

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0349b.2
(400 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
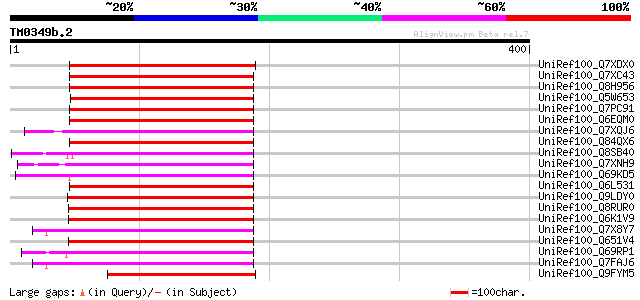
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q7XDX0 Contains similarity to ribosomal protein [Oryza... 154 3e-36
UniRef100_Q7XC43 Hypothetical protein [Oryza sativa] 154 4e-36
UniRef100_Q8H956 B protein [Oryza sativa] 152 1e-35
UniRef100_Q5W653 Hypothetical protein OSJNBb0052F16.9 [Oryza sat... 152 2e-35
UniRef100_Q7PC91 Putative transposase [Oryza sativa] 150 5e-35
UniRef100_Q6EQM0 Ribosomal protein-like [Oryza sativa] 150 5e-35
UniRef100_Q7XQJ6 OSJNBa0017B10.12 protein [Oryza sativa] 144 3e-33
UniRef100_Q84QX6 Hypothetical protein OSJNBa0093I13.9 [Oryza sat... 142 2e-32
UniRef100_Q8SB40 Putative ribosomal protein [Oryza sativa] 142 2e-32
UniRef100_Q7XNH9 OSJNBb0032D24.14 protein [Oryza sativa] 138 2e-31
UniRef100_Q69KD5 Ribosomal protein-like [Oryza sativa] 135 2e-30
UniRef100_Q6L531 Hypothetical protein OJ1005_B11.14 [Oryza sativa] 133 8e-30
UniRef100_Q9LDY0 Gb|AAF34840.1 [Arabidopsis thaliana] 129 2e-28
UniRef100_Q8RUR0 OSJNBa0026J14.30 protein [Oryza sativa] 128 2e-28
UniRef100_Q6K1V9 Ribosomal protein-like [Oryza sativa] 128 2e-28
UniRef100_Q7X8Y7 OSJNBa0085H03.2 protein [Oryza sativa] 125 3e-27
UniRef100_Q651V4 Ribosomal protein-like [Oryza sativa] 124 4e-27
UniRef100_Q69RP1 Ribosomal protein-like [Oryza sativa] 124 5e-27
UniRef100_Q7FAJ6 OSJNBb0078D11.4 protein [Oryza sativa] 121 4e-26
UniRef100_Q9FYM5 F21J9.3 [Arabidopsis thaliana] 120 5e-26
>UniRef100_Q7XDX0 Contains similarity to ribosomal protein [Oryza sativa]
Length = 308
Score = 154 bits (390), Expect = 3e-36
Identities = 69/143 (48%), Positives = 98/143 (68%)
Query: 47 RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRV 106
RK+++R+H GA+ +L DYF +P Y FRRR+RM++HVFL IV +L +YFT R+
Sbjct: 54 RKYINRNHEGAHDQLFADYFAKDPLYSAATFRRRFRMRRHVFLHIVDELGKWSSYFTHRI 113
Query: 107 DAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEY 166
+ + G SPL KCT +RMLA G AAD +DEY+K+ ++T LECL F +G++ ++ Y
Sbjct: 114 NCTGRLGHSPLQKCTAAIRMLAYGTAADTLDEYLKVPQSTALECLENFVEGVVEVFSSRY 173
Query: 167 LRAPTQEDLQRILHVNEMWGVPK 189
LR PT EDL+R+L V E G P+
Sbjct: 174 LRRPTAEDLERLLQVGESRGFPR 196
>UniRef100_Q7XC43 Hypothetical protein [Oryza sativa]
Length = 513
Score = 154 bits (389), Expect = 4e-36
Identities = 71/142 (50%), Positives = 96/142 (67%)
Query: 47 RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRV 106
RK+++R+H GA+ +L DYF +P Y FRRR+RM++HVFL IV +L +YFT RV
Sbjct: 54 RKYINRNHEGAHDQLFADYFAEDPLYSAATFRRRFRMRRHVFLHIVDELGKWSSYFTHRV 113
Query: 107 DAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEY 166
D G SPL KCT +RMLA G AAD +DEY+K+ ++T LECL F +G++ ++ Y
Sbjct: 114 DCTGCLGHSPLQKCTAAIRMLAYGTAADTLDEYLKVPQSTALECLENFVEGVVEVFSSRY 173
