSAVYLSANPALHN 1333
Query: 1293 RTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
R+KH D D H +RE++ G+++ I TLQ+AD+FTK L R F KL +
Sbjct: 1334 RSKHFDTDYHYIREQVALGLVETKHISATLQLADIFTKPLPRRAFIDLRIKLGV 1387
>UniRef100_Q9T0C5 Retrotransposon like protein [Arabidopsis thaliana]
Length = 1515
Score = 591 bits (1524), Expect = e-167
Identities = 359/977 (36%), Positives = 523/977 (52%), Gaps = 60/977 (6%)
Query: 418 LWHFRLGHLSHDRLLAL-HTVQNSISVSKSIVCDVCHLAKQKRKMFTVSVSKAQKCFDLV 476
+WH RLGH + + L L T ++ + S +C+ C + K R F S + + + +
Sbjct: 457 VWHQRLGHPNKEVLQHLIKTKAIVVNKTSSNMCEACQMGKVCRLPFVASEFVSSRPLERI 516
Query: 477 HMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFGQIV 536
H D+WGP S +Y++ +D+YSRF W L K + F LV+ Q+ +
Sbjct: 517 HCDLWGPAPVTSAQGFQYYVIFIDNYSRFTWFYPLKLKSDFFSVFVLFQQLVENQYQHKI 576
Query: 537 KAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLFQSK 593
+ D G EF+ A ++ GI SC TPQQNG ER+H+++ + +L+F SK
Sbjct: 577 AMFQCDGGGEFVSYKFVAHLASCGIKQLISCPHTPQQNGIAERRHRYLTELGLSLMFHSK 636
Query: 594 LPKKMWCYSVLHAVFLMNRIPSKLLK-NKSPYELLYREAVDLEMMKVFGSLCFATTLTNN 652
+P K+W + + FL N +PS L NKSPYE+L+ ++VFGS C+
Sbjct: 637 VPHKLWVEAFFTSNFLSNLLPSSTLSDNKSPYEMLHGTPPVYTALRVFGSACYPYLRPYA 696
Query: 653 RSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKG--------SN 704
++K DP++ C+FLGY KGY + T +++I R V+F E PY S
Sbjct: 697 KNKFDPKSLLCVFLGYNNKYKGYRCLHPPTGKVYICRHVLFDERKFPYSDIYSQFQTISG 756
Query: 705 SP---AW------TCLDLQTQAESTADKLNVASEIQSAKGDASVSTPCA----------S 745
SP AW T L +T + + D + SA +SV T CA
Sbjct: 757 SPLFTAWQKGFSSTALSRETPSTNVEDII-----FPSATVSSSVPTGCAPNIAETATAPD 811
Query: 746 IEDGVQHEM--PSGASIEDEIPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANF 803
++ H+M P +P TQ + + + E + + P + ++++ F
Sbjct: 812 VDVAAAHDMVVPPSPITSTSLP-TQPEESTSDQNHYSTDSETAISSAMTP--QSINVSLF 868
Query: 804 QFSALP-----IRSSTAHSIVHHYSDER----LSTAHRAYALNIAQDQ--EPSTFAEANK 852
+ S P I S+TA H R ++ + YAL + EP + EA K
Sbjct: 869 EDSDFPPLQSVISSTTAAPETSHPMITRAKSGITKPNPKYALFSVKSNYPEPKSVKEALK 928
Query: 853 DLHWREAMQAEIKALEKNGTWKLVDLPEGV-KPIGNKWVYRVKRNVDGTLARYKARLVAK 911
D W AM E+ + + TW LV PE V + +G KWV++ K N DG+L R KARLVA+
Sbjct: 929 DEGWTNAMGEEMGTMHETDTWDLVP-PEMVDRLLGCKWVFKTKLNSDGSLDRLKARLVAR 987
Query: 912 GYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIP 971
GY Q EG+DY +T+SPV + TVR IL +A W L QLDV NAFLH L E V+MT P
Sbjct: 988 GYEQEEGVDYVETYSPVVRSATVRSILHVATINKWSLKQLDVKNAFLHDELKETVFMTQP 1047
Query: 972 AGV--PSVKPNQVCKLLKSLYGLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSS 1029
G PS +P+ VCKL K++Y LKQA R W++K S++L GF + SD SLFV +G
Sbjct: 1048 PGFEDPS-RPDYVCKLKKAIYDLKQAPRAWFDKFSSYLLKYGFICSFSDPSLFVYLKGRD 1106
