

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0348.12
(180 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
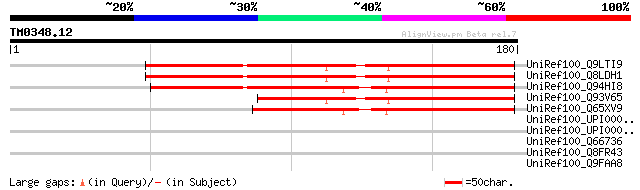
UniRef100_Q9LTI9 Arabidopsis thaliana genomic DNA, chromosome 5,... 120 1e-26
UniRef100_Q8LDH1 Hypothetical protein [Arabidopsis thaliana] 118 8e-26
UniRef100_Q94HI8 Hypothetical protein [Oryza sativa] 106 3e-22
UniRef100_Q93V65 AT5g59400/f2o15_60 [Arabidopsis thaliana] 86 4e-16
UniRef100_Q65XV9 Hypothetical protein P0016H04.9 [Oryza sativa] 79 7e-14
UniRef100_UPI0000311342 UPI0000311342 UniRef100 entry 35 0.68
UniRef100_UPI0000328D76 UPI0000328D76 UniRef100 entry 35 0.89
UniRef100_Q66736 Envelope protein [Equine infectious anemia virus] 34 1.5
UniRef100_Q8FR43 Hypothetical protein [Corynebacterium efficiens] 33 3.4
UniRef100_Q9FAA8 ORF-Y [Geobacillus thermoleovorans] 32 9.9
>UniRef100_Q9LTI9 Arabidopsis thaliana genomic DNA, chromosome 5, BAC clone:F2O15
[Arabidopsis thaliana]
Length = 282
Score = 120 bits (302), Expect = 1e-26
Identities = 67/136 (49%), Positives = 85/136 (62%), Gaps = 9/136 (6%)
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGI 108
W GS+S V YPRCS+ RQSTYADAE + L W + FGS AC+ P Y VG+
Sbjct: 106 WYGSKSVVKYPRCSLLRQSTYADAEDDASQVLLLA-TVWIMIFLFGSSACVLPTIYGVGL 164
Query: 109 AYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGFVIGYPLASAPVKVIQGLWRN 163
Y + F SGL + S L S+ + N I+ VLG GYP+AS+ V+V++GLWRN
Sbjct: 165 VYGGDPFDSGLVYSSQ---LSSSVPILSKFNGILLSVLGPAFGYPIASSAVRVLKGLWRN 221
Query: 164 DLVALRGACPNCGEEL 179
DL AL+G CPNCGEE+
Sbjct: 222 DLTALKGDCPNCGEEV 237
>UniRef100_Q8LDH1 Hypothetical protein [Arabidopsis thaliana]
Length = 299
Score = 118 bits (295), Expect = 8e-26
Identities = 65/136 (47%), Positives = 84/136 (60%), Gaps = 9/136 (6%)
Query: 49 WLGSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGI 108
W GS+S V YPRCS+ RQSTYADAE + L W + FGS AC+ P Y VG+
Sbjct: 106 WYGSKSVVKYPRCSLLRQSTYADAEDDASQVLLLA-TVWIMIFLFGSSACVLPTIYGVGL 164
Query: 109 AYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGFVIGYPLASAPVKVIQGLWRN 163
Y + F SGL + + L S+ + N I+ VLG GYP+AS+ V+ ++GLWRN
Sbjct: 165 VYGGDPFDSGLVYSNQ---LSSSVPILSKFNGILLSVLGPAFGYPIASSAVRALKGLWRN 221
Query: 164 DLVALRGACPNCGEEL 179
DL AL+G CPNCGEE+
Sbjct: 222 DLTALKGDCPNCGEEV 237
>UniRef100_Q94HI8 Hypothetical protein [Oryza sativa]
Length = 272
Score = 106 bits (264), Expect = 3e-22
Identities = 61/132 (46%), Positives = 87/132 (65%), Gaps = 8/132 (6%)
Query: 51 GSRSDV*YPRCSIRRQSTYADAEGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGIAY 110
GS S V YPRCS++RQSTYADAE + L W L L FG+ A L P + + +
Sbjct: 83 GSPSVVKYPRCSLKRQSTYADAEEDKSMFMALS-SIWMLLLLFGTSAFLVPSLCILSLTF 141
Query: 111 QNAFGSG-LSHGSHNPGLGFSII--VNNIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVA 167
+AFG+ L +G+ + F +I VN+++ I LG++IGYP++SA V +QGL N+LVA
Sbjct: 142 GDAFGARYLLYGAKS----FDVITRVNDMVLIGLGYLIGYPISSASVGALQGLLTNNLVA 197
Query: 168 LRGACPNCGEEL 179
L+G+CPNCGE++
Sbjct: 198 LKGSCPNCGEQV 209
>UniRef100_Q93V65 AT5g59400/f2o15_60 [Arabidopsis thaliana]
Length = 157
Score = 85.9 bits (211), Expect = 4e-16
Identities = 46/96 (47%), Positives = 61/96 (62%), Gaps = 8/96 (8%)
Query: 89 LFLAFGSLACLGPLSYAVGIAYQ-NAFGSGLSHGSHNPGLGFSIIV----NNIIFIVLGF 143
+ FGS AC+ P Y VG+ Y + F SGL + S L S+ + N I+ VLG
Sbjct: 1 MIFLFGSSACVLPTIYGVGLVYGGDPFDSGLVYSSQ---LSSSVPILSKFNGILLSVLGP 57
Query: 144 VIGYPLASAPVKVIQGLWRNDLVALRGACPNCGEEL 179
GYP+AS+ V+V++GLWRNDL AL+G CPNCGEE+
Sbjct: 58 AFGYPIASSAVRVLKGLWRNDLTALKGDCPNCGEEV 93
>UniRef100_Q65XV9 Hypothetical protein P0016H04.9 [Oryza sativa]
Length = 163