Query: 167 LRAPTQEDLQRILHVNEMWGVP 188
LR PT EDL+R+L V E G P
Sbjct: 174 LRRPTAEDLERLLQVGESRGFP 195
>UniRef100_Q8H956 B protein [Oryza sativa]
Length = 455
Score = 152 bits (385), Expect = 1e-35
Identities = 72/142 (50%), Positives = 96/142 (66%)
Query: 47 RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRV 106
RKH+ R A+Q L++DYF P Y IFRRR+RM + +FLRIV L +YFTQRV
Sbjct: 57 RKHIKRPREEAHQNLVNDYFSENPLYPSNIFRRRFRMYRPLFLRIVDALGQWSDYFTQRV 116
Query: 107 DAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEY 166
DAA ++G+SPL KCT +R LA G AD +DEY+KIGETT ++ ++ F KGI ++ + Y
Sbjct: 117 DAAGRQGLSPLQKCTAAIRQLATGSGADELDEYLKIGETTAMDAMKNFVKGIREVFGERY 176
Query: 167 LRAPTQEDLQRILHVNEMWGVP 188
LR PT ED +R+L + E G P
Sbjct: 177 LRRPTVEDTERLLELGERRGFP 198
>UniRef100_Q5W653 Hypothetical protein OSJNBb0052F16.9 [Oryza sativa]
Length = 444
Score = 152 bits (384), Expect = 2e-35
Identities = 74/141 (52%), Positives = 95/141 (66%)
Query: 48 KHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVD 107
K + RDH A+ RL+ DYF P Y + +FR R+RM K +FLRIV LS YFTQR D
Sbjct: 59 KTIRRDHVDAHSRLVADYFAEHPLYPERMFRTRFRMHKPLFLRIVEALSQWSPYFTQRGD 118
Query: 108 AAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYL 167
+ +SPL KCT +RMLA G ADA+DEY+KIG++T LECL F +G+I ++ YL
Sbjct: 119 CSGHTSLSPLQKCTAALRMLAYGTPADALDEYLKIGKSTALECLEMFSRGVIVVFGGTYL 178
Query: 168 RAPTQEDLQRILHVNEMWGVP 188
R PT+ED++ ILHVNE G P
Sbjct: 179 RRPTREDVEHILHVNESRGFP 199
>UniRef100_Q7PC91 Putative transposase [Oryza sativa]
Length = 482
Score = 150 bits (380), Expect = 5e-35
Identities = 71/142 (50%), Positives = 95/142 (66%)
Query: 47 RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRV 106
RKH+ R A+Q+L++DYF P Y IFRRR+RM + +FLRIV L YFTQRV
Sbjct: 91 RKHIKRPREEAHQQLVNDYFSENPLYPSKIFRRRFRMSRPLFLRIVEALGQWSVYFTQRV 150
Query: 107 DAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEY 166
DA ++G+SPL KCT +R LA G AD +DEY+KIGETT +E ++ F KG+ ++ + Y
Sbjct: 151 DAVNRKGLSPLQKCTAAIRQLATGSGADELDEYLKIGETTAMEAMKNFVKGLQDVFGERY 210
Query: 167 LRAPTQEDLQRILHVNEMWGVP 188
LR PT ED +R+L + E G P
Sbjct: 211 LRRPTMEDTERLLQLGEKRGFP 232
>UniRef100_Q6EQM0 Ribosomal protein-like [Oryza sativa]
Length = 447
Score = 150 bits (380), Expect = 5e-35
Identities = 71/142 (50%), Positives = 95/142 (66%)
Query: 47 RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRV 106
RKH+ R A+Q+L++DYF P Y IFRRR+RM + +FLRIV L YFTQRV
Sbjct: 56 RKHIKRPREEAHQQLVNDYFSENPLYPSKIFRRRFRMSRPLFLRIVEALGQWSVYFTQRV 115
Query: 107 DAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEY 166
DA ++G+SPL KCT +R LA G AD +DEY+KIGETT +E ++ F KG+ ++ + Y
Sbjct: 116 DAVNRKGLSPLQKCTAAIRQLATGSGADELDEYLKIGETTAMEAMKNFVKGLQDVFGERY 175
Query: 167 LRAPTQEDLQRILHVNEMWGVP 188
LR PT ED +R+L + E G P
Sbjct: 176 LRRPTMEDTERLLQLGEKRGFP 197