Query: 1030 FTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQR 1089
LL+YVDD+IL GN Q + + L F +KD+G L YFLG++ + G+ L Q
Sbjct: 1107 VMFLLLYVDDMILTGNNDVLLQQLLNILSTEFRMKDMGALHYFLGIQAHYHNDGLFLSQE 1166
Query: 1090 KYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDIS 1149
KY DLL G + TPL + L Q KP+ +P +RRL G+L YLT TRPDI
Sbjct: 1167 KYTSDLLVNAGMSDCSSMPTPLQ--LDLLQGNNKPFPEPTYFRRLAGKLQYLTLTRPDIQ 1224
Query: 1150 HATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDT 1209
A + Q M P F +R+L YLK + +G+ N+ ++ YSD+DWAGC DT
Sbjct: 1225 FAVNFVCQKMHAPTMSDFHLLKRILHYLKGTMTMGINLSSNTDSVLRCYSDSDWAGCKDT 1284
Query: 1210 RRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCT 1269
RRS G+C ++G +++SW AK+ TVS+SS EAEYR L++A E+ W+ +LLQ++ +
Sbjct: 1285 RRSTGGFCTFLGYNIISWSAKRHPTVSKSSTEAEYRTLSFAASEVSWIGFLLQEIGLPQQ 1344
Query: 1270 ATPVLFCDNQSALHIAANPVFHERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFT 1329
P ++CDN SA++++ANP H R+KH +D + VRE++ G L + IP + Q+AD+FT
Sbjct: 1345 QIPEMYCDNLSAVYLSANPALHSRSKHFQVDYYYVRERVALGALTVKHIPASQQLADIFT 1404
Query: 1330 KALQPRVFQGFATKLAM 1346
K+L F KL +
Sbjct: 1405 KSLPQAPFCDLRFKLGV 1421
>UniRef100_Q60DG5 Putative polyprotein [Oryza sativa]
Length = 1094
Score = 568 bits (1463), Expect = e-160
Identities = 333/947 (35%), Positives = 524/947 (55%), Gaps = 54/947 (5%)
Query: 419 WHFRLGHLSHDRL--LALHTVQNSISVSKSI--VCDVCHLAKQKRKMFTVSVS-KAQKCF 473
WH R GHL+ L LA + + V + +CD C KQ+R F +AQ
Sbjct: 180 WHARFGHLNFQALRRLAQAEMVRGLPVIDHVDQLCDGCLAGKQRRVPFPDKARFRAQDAL 239
Query: 474 DLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVKTQFG 533
+LVH D+ GP+A A+ KYFL ++DD SRF+W+ LL+ K E +K F V+ + G
Sbjct: 240 ELVHGDLCGPIAPATPGGRKYFLLLVDDMSRFMWIRLLSGKHEAAAAIKQFKAGVEMESG 299
Query: 534 QIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIARALLF 590
+ ++A+R+D G EF F + +G+ Q + +PQQNG VER++ ++ AR++L
Sbjct: 300 RKLRALRTDRGGEFTSVEFTEFCADRGVSRQLTAPYSPQQNGVVERRNLTVVAAARSMLK 359
Query: 591 QSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFATTLT 650
+ +P + W +V+ AV+L+NR P+K L +PYE + +E +KVFG + + T+
Sbjct: 360 AAGMPAQFWGEAVVAAVYLLNRSPTKSLDGVTPYEAWHGRRPSVEHLKVFGCVGYVKTVK 419
Query: 651 NNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSPAWTC 710
N KLD R + +F+GY+QG K Y + D +++ ++SRDV+F E + W
Sbjct: 420 PNLRKLDDRGTRMVFIGYEQGSKAYRMYDPVSRRFYVSRDVVFDEAAM---------WPW 470
Query: 711 LDLQTQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPNTQTV 770
D + + V G+ P +E G E + S T
Sbjct: 471 RDPEVTQTGGEEDFTVEFFSTPLGGNR---VPDVVVEHGGARETETAPS------PLATP 521
Query: 771 DEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIV--------HHY 822