Score = 78.6 bits (192), Expect = 7e-14
Identities = 42/96 (43%), Positives = 65/96 (66%), Gaps = 7/96 (7%)
Query: 87 WALFLAFGSLACLGPLSYAVGIAYQNAFGSG-LSHGSHNPGLGFSII--VNNIIFIVLGF 143
W L L FG+ A L P + + + +AFG+ L +G+ + F +I VN+++ I LG+
Sbjct: 9 WMLLLLFGTSAFLVPSLCILSLTFGDAFGARYLLYGAKS----FDVITRVNDMVLIGLGY 64
Query: 144 VIGYPLASAPVKVIQGLWRNDLVALRGACPNCGEEL 179
+IGYP++SA V +QGL N+LVAL+G+CPNCGE++
Sbjct: 65 LIGYPISSASVGALQGLLTNNLVALKGSCPNCGEQV 100
>UniRef100_UPI0000311342 UPI0000311342 UniRef100 entry
Length = 153
Score = 35.4 bits (80), Expect = 0.68
Identities = 36/124 (29%), Positives = 50/124 (40%), Gaps = 36/124 (29%)
Query: 59 PRCSIRRQSTYADA---EGRRIYLWFLH*QAWALFLAFGSLACLGPLSYAVGIAYQNAFG 115
PRCS+R ++TY+D EG Q WAL+ ++ LG + V +A + FG
Sbjct: 1 PRCSVRGEATYSDCVVDEG----------QMWALWTSY-----LGVFAIGVSLAVFD-FG 44
Query: 116 SGL--SHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVIQGLWRNDLVALRGACP 173
L S G+ P LG V G LA + +A+ G CP
Sbjct: 45 GALDPSEGARAPVLG---------------VAGCALARRGWDKLAQTMEGSTLAMTGECP 89
Query: 174 NCGE 177
C E
Sbjct: 90 ACHE 93
>UniRef100_UPI0000328D76 UPI0000328D76 UniRef100 entry
Length = 217
Score = 35.0 bits (79), Expect = 0.89
Identities = 22/92 (23%), Positives = 41/92 (43%)
Query: 85 QAWALFLAFGSLACLGPLSYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFV 144
Q+ ++ FG GPL + + +N++ ++ GL +I+++ I+ G V
Sbjct: 16 QSSEVYGGFGGCWDYGPLGVELKLNIKNSWWRSMTFREDIVGLDSTILMHPDIWKASGHV 75
Query: 145 IGYPLASAPVKVIQGLWRNDLVALRGACPNCG 176
G+ K + +R D + L CP CG
Sbjct: 76 DGFSDPLVDCKQCKERFREDQIDLNAPCPKCG 107
>UniRef100_Q66736 Envelope protein [Equine infectious anemia virus]
Length = 595
Score = 34.3 bits (77), Expect = 1.5
Identities = 20/51 (39%), Positives = 32/51 (62%), Gaps = 7/51 (13%)
Query: 114 FGSGL-SH-GSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVIQGLWR 162
FG L SH + NPGLG SII ++F+++ Y L ++P K+++ LW+
Sbjct: 344 FGKDLWSHIANWNPGLGASIIKYIVVFLLI-----YLLLTSPPKILRALWK 389
>UniRef100_Q8FR43 Hypothetical protein [Corynebacterium efficiens]
Length = 149
Score = 33.1 bits (74), Expect = 3.4
Identities = 32/104 (30%), Positives = 46/104 (43%), Gaps = 26/104 (25%)
Query: 79 LWFLH*QAWAL--FLAFGSLACLGPLSYAVGIA-YQNA------FGSGLSHGSHNPGLGF 129
+W L W ++ FG LAC+ ++ GIA ++ A FG + +H G G
Sbjct: 20 IWLLFGGIWLALGYVLFGILACILIITIPAGIASFRMANYALWPFGRTVVRKTHG-GNGL 78
Query: 130 SIIVNNIIFIVLGF----------------VIGYPLASAPVKVI 157
S I N I FI+ G +IG PLA A +K+I
Sbjct: 79 SAISNFIWFIIAGLWLAIGHITTAAAQAITIIGIPLAIANIKMI 122
>UniRef100_Q9FAA8 ORF-Y [Geobacillus thermoleovorans]
Length = 343
Score = 31.6 bits (70), Expect = 9.9
Identities = 20/67 (29%), Positives = 32/67 (46%), Gaps = 1/67 (1%)
Query: 99 LGPL-SYAVGIAYQNAFGSGLSHGSHNPGLGFSIIVNNIIFIVLGFVIGYPLASAPVKVI 157
+GPL SYA+G A G + H P LG +++ IF+++ + +P+ P K
Sbjct: 255 VGPLLSYALGWILPCAVGFLFAANRHRPVLGSVLVMIGAIFVLIVGISVFPMLLLPRKRA 314
Query: 158 QGLWRND 164
W D
Sbjct: 315 HVGWAGD 321
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.340 0.149 0.512
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 288,123,278
Number of Sequences: 2790947
Number of extensions: 10850903
Number of successful extensions: 32839
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 32824
Number of HSP's gapped (non-prelim): 10
length of query: 180
length of database: 848,049,833
effective HSP length: 119
effective length of query: 61
effective length of database: 515,927,140
effective search space: 31471555540
effective search space used: 31471555540
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 70 (31.6 bits)
Lotus: description of TM0348.12