>UniRef100_Q7XQJ6 OSJNBa0017B10.12 protein [Oryza sativa]
Length = 454
Score = 144 bits (364), Expect = 3e-33
Identities = 81/177 (45%), Positives = 102/177 (56%), Gaps = 6/177 (3%)
Query: 12 DLEAYLEKSEREDTYILNRFRDRRNLILEDSAPRGRKHLSRDHAGANQRLIDDYFLNEPT 71
D +Y+E ED + + R + RK++ R+ A RL YFL P
Sbjct: 38 DTSSYIETQIEEDISYQIKKQQR------GPPTQQRKYIRRERQDAEDRLKAYYFLENPM 91
Query: 72 YDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGV 131
Y+D FR RYRM+KH+FL IV LS F QR DA K G SPL KCT +RMLA G
Sbjct: 92 YNDTQFRWRYRMKKHLFLHIVQTLSHWSPVFQQRKDAFGKVGFSPLLKCTAALRMLAYGT 151
Query: 132 AADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVNEMWGVP 188
+AD +DE ++I ETT +E L FCKGII + +YLR PT ED+QR+LHV E G P
Sbjct: 152 SADILDENLQIAETTVIESLVNFCKGIIDCFGPKYLRRPTAEDIQRLLHVGEARGFP 208
>UniRef100_Q84QX6 Hypothetical protein OSJNBa0093I13.9 [Oryza sativa]
Length = 626
Score = 142 bits (358), Expect = 2e-32
Identities = 67/142 (47%), Positives = 96/142 (67%)
Query: 47 RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRV 106
R+++ R+ +N L+ +YF P Y D +FRRR+RM+K +FLRIV+ LS YFT R+
Sbjct: 241 RRYIPRNREASNADLVANYFSESPIYTDKMFRRRFRMRKPLFLRIVSALSEWSPYFTNRL 300
Query: 107 DAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEY 166
DA + G SPL KCT +RMLA G AD +DE +KIG T LECL +F +G+I ++ +EY
Sbjct: 301 DATGRAGHSPLQKCTAAIRMLAYGTPADQLDEVLKIGPNTALECLGKFAEGVIEIFRKEY 360
Query: 167 LRAPTQEDLQRILHVNEMWGVP 188
LRAP ++++R+L V + G P
Sbjct: 361 LRAPRSDEVERLLQVADSRGFP 382
>UniRef100_Q8SB40 Putative ribosomal protein [Oryza sativa]
Length = 915
Score = 142 bits (357), Expect = 2e-32
Identities = 81/198 (40%), Positives = 119/198 (59%), Gaps = 12/198 (6%)
Query: 2 ISQKMDPPE-FDLEAYLEKSEREDTYILNRFRDRRNLILEDSA---PRGR-------KHL 50
+ +++DP E + LE +L + E +++ + D+ +E S+ PR R +++
Sbjct: 473 VLEEIDPAEVYTLEDFLAEDEIMESF-RRKIGDKLKAKIEGSSSGPPRRRQRQSGPRRYI 531
Query: 51 SRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAK 110
R ++ L+ +YF P Y D FRRR+RM K +FLRIV LS+ D +FTQRVDA
Sbjct: 532 PRPREKGHEDLVANYFSANPIYTDEQFRRRFRMNKPLFLRIVNALSNWDQFFTQRVDATG 591
Query: 111 KEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAP 170
++ SPL KCT +RML G ADA+DE +KI +T+LECL +F GII + EYLR P
Sbjct: 592 RDSHSPLQKCTAAIRMLGYGTPADALDEVLKIAASTSLECLGKFAVGIIECFGSEYLRPP 651
Query: 171 TQEDLQRILHVNEMWGVP 188
T ++L++IL NE G P
Sbjct: 652 TSDELEKILQENEARGFP 669
>UniRef100_Q7XNH9 OSJNBb0032D24.14 protein [Oryza sativa]
Length = 450
Score = 138 bits (348), Expect = 2e-31
Identities = 77/182 (42%), Positives = 103/182 (56%), Gaps = 6/182 (3%)
Query: 7 DPPEFDLEAYLEKSEREDTYILNRFRDRRNLILEDSAPRGRKHLSRDHAGANQRLIDDYF 66
DP E LEA LE + + ++ N + R+++SRDH + RL YF
Sbjct: 31 DPMEAQLEARLEA--QLEARLMAHLAGSSNQL----GGYTRRYISRDHEDDHNRLFAKYF 84
Query: 67 LNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRM 126