D P +T+ V T P A+ + P R T ++++ Y
Sbjct: 522 DAAPVWSPVTSSSPAGVEFCTPPSD-----ASIESDGAPPRFRTVNNVLATTTPVLDFDY 576
Query: 823 SDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGV 882
DE L +QEP +F EA K+ W +AM+ E+ ++E N TW L DLP
Sbjct: 577 DDECLIA-----------EQEPFSFKEAEKEQCWMKAMEEEMSSIEGNNTWFLCDLPSDH 625
Query: 883 KPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAA 942
+ IG KWVY++KR+ +G + +YKARLVAKGY Q +G+DY + F+PVA++ TVR+++ALAA
Sbjct: 626 RAIGLKWVYKIKRSAEGEILKYKARLVAKGYVQQQGIDYEEVFAPVARMETVRLLVALAA 685
Query: 943 SQNWHLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQASRKWYE 1001
+ W +H +DV +AFL+G L+EDVY+ P G V K N+V KL K+LYGLKQA R WY
Sbjct: 686 HEGWQIHHMDVKSAFLNGELEEDVYVVQPPGFVVEHKENKVLKLKKALYGLKQAPRAWYA 745
Query: 1002 KLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAF 1061
KL + L L F ++A++++++ + +G++ + VYVDD+I+ G TE K+ + F
Sbjct: 746 KLDSTLANLDFVRSATENAVYTRGEGNARLVVGVYVDDLIITGALGTEIAKFKEQMRSMF 805
Query: 1062 GIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQ 1121
+ DLG+L Y+LG+EV + GI++ Q Y +L++ G G P P+D ++L ++
Sbjct: 806 SMSDLGLLSYYLGMEVKQTEEGITMSQAGYAGRILEKAGMQGCNPCQVPMDARLKL-KKG 864
Query: 1122 GKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASP 1181
+ D +R ++G L YL TRPD+S++ +S+YM NP H+ A + +LRY+ S
Sbjct: 865 VEDCIDATQFRSIIGSLRYLVNTRPDLSYSVGYVSRYMENPGAEHWAAMKHILRYVAGSL 924
Query: 1182 GLGLLFPRNST--INIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSS 1239
+GL F + + GYSD+D AG D R+S +G F +G +L++W+++KQ V+ SS
Sbjct: 925 NIGLKFRKGEEKFPRLVGYSDSDMAGDVDDRKSTTGVLFKLGENLITWQSQKQKIVALSS 984
Query: 1240 NEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDI 1299
EAEY A A C+ WL LL +L++ +L DN+SA+++ NPV H+R+KH+D
Sbjct: 985 CEAEYIAATTAACQGIWLARLLGELQMKKPCCAMLKVDNKSAINLCKNPVLHDRSKHIDT 1044
Query: 1300 DCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
H +RE+++ +++ + Q+AD+ TK L F KL +
Sbjct: 1045 RYHFIRERVERKEVEVEYTSSAEQLADILTKPLGKVRFLELRGKLGL 1091
>UniRef100_Q7XPB1 OSJNBb0026E15.10 protein [Oryza sativa]
Length = 1449
Score = 566 bits (1459), Expect = e-159
Identities = 331/944 (35%), Positives = 516/944 (54%), Gaps = 39/944 (4%)
Query: 415 PGALWHFRLGHLSHDRL--LALHTVQNSISVSKSI--VCDVCHLAKQKRKMF-TVSVSKA 469
P WH R GHL+ L LA + + + + + VCD C L KQ+R F T S +A
Sbjct: 524 PAWRWHARYGHLNFPALRKLAQQEMVRGLPLLQQVTQVCDGCLLGKQRRAAFPTQSKYRA 583
Query: 470 QKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVK 529
+ +LVH D+ GP+ A+ ++YFL ++DD SR++W+ ++ +K E +K+F +
Sbjct: 584 DEHLELVHGDLCGPIEPATPAGNRYFLLLVDDMSRYMWLTMIRSKDEAANAIKHFQARAE 643
Query: 530 TQFGQIVKAIRSDNGPEFLLPAF--YSAQ-GIVHQKSCVSTPQQNGRVERKHQHILNIAR 586