P Y D FRRR+RM++H+FLRIV L YF R DA K G+SPL KCT MRM
Sbjct: 85 SESPLYTDDQFRRRFRMRRHLFLRIVQALGVWSPYFRLRRDAFGKVGLSPLQKCTAAMRM 144
Query: 127 LANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVNEMWG 186
LA G AD +DE + E+T +EC+ F +G+ L+ ++YLR PT ED+QR+L E G
Sbjct: 145 LAYGTPADLMDETFGVAESTAMECMINFVQGVRHLFGEQYLRRPTVEDIQRLLQFGEAHG 204
Query: 187 VP 188
P
Sbjct: 205 FP 206
>UniRef100_Q69KD5 Ribosomal protein-like [Oryza sativa]
Length = 510
Score = 135 bits (341), Expect = 2e-30
Identities = 72/188 (38%), Positives = 108/188 (57%), Gaps = 4/188 (2%)
Query: 5 KMDPPE-FDLEAYLEKSEREDTYILNRFRDRRNLILEDSAP---RGRKHLSRDHAGANQR 60
++DP E + +E YL++ + + + + S+ R R++ RD G + R
Sbjct: 14 EVDPTEIYTMEDYLQRQQAWNAAAAQLASHLLHTLGPSSSQQPRRRRRYCRRDRIGGHAR 73
Query: 61 LIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKC 120
++ DYF P Y D FRRR+RM++ +FLRI+ L YFTQRVD + G+SP+ KC
Sbjct: 74 IMADYFCENPVYSDRDFRRRFRMRRPLFLRILNALGEWSEYFTQRVDGIGQLGLSPIQKC 133
Query: 121 TTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILH 180
T RMLA +AD VDEY +IG +T LE L+ FC+G+I ++ EYL P +D R+L
Sbjct: 134 TAVNRMLAYAASADQVDEYCRIGGSTALESLKLFCQGVIEIFGPEYLHQPNVDDTGRLLQ 193
Query: 181 VNEMWGVP 188
++ G P
Sbjct: 194 IHGSRGFP 201
>UniRef100_Q6L531 Hypothetical protein OJ1005_B11.14 [Oryza sativa]
Length = 443
Score = 133 bits (335), Expect = 8e-30
Identities = 64/142 (45%), Positives = 92/142 (64%)
Query: 47 RKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRV 106
R+++ R+ + L+ +YF P Y D +FRRR+RM++ +FLRIV +LS YFTQRV
Sbjct: 59 RRYIPRNRERGHDDLVANYFSANPIYTDEMFRRRFRMRRPLFLRIVHELSEWSPYFTQRV 118
Query: 107 DAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEY 166
DA + +SPL KC +RML G AAD +DE ++I +T LECL +F +G+I + EY
Sbjct: 119 DATGRNSLSPLQKCIAAIRMLGYGTAADQLDEVLRIAASTCLECLGKFAEGVIEKFGDEY 178
Query: 167 LRAPTQEDLQRILHVNEMWGVP 188
LR P ++L++IL NE G P
Sbjct: 179 LRLPRADELEKILQENEARGFP 200
>UniRef100_Q9LDY0 Gb|AAF34840.1 [Arabidopsis thaliana]
Length = 415
Score = 129 bits (323), Expect = 2e-28
Identities = 66/144 (45%), Positives = 92/144 (63%)
Query: 45 RGRKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQ 104
R R + R + L +DYF + + FRRR+RM+K +FLRIV LSS +F
Sbjct: 43 RKRAYFDRKREEGHVLLWNDYFSDNAIFPLQTFRRRFRMKKPLFLRIVDRLSSELMFFQH 102
Query: 105 RVDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQ 164
R DA + G SP+ KCT +R+LA G A+DAVDEY+++GETT + CL F KGII +
Sbjct: 103 RRDATGRFGHSPIQKCTAAIRLLAYGYASDAVDEYLRMGETTAMSCLENFTKGIISFFGD 162
Query: 165 EYLRAPTQEDLQRILHVNEMWGVP 188
EYLRAPT +L+R+L++ ++ G P
Sbjct: 163 EYLRAPTATNLRRLLNIGKIRGFP 186
>UniRef100_Q8RUR0 OSJNBa0026J14.30 protein [Oryza sativa]
Length = 626
Score = 128 bits (322), Expect = 2e-28
Identities = 64/143 (44%), Positives = 90/143 (62%)
Query: 46 GRKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQR 105
G + + R+ + RL DYF N PTY +FRRR RM + +FLRI+ + D+YF Q+
Sbjct: 248 GHEVIDRNREARHLRLYQDYFSNNPTYGPVLFRRRNRMSRPLFLRIMNAIEDHDDYFVQK 307
Query: 106 VDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQE 165
+AA G S K T MR LA G+AADA+DEY+ I E+T +E LRRF K +++++E E
Sbjct: 308 RNAAGLIGFSCHQKVTAAMRQLAYGIAADALDEYLGIAESTAIESLRRFVKAVVQVFEHE 367
Query: 166 YLRAPTQEDLQRILHVNEMWGVP 188
YLR+P + D R+L + E G P
Sbjct: 368 YLRSPNENDTTRLLELGEDRGFP 390
>UniRef100_Q6K1V9 Ribosomal protein-like [Oryza sativa]
Length = 916
Score = 128 bits (322), Expect = 2e-28
Identities = 64/143 (44%), Positives = 90/143 (62%)
Query: 46 GRKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQR 105
G + + R+ + RL DYF N PTY +FRRR RM + +FLRI+ + D+YF Q+
Sbjct: 538 GHEVIDRNREARHLRLYQDYFSNNPTYGPVLFRRRNRMSRPLFLRIMNAIEDHDDYFVQK 597
Query: 106 VDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQE 165
+AA G S K T MR LA G+AADA+DEY+ I E+T +E LRRF K +++++E E
Sbjct: 598 RNAAGLIGFSCHQKVTAAMRQLAYGIAADALDEYLGIAESTAIESLRRFVKAVVQVFEHE 657
Query: 166 YLRAPTQEDLQRILHVNEMWGVP 188
YLR+P + D R+L + E G P
Sbjct: 658 YLRSPNENDTTRLLELGEDRGFP 680
>UniRef100_Q7X8Y7 OSJNBa0085H03.2 protein [Oryza sativa]
Length = 440
Score = 125 bits (313), Expect = 3e-27
Identities = 65/175 (37%), Positives = 100/175 (57%), Gaps = 4/175 (2%)
Query: 18 EKSEREDTYI----LNRFRDRRNLILEDSAPRGRKHLSRDHAGANQRLIDDYFLNEPTYD 73
E S+ +D YI L +R + GR+ + RD N R++ DYF + P Y
Sbjct: 27 ELSDDDDYYIAAALLTDIEHKRTKRKHHGSVPGREIIHRDRFAGNLRIVADYFADPPIYS 86
Query: 74 DGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGVAA 133
+FRRR+RM + +FLRIVA + + D+YF QR +A G + L K +RMLA + A
Sbjct: 87 AKLFRRRFRMSRELFLRIVASVEAHDDYFRQRPNAVGLLGATALQKVYGAIRMLAYDIPA 146
Query: 134 DAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVNEMWGVP 188
D++DE ++I E+T +E + F K ++ ++ +YLRAPT ED R++ +N G P
Sbjct: 147 DSLDEVVRISESTMIEAFKHFVKAVVDVFADQYLRAPTAEDTARLMAINTPRGFP 201
>UniRef100_Q651V4 Ribosomal protein-like [Oryza sativa]
Length = 373
Score = 124 bits (312), Expect = 4e-27
Identities = 58/143 (40%), Positives = 97/143 (67%)
Query: 46 GRKHLSRDHAGANQRLIDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQR 105
G K RD A+ +L +DYF+ P Y++ FRRR+RM++ +FLRIV ++++ + +FTQR
Sbjct: 45 GHKTYKRDRIAADWQLNEDYFVERPLYNEEHFRRRFRMRRELFLRIVDEVTAKNRFFTQR 104
Query: 106 VDAAKKEGISPLAKCTTTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQE 165
+AA + G S L KCT ++MLA G A+ +D+++K+GE+T L+ L+ F +I+++ +E
Sbjct: 105 RNAAGQLGFSALHKCTVALKMLAYGGPANELDDHLKMGESTALKTLKEFIMTVIKVFGKE 164
Query: 166 YLRAPTQEDLQRILHVNEMWGVP 188
+LR P E+++ IL +N G P
Sbjct: 165 FLRPPRSEEIEHILSINLARGFP 187
>UniRef100_Q69RP1 Ribosomal protein-like [Oryza sativa]
Length = 258
Score = 124 bits (311), Expect = 5e-27
Identities = 70/187 (37%), Positives = 106/187 (56%), Gaps = 10/187 (5%)