+ G+ ++A+R D G EF F Y A G+ Q + +PQQNG VER++Q I+ AR
Sbjct: 644 VESGRKLRALRMDRGSEFTSIEFGEYCANLGVGRQLTAPYSPQQNGVVERRNQTIVATAR 703
Query: 587 ALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFA 646
+++ +P + W ++ AVFL+NR P+K L N++PYE Y + + ++ FG +
Sbjct: 704 SMMKAKGVPGRFWGEAMSTAVFLLNRSPTKSLDNQTPYEAWYGQWPAVHFLRTFGCVGHV 763
Query: 647 TTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSP 706
KLD R+ + LGY+QG K Y L D +++ + +SRDV+F E +
Sbjct: 764 KITKPGLKKLDDRSAPMVLLGYEQGSKAYRLYDPVSERVHVSRDVVFDEDI--------- 814
Query: 707 AWTCLDLQTQAESTADKLNVASEIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIPN 766
AW + + V + + G A S+P PS S P
Sbjct: 815 AWDWGPVTPDGAPQLEPFTVEQVVTTTIGTAPASSPTP----------PSPPSPAPSAPT 864
Query: 767 TQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHSIVHHYSDER 826
T P P + V T P + + A+ +P + + +
Sbjct: 865 TPAPPSPPSPEA--------VEFVTPPTQDSILDADADDDVVPRYRLVDNLLGNASPPGH 916
Query: 827 LSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVDLPEGVKPIG 886
L++ EP++ AEA D WR AMQ E+ A+ N TW L DLP G + IG
Sbjct: 917 APRVLEQLELHVVSADEPASLAEAEADPSWRGAMQDELNAIVDNDTWSLTDLPHGHRAIG 976
Query: 887 NKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVILALAASQNW 946
KWVY++KR+ G + RYKARLVAKGY Q +G+D+ + F+ VA+L +VR++LA+AA Q W
Sbjct: 977 LKWVYKLKRDEQGAIVRYKARLVAKGYVQRQGVDFDEVFALVARLESVRLLLAVAAHQGW 1036
Query: 947 HLHQLDVDNAFLHGNLDEDVYMTIPAG-VPSVKPNQVCKLLKSLYGLKQASRKWYEKLSA 1005
+H +DV +AFL+G L E+VY++ P G V N+V +L K+LYGL+QA R W KL +
Sbjct: 1037 QVHHMDVKSAFLNGELLEEVYVSQPPGFVDDNHKNKVYRLHKALYGLRQAPRAWNAKLDS 1096
Query: 1006 HLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDSLHQAFGIKD 1065
L +LGF +++S+H ++ + +G + VYVDD+I+ G+ E + K + + F + D
Sbjct: 1097 SLLSLGFHRSSSEHGVYTRTRGGRRLMVGVYVDDLIITGDHDDEIRSFKGEMMKLFKMSD 1156
Query: 1066 LGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIRLSQEQGKPY 1125
LG L+Y+LG+EV + GI+L Q Y +L++ G P TP++ ++L + P
Sbjct: 1157 LGALRYYLGIEVTQDSDGITLGQAAYAGKILEKAGLKDCNPCQTPMEVRLKLRKGSDFPL 1216
Query: 1126 DDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRYLKASPGLGL 1185
D YR LVG L YL TRPD++ + +S++M +P + H A +R+LRY+ + G+
Sbjct: 1217 VDATLYRSLVGSLRYLVNTRPDLAFSVGYVSRFMESPREDHLAAVRRILRYVAGTRCWGI 1276
Query: 1186 LF---PRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVSRSSNEA 1242
F R + + GYSD+D AG PD R+S SG F+I V+W++ KQ V+ SS EA
Sbjct: 1277 RFGPGARCALPMLVGYSDSDLAGDPDERKSTSGQIFFINGGPVTWQSSKQKVVALSSCEA 1336
Query: 1243 EYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKHLDIDCH 1302
EY A A ATC+ WL LL ++ P+L DNQS + + NPV H+R+KH+D+ H
Sbjct: 1337 EYIAAAAATCQGVWLARLLAEVLGDEITAPLLKVDNQSTISLIKNPVHHDRSKHIDVKYH 1396
Query: 1303 VVREKLQAGILKLLPIPTTLQVADVFTKALQPRVFQGFATKLAM 1346
+RE + +++++ + T Q+ D+FTK+L FQ +K+ +
Sbjct: 1397 YIRECAEKKLIEMMFVGTAEQLGDIFTKSLGRTRFQELRSKIGV 1440
>UniRef100_Q9XEJ4 Copia-type pol polyprotein [Zea mays]
Length = 1063
Score = 560 bits (1444), Expect = e-158
Identities = 340/947 (35%), Positives = 496/947 (51%), Gaps = 44/947 (4%)
Query: 416 GALWHFRLGHLS----HDRLLALHTVQ-NSISVSKSIVCDVCHLAKQKRKMFT-VSVSKA 469
G LWH RL H+ H L H + ++ K +C C KQ ++
Sbjct: 119 GWLWHRRLAHVGMKNLHKLLKGEHILGLTNVHFEKDRICSACQAGKQVGTHHPHKNIMTT 178
Query: 470 QKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVK 529
+ +L+HMD++GP+A S+ KY L ++DDYSRF WV L K + Q+ +K F+ +
Sbjct: 179 DRPLELLHMDLFGPIAYISIGGSKYCLVIVDDYSRFTWVFFLQEKSQTQETLKGFLRRAQ 238
Query: 530 TQFGQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIAR 586
+FG +K IRSDNG EF + +F +GI H+ S TPQQNG VERK++ +L++AR
Sbjct: 239 NEFGLRIKKIRSDNGTEFKNSQIESFLEEEGIKHEFSSPYTPQQNGVVERKNRTLLDMAR 298
Query: 587 ALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFA 646
+L + K P + W +V A + +NR+ + K+ YELL + ++ +VFGS CF
Sbjct: 299 TMLDEYKTPDRFWAEAVNTACYAINRLYLHRILKKTSYELLTGKKPNISYFRVFGSKCFI 358
Query: 647 TTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSP 706
+SK P+ + LGY + Y + + T + +S DV+F E +N
Sbjct: 359 LIKRGRKSKFAPKTVEGFLLGYDSNTRAYRVFNKSTGLVEVSCDVVFDE-------TNGS 411
Query: 707 AWTCLDLQTQAESTADKLNVAS---------------EIQSAKGDASVSTPCASIEDGVQ 751
+DL E A + + + Q + ++P ED Q
Sbjct: 412 QVEQVDLDEIGEEQAPCIALRNMSIGDVCPKESEEPPSTQDQPSSSMQASPPTQNEDEAQ 471
Query: 752 HEMPSGASIEDEIPNTQTVDEIPQPGSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIR 811
++ EDE P + D+ G T + E E RP VH A + +
Sbjct: 472 NDEEQNQ--EDEPPQDDSNDQ----GGDTNDQEKEDEEEPRPPHPRVHQAIQRDHPV--- 522
Query: 812 SSTAHSIVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNG 871
T +H R AH + EP EA +D W AMQ E+ +N
Sbjct: 523 -DTILGDIHKGVTTRSRVAHFCEHYSFVSSIEPHRVEEALQDSDWVVAMQEELNNFTRNE 581
Query: 872 TWKLVDLPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKL 931
W LV P +G KWV+R K++ G + R KARLVAKGY+QVEGLD+ +T++PVA+L
Sbjct: 582 VWHLVPRPNQ-NVVGTKWVFRNKQDEHGVVTRNKARLVAKGYSQVEGLDFGETYAPVARL 640
Query: 932 TTVRVILALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVP-SVKPNQVCKLLKSLY 990
++R++LA A + L+Q+DV +AFL+G + E+VY+ P G S PN V +L K+LY
Sbjct: 641 ESIRILLAYATYHGFKLYQMDVKSAFLNGPIKEEVYVEQPPGFEDSEYPNHVYRLSKALY 700
Query: 991 GLKQASRKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEF 1050
GLKQA R WYE L L GFK +D +LF K + +YVDD+I +
Sbjct: 701 GLKQAPRAWYECLRDFLIANGFKVGKADPTLFTKTLENDLFVCQIYVDDIIFGSTNKSTC 760