Query: 10 EFDLEAYLEKSEREDTYILNRFRDRRNLILEDS--------APRGRKHLSRDHAGANQRL 61
EF + ++ S ++ + L F +I+E+S + G + + RD + L
Sbjct: 31 EFVMNELIDPSSSDEEHDL--FFGAAQMIIEESVNNPGRIGSVHGHEVVHRDRLLWHNLL 88
Query: 62 IDDYFLNEPTYDDGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCT 121
DYF + PT+ IFRRR+RM+ HVFLRI+ + D+YF Q+ +AA G+S L K
Sbjct: 89 YKDYFSDNPTFGPNIFRRRFRMRIHVFLRIMKAVEEHDDYFKQKRNAAGVLGLSCLQKVV 148
Query: 122 TTMRMLANGVAADAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHV 181
MLA GV ADA+DEYI+IGE+T LE LR+F ++ ++ EY+R P ++D R+L +
Sbjct: 149 AAFCMLAYGVPADALDEYIRIGESTALEALRKFVAAVVEVFGPEYMRKPNEQDTARLLTI 208
Query: 182 NEMWGVP 188
G P
Sbjct: 209 GASRGFP 215
>UniRef100_Q7FAJ6 OSJNBb0078D11.4 protein [Oryza sativa]
Length = 420
Score = 121 bits (303), Expect = 4e-26
Identities = 64/175 (36%), Positives = 99/175 (56%), Gaps = 4/175 (2%)
Query: 18 EKSEREDTYI----LNRFRDRRNLILEDSAPRGRKHLSRDHAGANQRLIDDYFLNEPTYD 73
E S+ +D YI L +R + GR+ + RD N R++ DYF + P Y
Sbjct: 22 ELSDDDDYYIAAALLTDIEHKRTKRKRRGSVPGREIIHRDRFAGNLRIVADYFADPPVYS 81
Query: 74 DGIFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGVAA 133
+FRRR+RM + +FLRIVA + + D+YF QR +A G + L K + MLA + A
Sbjct: 82 AKLFRRRFRMSRELFLRIVASVEAHDDYFRQRPNAVGLLGATALQKVYGAICMLAYDIPA 141
Query: 134 DAVDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVNEMWGVP 188
D++DE ++I E+T +E + F K ++ ++ +YLRAPT ED R++ +N G P
Sbjct: 142 DSLDEVVRISESTMIEAFKHFVKAMVDVFADQYLRAPTAEDTTRLMAINTPRGFP 196
>UniRef100_Q9FYM5 F21J9.3 [Arabidopsis thaliana]
Length = 457
Score = 120 bits (302), Expect = 5e-26
Identities = 58/114 (50%), Positives = 76/114 (65%)
Query: 76 IFRRRYRMQKHVFLRIVADLSSSDNYFTQRVDAAKKEGISPLAKCTTTMRMLANGVAADA 135
+FRRR+RM + +FLRI + DNYF QR D K G+S L K T RMLA GV AD+
Sbjct: 1 MFRRRFRMSRSLFLRIYDAIQRHDNYFVQRRDRVGKLGLSGLQKMTAAFRMLAYGVPADS 60
Query: 136 VDEYIKIGETTTLECLRRFCKGIIRLYEQEYLRAPTQEDLQRILHVNEMWGVPK 189
DEYIKIGE+T LE L+RFC+ I+ ++ YLR+P D+ R+LH+ E G P+
Sbjct: 61 TDEYIKIGESTALESLKRFCRAIVEVFACRYLRSPDANDVARLLHIGESRGFPR 114
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.345 0.152 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 592,405,580
Number of Sequences: 2790947
Number of extensions: 21684023
Number of successful extensions: 80927
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 41
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 80880
Number of HSP's gapped (non-prelim): 50
length of query: 400
length of database: 848,049,833
effective HSP length: 129
effective length of query: 271
effective length of database: 488,017,670
effective search space: 132252788570
effective search space used: 132252788570
T: 11
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.6 bits)
S2: 76 (33.9 bits)
Lotus: description of TM0349b.2