Query: 1051 QLVKDSLHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATP 1110
+ + Q F + +G LKYFLG +V G +CQ KY D+L + G +KP+ TP
Sbjct: 761 EEFSRIMTQKFEMSMMGELKYFLGFQVKQLQEGTFICQTKYTQDILTKFGMKDAKPIKTP 820
Query: 1111 LDPSIRLSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAA 1170
+ + L + G D YR ++G LLYL +RPDI + +++ S+P + H A
Sbjct: 821 MGTNGHLDLDTGGKSVDQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKESHLTAV 880
Query: 1171 QRVLRYLKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAK 1230
+R+LRYL +P GL +PR ST ++ GYSDADWAGC R+S SG C ++GRSLVSW +K
Sbjct: 881 KRILRYLAYTPKFGLWYPRGSTFDLIGYSDADWAGCKINRKSTSGTCQFLGRSLVSWASK 940
Query: 1231 KQTTVSRSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVF 1290
KQ +V+ S+ EAEY A + +L W+ L D T P+L CDN+SA+ +A NPV
Sbjct: 941 KQNSVALSTAEAEYIAAGHCCAQLLWMRQTLLDYGYKLTKVPLL-CDNESAIKMADNPVE 999
Query: 1291 HERTKHLDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVF 1337
H RTKH+ I H +R+ Q G +++ I T Q+AD+FTK L + F
Sbjct: 1000 HSRTKHIAIRYHFLRDHQQKGDIEISYINTKDQLADIFTKPLDEQSF 1046
>UniRef100_Q8H6I8 Putative gag-pol polyprotein [Zea mays]
Length = 1892
Score = 555 bits (1431), Expect = e-156
Identities = 337/941 (35%), Positives = 493/941 (51%), Gaps = 32/941 (3%)
Query: 416 GALWHFRLGHLS----HDRLLALHTVQ-NSISVSKSIVCDVCHLAKQKRKMFT-VSVSKA 469
G LWH RL H+ H L H + ++ K +C C KQ ++
Sbjct: 948 GWLWHRRLAHVGMKNLHKLLKGEHILGLTNVHFEKDRICSACQAGKQVGTHHPHKNIMTT 1007
Query: 470 QKCFDLVHMDIWGPLAQASVHNHKYFLTVLDDYSRFVWVVLLNNKGEVQQQVKNFITLVK 529
+ +L+HMD++GP+A S+ KY L ++DDYSRF WV L K + Q+ +K F+ +
Sbjct: 1008 DRPLELLHMDLFGPIAYISIGGSKYCLVIVDDYSRFTWVFFLQEKSQTQETLKGFLRRAQ 1067
Query: 530 TQFGQIVKAIRSDNGPEFL---LPAFYSAQGIVHQKSCVSTPQQNGRVERKHQHILNIAR 586
+FG +K IRSDNG EF + +F +GI H+ S TPQQNG VERK++ +L++AR
Sbjct: 1068 NEFGLRIKKIRSDNGTEFKNSQIESFLEEEGIKHEFSSPYTPQQNGVVERKNRTLLDMAR 1127
Query: 587 ALLFQSKLPKKMWCYSVLHAVFLMNRIPSKLLKNKSPYELLYREAVDLEMMKVFGSLCFA 646
+L + K P + W +V A + +NR+ + K+ YELL + ++ +VFGS CF
Sbjct: 1128 TMLDEYKTPDRFWAEAVNTACYAINRLYLHRILKKTSYELLTGKKPNISYFRVFGSKCFI 1187
Query: 647 TTLTNNRSKLDPRARKCIFLGYKQGMKGYVLMDLITQEIFISRDVIFHEHVLPYKGSNSP 706
+SK P+ + LGY + Y + + T + +S DV+F E +N
Sbjct: 1188 LIKRGRKSKFAPKTVEGFLLGYDSNTRAYRVFNKSTGLVEVSCDVVFDE-------TNGS 1240
Query: 707 AWTCLDLQTQAESTADKLNVAS-EIQSAKGDASVSTPCASIEDGVQHEMPSGASIEDEIP 765
+DL E A + + + I S P + + EDE
Sbjct: 1241 QVEQVDLDEIGEEQAPCIALRNMSIGDVCPKESEEPPSTQDQPPSSMQASPPTQNEDEAQ 1300
Query: 766 NTQTVDEIPQP--------GSITAEGEVEVRRSTRPLKRPVHLANFQFSALPIRSSTAHS 817
N + ++ +P G T + E E RP VH A + + T
Sbjct: 1301 NDEEQNQEVKPPQDKSNDQGGDTNDQEKEDEEEPRPPHPRVHQAIQRDHPV----DTILG 1356
Query: 818 IVHHYSDERLSTAHRAYALNIAQDQEPSTFAEANKDLHWREAMQAEIKALEKNGTWKLVD 877
+H R AH + EP EA +D W AMQ E+ +N W LV
Sbjct: 1357 DIHKGVTTRSRVAHFCEHYSFVSSIEPHRVEEALQDSDWVVAMQEELNNFTRNEVWHLVP 1416
Query: 878 LPEGVKPIGNKWVYRVKRNVDGTLARYKARLVAKGYNQVEGLDYFDTFSPVAKLTTVRVI 937
P +G KWV+R K++ G + R KARLVAKGY+QVEGLD+ +T++PVA+L ++R++
Sbjct: 1417 RPNQ-NVVGTKWVFRNKQDEHGVVTRNKARLVAKGYSQVEGLDFDETYAPVARLESIRIL 1475
Query: 938 LALAASQNWHLHQLDVDNAFLHGNLDEDVYMTIPAGVP-SVKPNQVCKLLKSLYGLKQAS 996
LA A + L+Q+DV +AFL+G + E+VY+ P G S PN V +L K+LYGLKQA
Sbjct: 1476 LAYATYHGFKLYQMDVKSAFLNGPIKEEVYVEQPPGFEDSEYPNHVYRLSKALYGLKQAP 1535
Query: 997 RKWYEKLSAHLETLGFKQTASDHSLFVKFQGSSFTGLLVYVDDVILFGNTVTEFQLVKDS 1056
R WYE L L GFK +D +LF K + +YVDD+I + +
Sbjct: 1536 RAWYECLRDFLIANGFKVGKADPTLFTKTLENDLFVCQIYVDDIIFGSTNESTCEEFSRI 1595
Query: 1057 LHQAFGIKDLGVLKYFLGLEVAHSTSGISLCQRKYCLDLLQETGTLGSKPVATPLDPSIR 1116
+ Q F + +G LKYFLG +V G + Q KY D+L + G +KP+ TP+ +
Sbjct: 1596 MTQKFEMSMMGELKYFLGFQVKQLREGTFISQTKYTQDILAKFGMKDAKPIKTPMGTNGH 1655
Query: 1117 LSQEQGKPYDDPAAYRRLVGRLLYLTTTRPDISHATQQLSQYMSNPMDGHFKAAQRVLRY 1176
L + G D YR ++G LLYL +RPDI + +++ S+P + H A +R+LRY
Sbjct: 1656 LDLDTGGKSVDQKVYRSMIGSLLYLCASRPDIMLSVCMCARFQSDPKESHLTAVKRILRY 1715
Query: 1177 LKASPGLGLLFPRNSTINIQGYSDADWAGCPDTRRSISGYCFYIGRSLVSWKAKKQTTVS 1236
L +P GL +PR ST ++ GYSDADWAGC R+S SG C ++GRSLVSW +KKQ +V+
Sbjct: 1716 LAYTPKFGLWYPRGSTFDLIGYSDADWAGCKINRKSTSGTCQFLGRSLVSWASKKQNSVA 1775
Query: 1237 RSSNEAEYRALAYATCELQWLLYLLQDLKVTCTATPVLFCDNQSALHIAANPVFHERTKH 1296
S+ EAEY A + +L W+ L+D T P+L CDN+SA+ +A NPV H RTKH
Sbjct: 1776 LSTAEAEYIAAGHCCAQLLWMRQTLRDYGYKLTKVPLL-CDNESAIKMADNPVEHSRTKH 1834
Query: 1297 LDIDCHVVREKLQAGILKLLPIPTTLQVADVFTKALQPRVF 1337
+ I H +R+ Q G +++ I T Q+AD+FTK L + F
Sbjct: 1835 IAIRYHFLRDHQQKGDIEISYINTKDQLADIFTKPLDEQSF 1875
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.335 0.144 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,982,332,543
Number of Sequences: 2790947
Number of extensions: 78407287
Number of successful extensions: 300285
Number of sequences better than 10.0: 2095
Number of HSP's better than 10.0 without gapping: 1410
Number of HSP's successfully gapped in prelim test: 685
Number of HSP's that attempted gapping in prelim test: 293229
Number of HSP's gapped (non-prelim): 3550
length of query: 1346
length of database: 848,049,833
effective HSP length: 140
effective length of query: 1206
effective length of database: 457,317,253
effective search space: 551524607118
effective search space used: 551524607118
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 81 (35.8 bits)
Lotus: description of TM0366.1


